Re-expression of SMARCA4/BRG1 in small cell carcinoma of ovary, hypercalcemic type (SCCOHT) promotes an epithelial-like gene signature through an AP-1-dependent mechanism
Figures
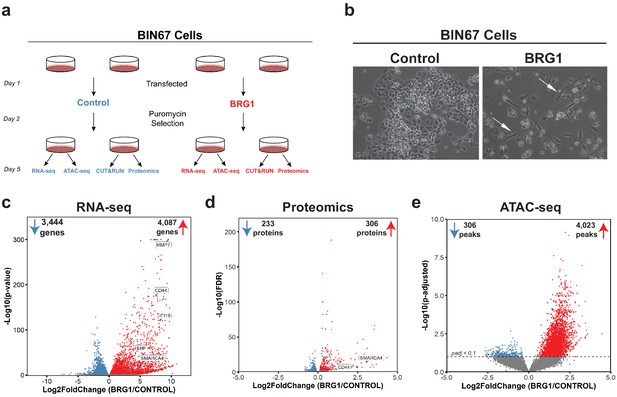
BRG1 reexpression induces global morphological, transcriptomic, and epigenetic changes in BIN67 cells.
(a) Experimental design for BRG1 reexpression in BIN67 cells and multi-omic data analysis. (b) Following the experimental design in 1a, pictures were taken at 10X phase. BIN67 cells transfected with BRG1 show an elongated morphology (white arrows) relative to control. Additional fields and controls shown in Figure 1—figure supplement 1. (c) Volcano plot results of RNA-seq differential gene expression (BRG1/Control) for protein coding genes using DESeq2 (n = 18,507 genes total). Significantly upregulated genes (padj <0.05 and Log2FoldChange > 0) are colored in red (n = 4087 genes). Significantly downregulated genes (padj <0.05 and Log2FoldChange < 0) are colored in blue (n = 3444 genes). Non-significant genes are colored in gray (n = 10,976 genes). SMARCA4/BRG1 and BRG1 target genes are identified. (d) Volcano plot of differential expressed proteins for BIN67 +/- BRG1 reexpression by PECA analysis (n = 5726 total proteins identified). Significantly upregulated proteins (p.fdr <0.05 and Log2FoldChange > 0) are colored in red (n = 306 proteins). Significantly downregulated proteins (p.fdr <0.05 and Log2FoldChange < 0) are colored in blue (n = 233 proteins). Non-significant proteins are identified in gray (n = 5187 proteins). Proteins of SMARCA4/BRG1 and CD44 are identified. (e) Differential peak analysis for ATAC-seq data using DESeq2 (n = 62,308 peaks total). Significantly gained ATAC-seq peaks (padj <0.10 and Log2FoldChange > 0) are identified in red (Gained; n = 4,023). Significantly lost ATAC-seq peaks (padj <0.10 and Log2FoldChange < 0) are identified in blue (Lost; n = 306). Non-significant peaks are identified in Gray (Static/N.S.; n = 57,979). Source data is available in Figure 1—source data 1.
-
Figure 1—source data 1
Raw data for Figure 1.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig1-data1-v2.xlsx
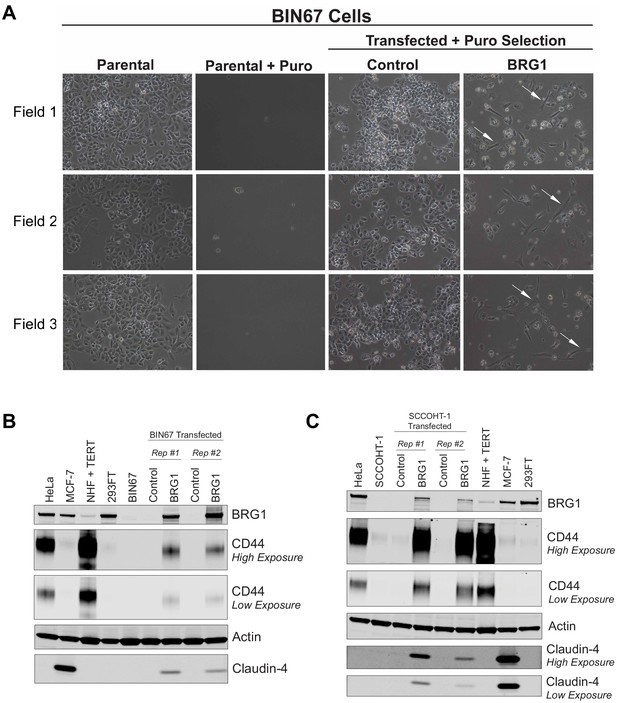
Morphology and protein expression changes in SCCOHT cells following BRG1 reexpression.
(A) Three different fields taken randomly across two biological replicates of BIN67 cells +/- BRG1 reexpression flowing experimental design in Figure 1a. Pictures were taken at 10X phase prior to harvest. BIN67 cells transfected with BRG1 show an elongated morphology (white arrows) relative to control. BIN67 (parental) cells at similar passage are shown for morphology comparison. Untransfected controls treated with puromycin (Parental + Puro) are shown for each experiment. Control and BRG1 transfected for Field one are shown in Figure 1b. Field one is from biological replicate #1 and Field two and Field three are from biological replicate #2. (B, C) Western blots for (B) BIN67 and (C) SCCOHT-1 cells +/- BRG1 compared to other cancer cell lines. Reexpression studies were performed in biological duplicate.
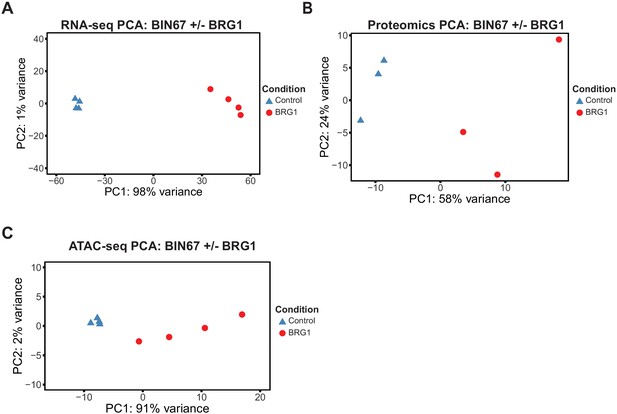
Unsupervised analysis for multi-omics samples.
Principal component analysis (PCA) for RNA-seq (A), Proteomics (B), ATAC-seq (C). A, PCA for RNA-seq samples was performed by DESeq2 on rlog transformed counts. (B) PCA was performed by prcomp on log2 transformed counts for proteomics samples. (C) PCA was performed on rlog transformed counts under called ATAC-seq peaks by DESeq2.
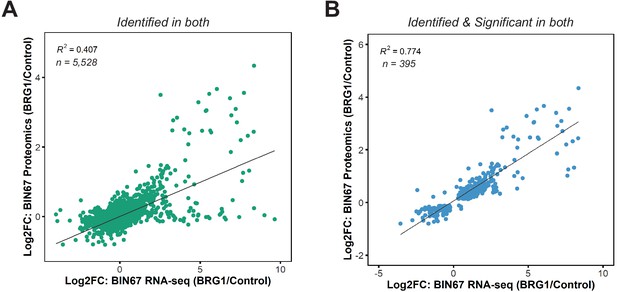
Correlation of RNA-seq and proteomics data in BIN67 cells.
(A) Scatterplot of gene expression changes and protein expression changes for proteins and genes identified in both (n = 5528). For instances where multiple proteins mapped to a particular gene, the average log2foldchange of protein abundance was calculated. (B) Scatterplot of gene expression changes and protein expression changes for proteins and genes identified as significant in both (RNA-seq: padj <0.05, Proteomics: FDR < 0.05). For instances where multiple proteins mapped to a particular gene, the average log2foldchange of protein abundance was calculated.
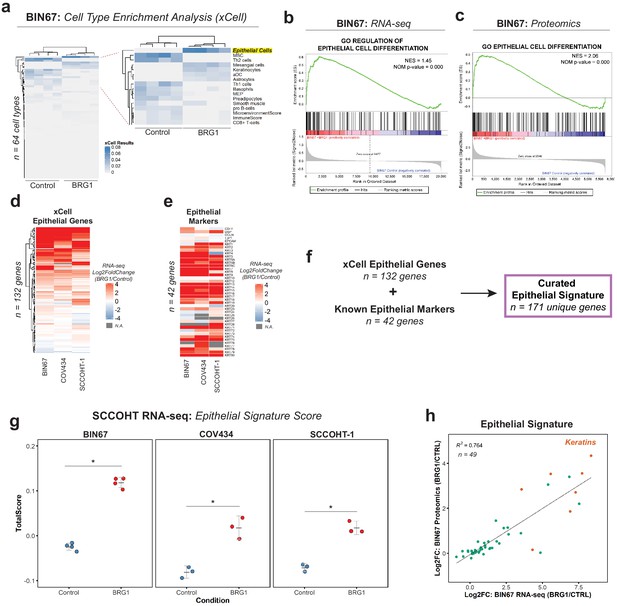
BRG1 reexpression induces an epithelial-like expression signature and chromatin profile.
(a) Heatmap of xCell results from RNA-seq data (gene and library scaled TPMs for input). Epithelial cell category highlighted in yellow. Rows and columns clustered by Pearsons correlation. (b) Enrichment plots for epithelial related Gene Ontology (GO) terms in RNA-seq data (VST normalized TPMs) using Gene Set Enrichment Analysis (GSEA; MSigDB GO gene set - C5 all v6.0). (c) Enrichment plots for epithelial related GO terms in proteomics data (log2 transformed) using GSEA (MSigDB GO gene set - C5 all v6.0). (d) Heatmap of RNA-seq Log2FoldChange (BRG1/Control) for xCell Epithelial Gene set (n = 132) in BIN67, SCCOHT-1, and COV434 cells clustered by Euclidean correlation. (e) Heatmap of RNA-seq Log2FoldChange (BRG1/Control) for epithelial cell markers in BIN67, SCCOHT-1, and COV434 cells clustered by Euclidean correlation. For genes that no measurable Log2FoldChange was calculated by DESeq2 are colored in gray. (f) Definition of the curated epithelial gene signature. (g) Scoring of the epithelial signature in RNA-seq data from SCCOHT cell lines using singscore. (h) Scatterplot of gene expression change and associated protein expression changes in BIN67 cells for genes and proteins identified in common from the epithelial signature. Source data is available in Figure 2—source data 1.
-
Figure 2—source data 1
Raw data for Figure 2.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig2-data1-v2.xlsx
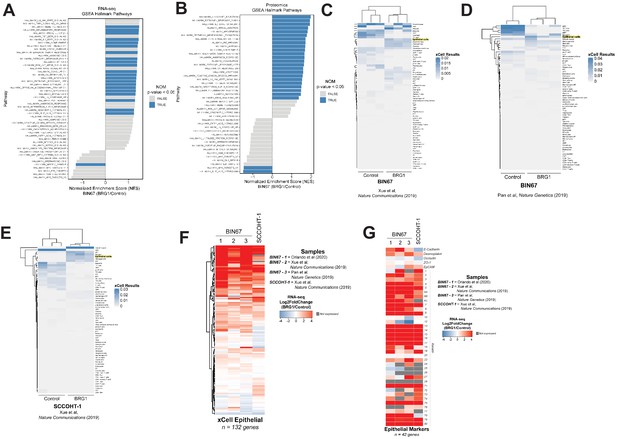
Epithelial genes are upregulated following BRG1 expression in SCCOHT cell lines.
(A) Gene set enrichment analysis (GSEA) performed on RNA-seq data (VST normalized TPMs) using the Hallmark gene set (MSigDB Hallmark v6.0). (B) GSEA performed on Proteomics data (log2 transformed) using the Hallmark gene set (MSigDB Hallmark v6.0). (C,D,E) Heatmap of xCell results for previously published data sets of BIN67 +/- BRG1 and SCCOHT-1 +/- BRG1 and clustered by Euclidean distance. Epithelial cell category highlighted in yellow. (F,G) Publically available BIN67 RNA-seq data (+/- BRG1; columns 2 and 3), SCCOHT-1 cell line (+/- BRG1) was compared to BIN67 cells (+/- BRG1) in this study (column 1) for Log2FoldChange (BRG1/Control) for genes in the xCell epithelial list (F) and epithelial cell marker list (G).
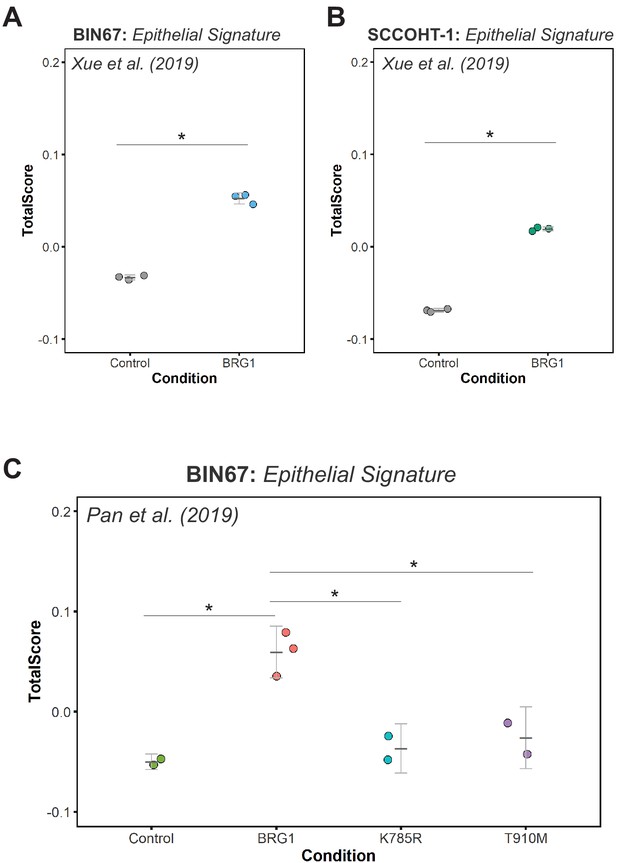
Epithelial signature scoring in previously published SCCOHT datasets.
Scoring of epithelial signature in RNA-seq data using singscore for: (A) BIN67 +/- BRG1; Xue et al., 2019b, (*Welsh Two Sample t-test p-value=0.00028) (B) SCCOHT-1 +/- BRG1; Xue et al., 2019b, (*Welsh Two-Sample t-test p-value=4.367 e-07) and (C) BIN67 +/- BRG1/K785R/T910M; Pan et al., 2019 (Welsh Two-Sample t-test p-values: Control compared to wild-type BRG1 (p-value=0.0097)), BRG1 wildtype to K785R mutant (p-value=0.0162) and BRG1 wild-type to T910M mutant (p-value=0.0405).
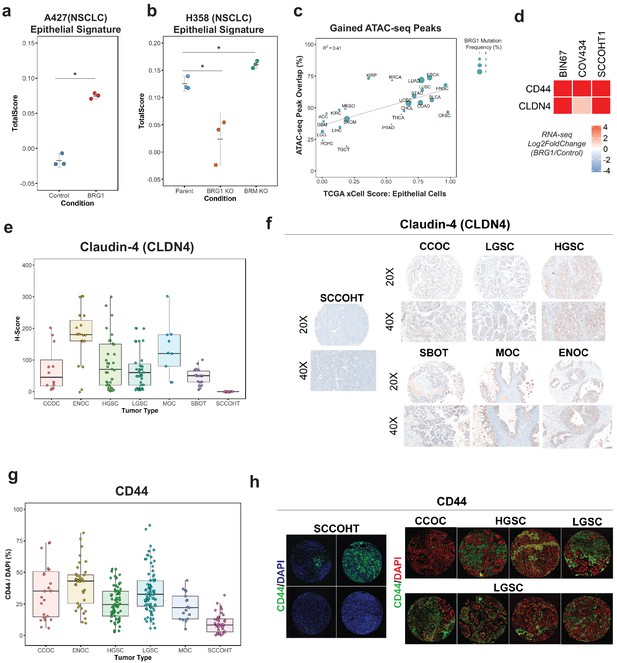
Loss of the epithelial signature is broadly applicable to BRG1-deficient NSCLC cell lines and SCCOHT primary tumors.
(a, b) Scoring of epithelial signature in RNA-seq data from NSCLC cell lines +/- BRG1 expression (A427) and +/- BRG1/BRM knockout (H358). (C) Scatterplot of percent peak overlap of gained ATAC-seq peaks with TCGA ATAC-seq peaks versus the averaged xCell calculated epithelial cell score for each TCGA tumor type. The frequency of BRG1 mutations identified per tumor type (reported as a percentage) is indicated by the size of the point. (d) Heatmap of CD44 and Claudin-4 (CLDN4) gene expression changes following BRG1 expression in SCCOHT cell lines. (e, f) Immunohistochemistry scoring and representative images for Claudin-4 expression from a TMA containing gynecological tumors. (g, h) Immunofluorescence scoring and representative images for CD44 expression at ×20 magnification from a TMA containing gynecological tumors. In both Claudin-4 and CD44 staining (e–h), multiple core samples (between 2 and 6 cores) were taken per tumor and reported as single points in boxplots. Total numbers reported as the number of cores taken per tumor type. Small Cell Carcinoma of the Ovary, Hypercalcemic Type (SCCOHT; n = 16 cores for CLDN4, n = 39 cores for CD44). Clear Cell Ovarian Carcinoma (CCOC; n = 12 cores for CLDN4, n = 23 cores for CD44). High-Grade Serous Ovarian Carcinoma (HGSC; n = 33 cores for CLDN4, n = 64 cores for CD44). Low-Grade Serous Ovarian Carcinoma (LGSC; n = 38 cores for CLDN4, n = 83 cores for CD44). Endometriod Ovarian Carcinoma (ENOC; n = 19 cores for CLDN4, n = 37 cores for CD44). Mucinous Ovarian Carcinoma (MOC; n = 9 cores for CLDN4, n = 17 cores for CD44). Serous Borderline Ovarian Tumor, pre-cursor legion of LGSC (SBOT; n = 18 cores for CLDN4; CD44 staining not performed for this tumor type). Source data is available in Figure 3—source data 1.
-
Figure 3—source data 1
Raw data for Figure 3.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig3-data1-v2.xlsx
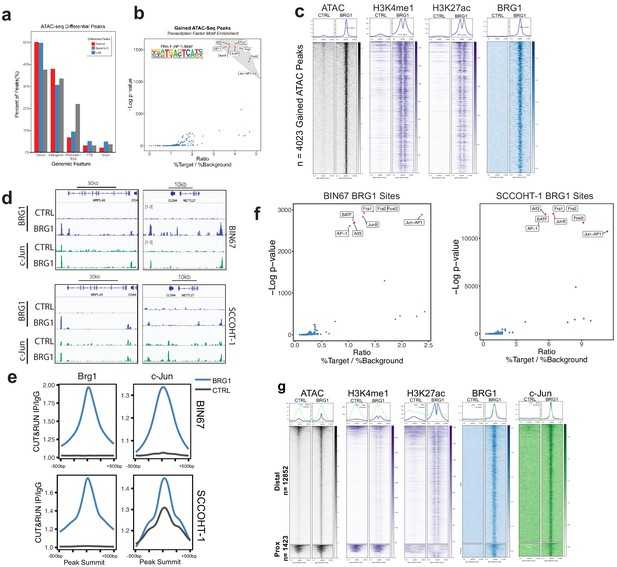
AP-1 motifs are enriched at BRG1 recruited chromatin and gained chromatin accessibility.
(a) Genomic annotations by HOMER for differential ATAC-seq peaks on standard chromosomes (Gained: n = 4023 peaks; Static/N.S.: n = 57,747 peaks; and Lost: n = 306 peaks). (b) Known transcription factor motif analysis for ATAC-seq gained peaks (n = 4023 peaks) performed by HOMER with Static/N.S. ATAC-seq peaks used as background. AP-1 family members indicated by red points and highlighted in gray. (c) Heatmap of ATAC-seq signal, H3K27Ac signal, and H3K4me1 signal, and BRG1 signal in CUT and RUN experiments +/- BRG1 reexpression at the gained ATAC-seq peaks in BIN67 cells. (d) Examples of browser tracks showing enrichment (IP/IgG) of CUT and RUN data for BRG1 and Jun in BIN67 and SCCOHT-1 cells. (e) Metaplots showing BRG1 or c-Jun signal relative to Brg1 peaks (+/- 500 bp) in BIN67 or HT1 cells comparing Control or BRG1 reexpressed cells. (f) Known transcription factor motif analysis for BRG1 peaks identified in BIN67 or SCCOHT-1 cells reexpressing BRG1 (n = 16,283, 31,379). (g) ATAC-seq signal, H3K27ac, H3K4me1 ChIP-seq signal BRG1 and c-Jun CUT and RUN signal (+/- BRG1 reexpression) at FRA1 motifs contained in open chromatin in the genome. Promoter regions defined as +/- 1 kb of TSS (n = 1423 motif sites) and all remaining sites defined as distal (n = 12,852 motif sites). Heatmaps in all panels are in the same sorted order. Source data is available in Figure 4—source data 1.
-
Figure 4—source data 1
Raw data for Figure 4.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig4-data1-v2.xlsx
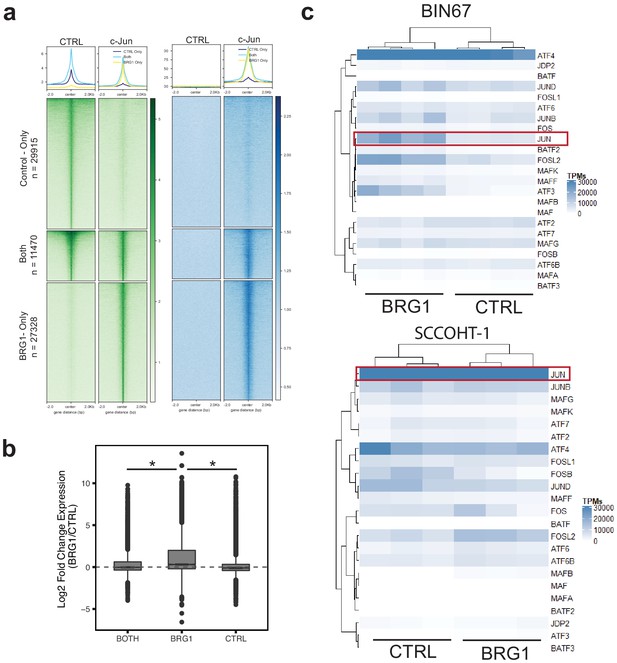
c-Jun is redistributed in SCCOHT-1 following BRG1 reexpression.
(a) CUT and RUN signal of c-Jun and BRG1 in SCCOHT-1 cells reexpressing BRG1 at peaks that are present only in cells lacking BRG1 n = 29,915, present only in cells expressing BRG1 n = 27,328, or found in both conditions n = 11470. (b) Expression of genes assigned to peaks found in each of the three previous categories. pValues are BRG1-Both. = 4.35e−63, BRG1-CTRL, 3.63e−168 by Wilcoxon signed-rank test. (c) Length scaled transcripts per million for AP-1 family members in BIN67 and SCCOHT-1 cells. Source data is available in Figure 4—figure supplement 1—source data 1.
-
Figure 4—figure supplement 1—source data 1
Raw data for Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig4-figsupp1-data1-v2.xlsx
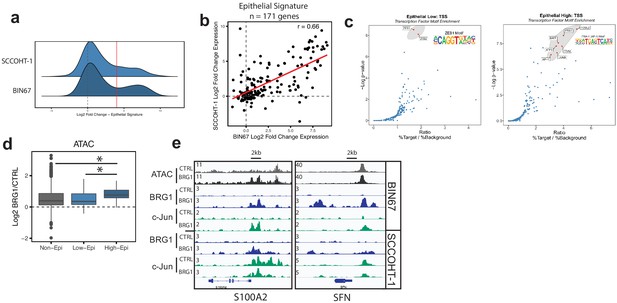
Epithelial genes dependent on AP-1 are highly upregulated by BRG1 expression.
(a) Density plot of RNA-seq Log2FoldChange (BRG1/Control) for epithelial signature genes (n = 171 genes). Red dotted line indicates cutoff point for ‘High’ category (Log2FoldChange > 4). (b) Scatterplot of log2FoldChanges (+/- BRG1) in BIN67 (x-axis) and SCCOHT-1(y-axis). Pearson’s correlation. = 0.66. (c) Known transcription factor motif analysis at the promoter of (b) low epithelial gene and (c) high epithelial genes from BIN67 xCell gene set. Default settings defined by HOMER were used for promoter region (−300 bp, +50 bp from Transcription Start Site, TSS) and background. Transcription factors that were also expressed in RNA-seq data were plotted. Transcription factors that were not expressed at a mean count across all samples greater than four TPMs were eliminated. Highest ranking based on -log(p-value) and Ratio (% Target / % Background) and of similar TF families plotted in red and encircled in gray. (d) Box plot of ATAC-seq signal (Log2FoldChange; BRG1/Control) at the BRG1 CUT and RUN peaks annotated to all high epithelial genes (n = 48 peaks), all low epithelial genes (n = 94 peaks) and non-epithelial genes (n = 15,562 peaks). Non-epithelial genes defined as a BRG1 CUT and RUN-seq peak annotated to nearest gene not contained in either high or low group. Statistical analysis was performed by Wilcoxon Rank Sum Test on low epithelial versus non-epithelial (N.S., p-value=0.907), low epithelial versus high epithelial (p-value=3.5e−4), and high epithelial versus non-epithelial (p-value=5.4e−5). (e) IGV browser tracks of two epithelial signature loci. Source data is available in Figure 5—source data 1.
-
Figure 5—source data 1
Raw data for Figure 5.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig5-data1-v2.xlsx
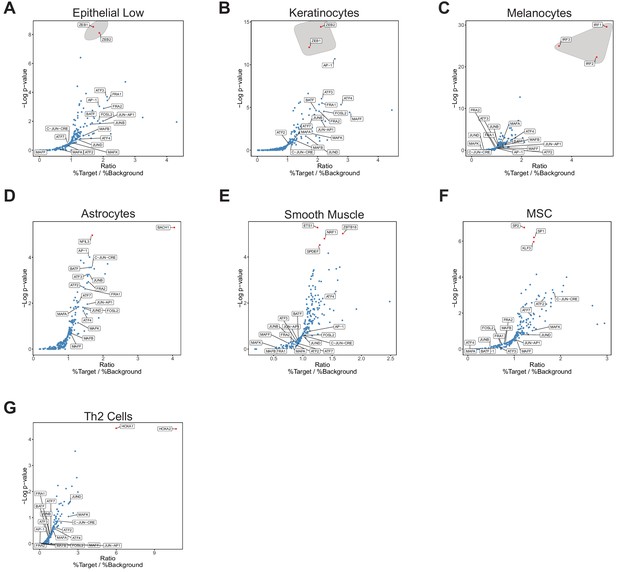
Transcription factor motif analysis at promoter of xCell gene signatures.
Known transcription factor motif analysis at the promoter of genes that compromise xCell signatures was performed by HOMER. Default settings defined by HOMER were used for promoter region (−300 bp, +50 bp from Transcription Start Site) and background. Transcription factors that were also expressed in RNA-seq data were plotted. Transcription factors that were not expressed at a mean across all samples greater than four TPMs were eliminated. Highest ranking based on -log(p-value) and Ratio (% Target / % Background) are colored red. Those of similar family are circled in gray. AP-1 family members are labeled. (A) Epithelial low group (defined in BIN67) is shown the same as Figure 5b with AP-1 family members additionally pointed out (n = 100 genes in category). (B) Keratinocytes (n = 121 genes in signature). (C) Melanocytes (n = 212 genes in signature). (D) Astrocytes (n = 219 genes in signature). (E) Smooth muscle (n = 308 genes in signature). (F) Mesenchymal stem cells (MSC; n = 105 genes in signature). (G) Th2 cells (n = 22 genes in signature).
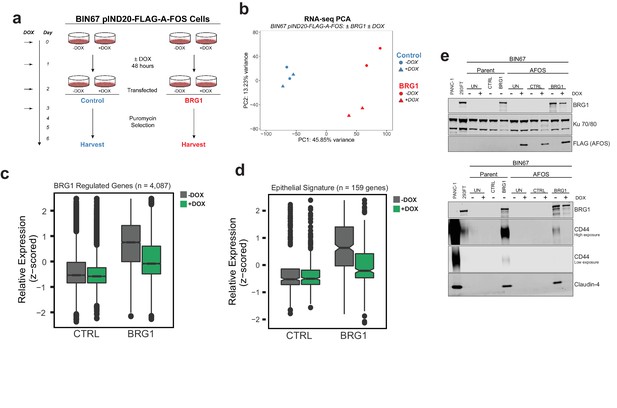
AP-1 activity is necessary for BRG1 induction of epithelial targets.
(a) Experimental design for BRG1 reexpression and A-FOS induction. BIN67 pIND20-FLAG-A-FOS cells were induced for 2 days with DOX prior to BRG1 reexpression. BRG1 reexpression followed previous reexpression studies (Figure 1a). (b) PCA analysis of BIN67 pIND20-FLAG-A-FOS (+/- DOX, +/- BRG1) RNA-seq results. (c) Boxplot of gene expression (z-scored) in BIN67 pIND20-FLAG-A-FOS (+/- DOX, +/- BRG1) for the significantly increased genes identified by BRG1 expression in parent BIN67 (Figure 1C; n = 4087), p-values ~0 between Control samples and 2.4e−6 between BRG1 samples by (Wilcoxon signed-rank test). (d) Boxplot of gene expression (z-scored) in BIN67 pIND20-FLAG-A-FOS (+/- DOX, +/- BRG1) for the genes in the epithelial signature that were measurable by RNA-seq n = 159. (p-values 0.96 between Control samples, and 4.95e−22 between BRG1 samples by Wilcoxon signed-rank test). (e) Western blots for BIN67 +/- BRG1 +/- AFOS following experimental design in (a). CD44 shown at low/high exposure by LiCOR detection. Blots are technical duplicates of the same protein sample run in parallel. Par = Parent. AFOS = BIN67 pIND20-FLAG-A-FOS cells. CTRL = control transfected. UN = untransfected. PANC-1 serves as a positive control for CD44 and Claudin-4 protein expression. Ku 70/80 serves as an internal loading control. Source data is available in Figure 6—source data 1.
-
Figure 6—source data 1
Raw data for Figure 6.
- https://cdn.elifesciences.org/articles/59073/elife-59073-fig6-data1-v2.xlsx
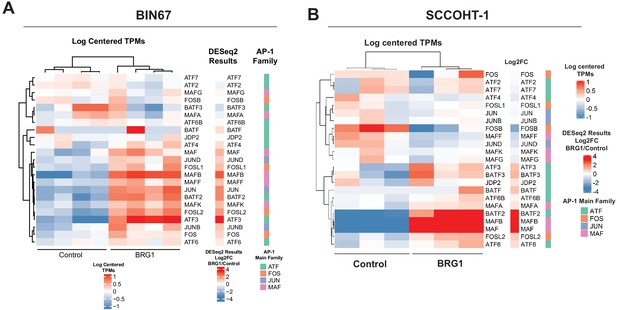
Gene expression of AP-1 family members in BIN67 and SCCOHT-1 cells.
Heatmap of RNA-seq counts of AP-1 family members for (A) BIN67 and (B) SCCOHT-1. Counts (gene and library scaled TPMs) are log scaled and mean centered. Individual replicates are shown. Heatmap of DESeq2 log2FoldChange (log2FC) results for AP-1 family member gene expression (RNA-seq). Genes are in same row order as heatmap 4A. AP-1 main family defined for each gene. Rows are ordered the same as other heatmaps.
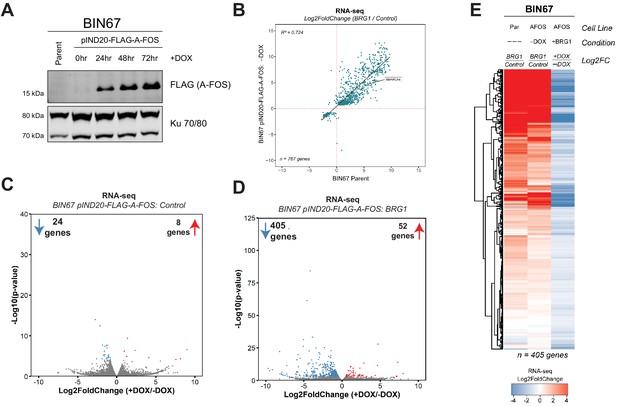
A-FOS induction in BIN67 pIND20-FLAG-A-FOS cells.
(A) BIN67 pIND20-FLAG-A-FOS cells were induced with doxycycline (DOX; [1 µg/mL]) for 0, 24, 48, and 72 hr (hr). Media and DOX was replaced daily. Protein was extracted by RIPA High-Salt buffer and 30 µg of protein was loaded for western blotting analysis. Ku 70/80 serves as an internal loading control. (B) Scatter plot of RNA-seq gene expression (Log2FoldChange = BRG1/Control) for BIN67 pIND20-FLAG-A-FOS (-DOX condition) versus BIN67 parent. Genes that were significantly changing in BIN67 pIND20-FLAG-A-FOS (-DOX condition; BRG1 versus control) were plotted (n = 767 genes). (C) Volcano plot of RNA-seq differential gene expression in BIN67 pIND2-FLAG-A-FOS, Control transfected (+DOX/-DOX). (D) Volcano plot of RNA-seq differential gene expression in BIN67 pIND20-FLAG-A-FOS, BRG1 transfected (+DOX/-DOX). (E) RNA-seq heatmap of the 405 downregulated genes (BIN67 pIND20-FLAG-A-FOS, BRG1) transfected (+DOX/-DOX; Figure 5e). Par = Parent. AFOS = pIND20 FLAG-A-FOS. Not expressed defined as NA log2FoldChange by DESeq2.
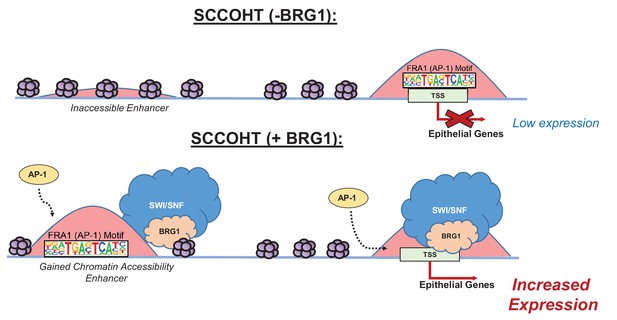
Model of BRG1 and AP-1 driving epithelial cell signature in SCCOHT tumorigenesis.
Following BRG1 reexpression in SCCOHT cells, BRG1 is recruited to and opens distal chromatin enriched in AP-1 motifs. BRG1 is also recruited to promoters of epithelial genes that have AP-1 motifs. These epithelial genes are upregulated in the presence of both BRG1 and AP-1 at the gene and protein level.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Homo sapiens) | BRG1 | Ensembl | ENSG00000127616 | SMARCA4 |
| Gene (Homo sapiens) | BRM | Ensembl | ENSG00000080503 | SMARCA2 |
| Gene (Homo sapiens) | JUN | Ensembl | ENSG00000177606 | JUN |
| Cell line (Homo sapiens) | BIN67 | Gamwell et al. (2013) doi: 10.1186/1750-1172-8-33 | SCCOHT Cell Line | Female |
| Cell line (Homo sapiens) | SCCOHT-1 | Otte et al. (2014) doi: 10.1186/s13023-014-0126-4 | SCCOHT Cell Line | Female |
| Cell line (Homo sapiens) | COV434 | Karnezis et al., 2020. doi: 10.1016/j.ygyno.2020.12.004 | SCCOHT Cell Line | Female |
| Cell Line (Homo sapiens) | A427 | ATCC HTB-53 | Non-Small Cell Lung Cancer Cell Line | Male |
| Transfected construct (Homo sapiens) | pBabe-puro | https://www.addgene.org/1764/ | RRID:Addgene_1764 | |
| Transfected construct (Homo sapiens) | pBJ5-BRG1 | https://www.addgene.org/17873/ | RRID:Addgene_17873 | |
| Transfected construct (Homo sapiens) | pCDNA3(+)-neo | Invitrogen | ||
| Lentiviral Construct (Homo sapiens) | pINDUCER20-FLAG-A-FOS | Ibrahim & Abudu et al. (2018). doi: 10.18632/oncotarget.26047 | ||
| Lentiviral Construct (Homo sapiens) | pINDUCER20-BRG1 | pINDUCER20 Backbone: https://www.addgene.org/44012/ | ||
| Antibody | Anti-Brg1 (rabbit monoclonal) | Abcam | Ab110641, RRID:AB_10861578 | CUT and RUN (1:25), WB (1:10,000) |
| Antibody | Anti-Jun (rabbit monoclonal) | Cell Signaling Technologies | 9165,RRID:AB_2130165 | CUT and RUN (1:25) |
| Antibody | Anti-Jun (rabbit ) | ThermoFisher | MA5-15172, RRID:AB_10979794 | CUT and RUN (1:25) |
| Antibody | Anti-T7 (rabbit monoclonal) | Cell Signaling Technologies | 13246,RRID:AB_2798161 | CUT and RUN (1:25) |
| Antibody | Anti-Rabbit IgG (Rabbit Monoclonal) | Cell Signaling Technologies | 3900, RRID:AB_1550038 | CUT and RUN (1:25) |
| Antibody | Anti-CD44 (Mouse Monoclonal) | Larry S. Sherman Lab and ATCC Hermes-3 | WB (1:200); IF (1:50) | |
| Antibody | Anti-Claudin-4 (Mouse Monoclonal Clone 3E2C1) | ThermoFisher (Invitrogen) | 32–9400, RRID:AB_2533096 | WB (1:1000); IHC (1:50) |
| Antibody | Anti-Ku 70/80 (Rabbit Polyclonal) | Dale Ramsden Lab - UNC Chapel Hill | WB (1:10,000) | |
| Antibody | Anti-Actin (Rabbit Monoclonal) | Sigma | A2066 | WB (1:1000) |
| Antibody | anti-FLAG (Rabbit Monoclonal) | Cell Signaling Technologies | 2368S RRID:AB_2217020 | WB (1:1000) |
| Commercial assay or kit | Fugene 6 | Promega | E269A | |
| Commercial assay or kit | Kapa Hyperprep | Roche | KK8504 | |
| Commercial assay or kit | CUT and RUN reagents | Cell Signaling Technologies | 40366 | |
| Software, algorithm | Deeptools v3.2.0 | 10.1093/nar/gkw257 | RRID:SCR_016366 | |
| Software, algorithm | Bowtie2 v 2.3.4.1 | 10.1038/nmeth.1923 | RRID:SCR_005476 | |
| Software, algorithm | Samtools v 1.9 | 10.1093/bioinformatics/btp352 | RRID:SCR_002105 | |
| Software, algorithm | Macs2 2.1.2 | 10.1186/gb-2008-9-9-r137 | RRID:SCR_013291 | |
| Software, algorithm | STAR (v 2.5.3a) | 10.1093/bioinformatics/bts635 | RRID:SCR_015899 | |
| Software, algorithm | FASTQC (v 0.11.5) | Andrews, 2010:http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ | RRID:SCR_014583 | |
| Software, algorithm | DESeq2 (v1.22.2) | 10.1186/s13059-014-0550-8 | RRID:SCR_015687 | |
| Software, algorithm | HOMER (v4.10) | 10.1016/j.molcel.2010.05.004 | RRID:SCR_010881 | |
| Software, algorithm | Picard (v2.10.3 and v 2.20) | http://broadinstitute.github.io/picard/ | RRID:SCR_006525 | |
| Software, algorithm | Bedtools (v2.26.0) | 10.1093/bioinformatics/btq033 | RRID:SCR_006646 | |
| Software, algorithm | tximport(v1.14.0) | 10.18129/B9.bioc.tximport | RRID:SCR_016752 | |
| Software, algorithm | salmon(v0.8.2) | 10.1038/nmeth.4197; https://combine-lab.github.io/salmon/ | RRID:SCR_017036 |
Additional files
-
Supplementary file 1
RNA-seq and Proteomics differential expression results for BIN67 +/- BRG1 reexpression.
A, Table of DESeq2 results for RNA-seq BIN67 +/- BRG1 samples. Log2FoldChange = BIN67/Control. B, Table of PECA analysis results for proteomics BIN67 +/- BRG1.
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp1-v2.xlsx
-
Supplementary file 2
Transcription factor motif results for ATAC-seq gained peaks.
Table of transcription factor motif analysis results for ATAC-seq gained peaks described in Figure 3e
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp2-v2.xlsx
-
Supplementary file 3
RNA-seq differential expression results for BIN67 +/- BRG1 +/- A FOS.
A, Table of DESeq2 results for BIN67 pIND20-FLAG-A-FOS, -DOX Conditions (absent A-FOS), +/- BRG1used in volcano plot Figure 6. Log2Foldchange = BRG1/Control. B, Table of DESeq2 results for BIN67 pIND20-FLAG-A-FOS, Control transfected, +/- DOX (A-FOS) used in volcano plot Figure 6. Log2Foldchange = +DOX/-DOX. B, Table of DESeq2 results for BIN67 pIND20-FLAG-A-FOS, BRG1 transfected, +/- DOX (A-FOS) used in volcano plot Figure 6. Log2Foldchange = +DOX/-DOX.
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp3-v2.xlsx
-
Supplementary file 4
RNA-seq differential expression results for SCCOHT-1 and COV434 +/- BRG1 reexpression.
A. Table of DESeq2 Results for SCCOHT-1 cells +/- BRG1. B. Table of DESeq2 results for COV434 +/- BRG1
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp4-v2.xlsx
-
Supplementary file 5
ATAC sites used in analysis of BRG1 and c-Jun localization.
A. ATAC sites gained following expression of BRG1. B. ATAC sites that overlap a Fra1 motif, used to identify protein localization relative to motif location.
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp5-v2.xlsx
-
Supplementary file 6
Peaks identified in CUT and RUN analysis.
Table of output from macs2 peak calling on each CUT-and-RUN experiment for BRG1 and c-Jun in BIN67 and SCCOHT-1 cells.
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp6-v2.tsv
-
Supplementary file 7
Transcription factor motif results for BRG1 peaks found in BIN67 and SCCOHT-1.
Motif analysis results from homer to identify known transcription factor motifs enriched at BRG1 peak locations.
- https://cdn.elifesciences.org/articles/59073/elife-59073-supp7-v2.tsv
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/59073/elife-59073-transrepform-v2.docx





