Male-predominant galanin mediates androgen-dependent aggressive chases in medaka
Figures
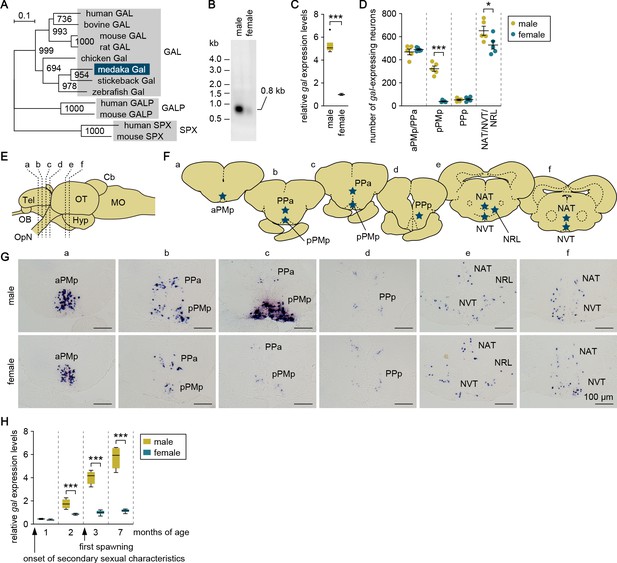
Male-biased sexual dimorphism exists in gal expression in the MPOA.
(A) Phylogenetic tree showing the relationship of medaka Gal to other known GAL family proteins. The number at each node indicates bootstrap values for 1000 replicates. Scale bar represents 0.1 substitutions per site. GALP, galanin-like peptide; SPX, spexin hormone. For species names and GenBank accession numbers, see Supplementary file 2. (B) Detection of gal transcript in the whole brain of adult males and females by Northern blot analysis. Sizes (in kb) of RNA markers and the detected band are indicated on the left and right, respectively. (C) Confirmation of male-biased expression of gal in adult whole brain by real-time PCR (n = 8 per sex). ***p<0.001 (unpaired t-test with Welch’s correction). (D) Number of gal-expressing neurons in each brain nucleus of adult males and females (n = 5 per sex). *p<0.05; ***p<0.001 (unpaired t-test). (E) Line drawing of a lateral view (anterior to the left) of the medaka brain showing the approximate levels of sections in panel G. (F) Line drawing of coronal sections showing the location of brain nuclei containing gal-expressing neurons (stars). (G) Representative micrographs showing gal-expressing neurons in each brain nucleus of adult males (upper panels) and females (lower panels). Scale bars represent 100 μm. (H) Levels of gal expression in the whole brain of males and females during growth and sexual maturation (n = 8 per sex and stage). ***p<0.001 (Bonferroni’s post hoc test). For abbreviations of brain regions and nuclei, see Supplementary file 1. See also Figure 1—figure supplement 1.

Sequence information for medaka gal.
(A) Nucleotide and deduced amino acid sequences of the medaka gal cDNA. The predicted signal peptide is underlined and the mature Gal polypeptide is boxed. Asterisk indicates stop codon. Nucleotide numbers are shown at the right of each sequence line. (B) Comparison of mature GAL polypeptide sequences from medaka and other vertebrate species. Amino acids identical in all sequences are shaded in beige. For species names and GenBank accession numbers, see Supplementary file 2.
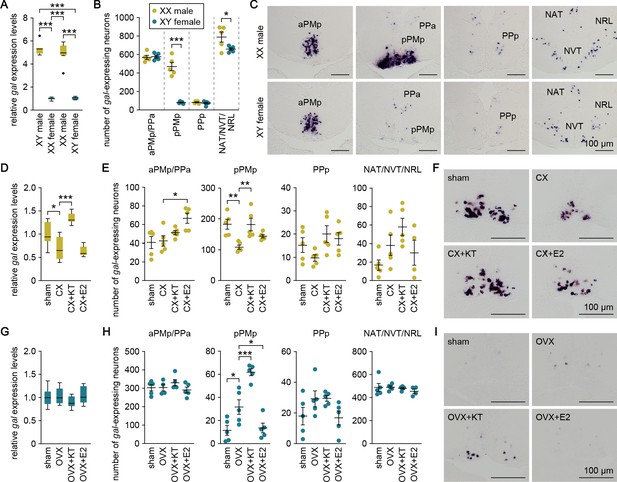
Sexually dimorphic gal expression is dependent on adult sex steroids.
(A) Levels of gal expression in the whole brain of sex-reversed adult XX males and XY females versus typical adult XY males and XX females (n = 8 per group). ***p<0.001 (Bonferroni’s post hoc test). (B) Number of gal-expressing neurons in each brain nucleus of sex-reversed adult XX male and XY females (n = 5 per group). *p<0.05; ***p<0.001 (unpaired t-test). (C) Representative micrographs showing gal-expressing neurons in each brain nucleus of sex-reversed adult XX males (upper panels) and XY females (lower panels). (D) Levels of gal expression in the whole brain of sham-operated males (sham; n = 8) and castrated males exposed to vehicle alone (CX; n = 8), KT (CX+KT; n = 9), or E2 (CX+E2; n = 6). *p<0.05; ***p<0.001 (Bonferroni’s post hoc test). (E) Number of gal-expressing neurons in each brain nucleus of sham, CX, CX+KT, and CX+E2 males (n = 5 per group). *p<0.05; **p<0.01 (Bonferroni’s post hoc test). (F) Representative micrographs showing gal expression in pPMp of sham, CX, CX+KT, and CX+E2 males. (G) Levels of gal expression in the whole brain of sham females and ovariectomized females exposed to vehicle alone (OVX), KT (OVX+KT), or E2 (OVX+E2) (n = 12 per group). (H) Number of gal-expressing neurons in each brain nucleus of sham, OVX, OVX+KT, and OVX+E2 females (n = 5 per group). *p<0.05; ***p<0.001 (Bonferroni’s post hoc test). (I) Representative micrographs showing gal expression in pPMp of sham, OVX, OVX+KT, and OVX+E2 females. Scale bars represent 100 μm. For abbreviations of brain nuclei, see Supplementary file 1. See also Figure 2—figure supplement 1.
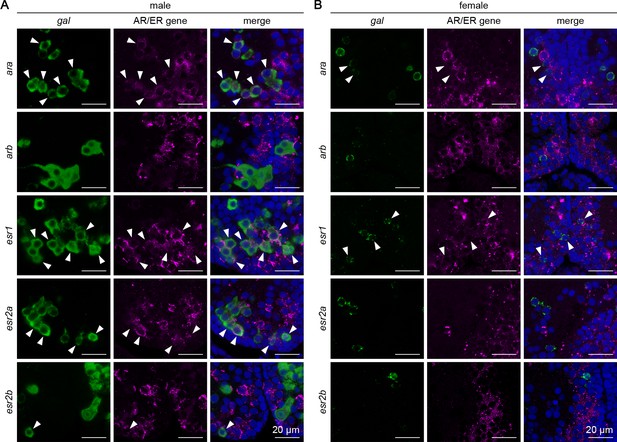
Expression of sex steroid receptors in sexually dimorphic gal neurons in pPMp.
Representative micrographs showing the expression of androgen receptor (AR) and estrogen receptor (ER) genes in pPMp, where sexually dimorphic gal-expressing neurons reside, in males (A) and females (B). In each row, left and middle panels show images of, respectively, gal (green) and ER/AR gene (magenta) expression in the same section; right panels show the merged images with nuclear counterstaining (blue). Arrowheads indicate representative neurons coexpressing gal and ER/AR. Scale bars represent 20 μm.
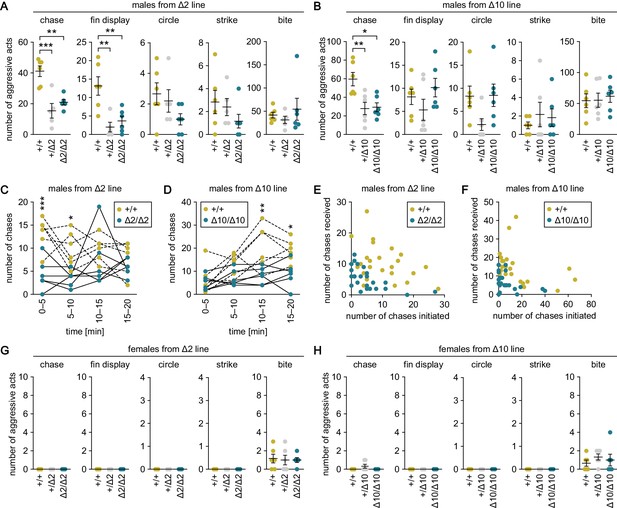
Genetic ablation of gal specifically suppresses male–male chases.
(A, B) Sum of each aggressive behavioral act (chase, fin display, circle, strike, and bite) performed by wild-type (+/+), heterozygous (+/Δ2 and +/Δ10), and homozygous (Δ2/Δ2 and Δ10/Δ10) males from Δ2 (A) and Δ10 (B) gal knockout lines. n = 6 per group, except +/Δ2 males, where n = 5. **p<0.01; ***p<0.001 (Bonferroni’s post hoc test). (C, D) Number of chases performed by wild-type and homozygous males from Δ2 (C) and Δ10 (D) gal knockout lines for each 5-min interval. n = 6 per genotype. Asterisks indicate significant differences between the two genotypes in the same time interval. *p<0.05; **p<0.01; ***p<0.001 (Bonferroni’s post hoc test). (E, F) Number of chases initiated (x-axis) and received (y-axis) by each wild-type and homozygous male from Δ2 (E) and Δ10 (F) gal knockout lines. (G, H) Sum of each aggressive behavioral act (chase, fin display, circle, strike, and bite) performed by wild-type, heterozygous, and homozygous females from Δ2 (G) and Δ10 (H) gal knockout lines. n = 6 per group. See also Figure 3—figure supplement 1 and Figure 3—figure supplement 2.
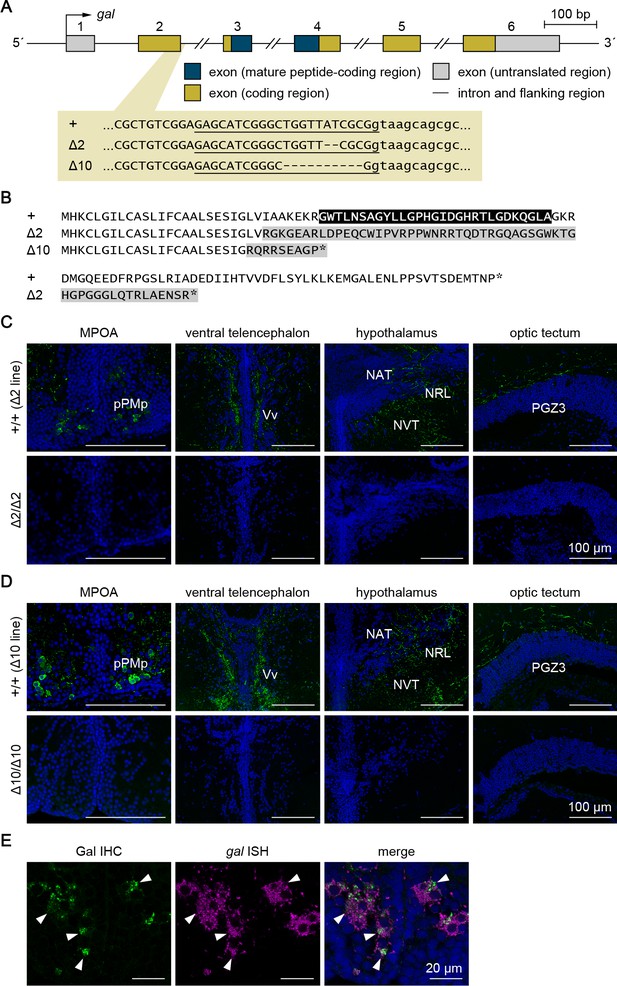
Generation and verification of gal knockout medaka.
Two different lines of gal knockout medaka (Δ2 and Δ10) were generated by CRISPR/Cas9-mediated genome editing. (A) Gene structure of gal showing the location of the CRISPR target site, which is enlarged to show the nucleotide sequences of the wild-type (+) and targeted (Δ2 and Δ10) alleles. Exon numbers are indicated above each exon. Exon and intron sequences are given in uppercase and lowercase letters, respectively. The target sequence complementary to the CRISPR RNA is underlined, and deleted nucleotides are indicated by dashes. (B) Comparison of the deduced Gal precursor protein sequences of the +, Δ2, and Δ10 alleles. The mature Gal polypeptide is indicated in white letters on a black background. The altered sequence caused by a frameshift is shaded in gray. Asterisks indicate stop codons. (C, D) Successful ablation of gal in the Δ2 (C) and Δ10 (D) knockout lines was confirmed by the observation of Gal immunoreactivity (green) in +/+ males but not in Δ2/Δ2 or Δ10/Δ10 males (n = 4 per group). Shown are representative micrographs of the MPOA containing pPMp, where the cell bodies of Gal-expressing neurons reside, and the ventral telencephalon, hypothalamus, and optic tectum, where Gal-containing axons are densely distributed. Blue color indicates nuclear counterstaining. Scale bars represent 100 μm. For abbreviations of brain nuclei, see Supplementary file 1. (E) The specificity of the anti-GAL antibody was verified by a pattern of labeling consistent with gal-expressing neurons detected by in situ hybridization (ISH). Left and middle panels show images of, respectively, immunohistochemistry (IHC) using anti-GAL antibody (green) and ISH detecting gal expression (magenta) in the same pPMp section; right panel shows the merged image with nuclear counterstaining (blue). Arrowheads indicate representative neuronal cell bodies labeled by both IHC and ISH. Scale bars represent 20 μm.
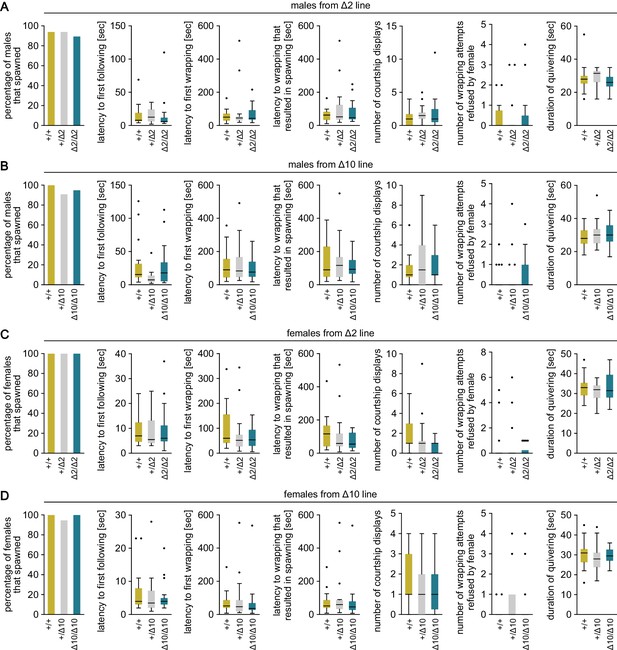
Mating behavior of gal knockout medaka.
Various parameters in the mating behavior of males (A, B) and females (C, D) from Δ2 (A, C) and Δ10 (B, D) gal knockout lines. Results were compared among wild-type (+/+), heterozygous (+/Δ2 and +/Δ10), and homozygous (Δ2/Δ2 and Δ10/Δ10) genotypes. n = 17–22 per group.
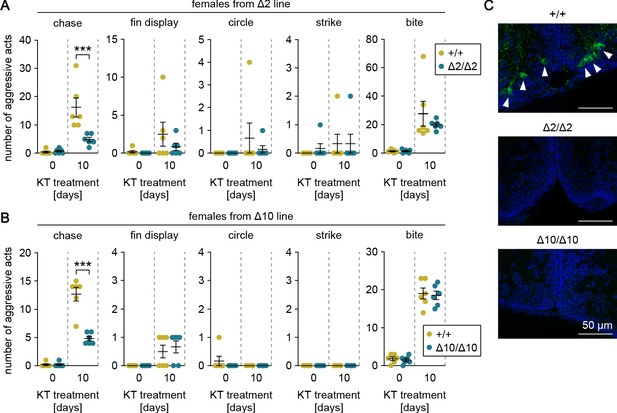
Genetic ablation of gal attenuates androgen-induced chases in females.
(A, B) Sum of each aggressive behavioral act (chase, fin display, circle, strike, and bite) performed by KT-treated females from Δ2 (A) and Δ10 (B) gal knockout lines. Asterisks indicate significant differences between wild-type (+/+) and homozygous (Δ2/Δ2 and Δ10/Δ10) genotypes on the same day. n = 6 per group. ***p<0.001 (Bonferroni’s post hoc test). (C) Representative micrographs of coronal pPMp sections from KT-treated females of Δ2 and Δ10 gal knockout lines showing the induction of Gal-expressing neurons in wild-type but not homozygous knockout females. Arrowheads indicate Gal-immunoreactive (green) neuronal cell bodies. Blue color indicates nuclear counterstaining. Scale bars represent 50 μm.
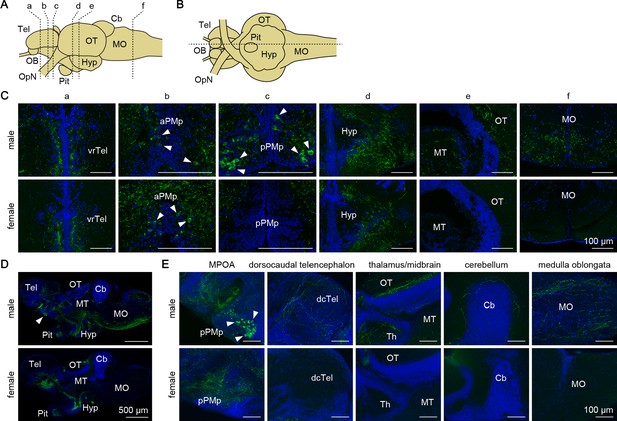
Gal peptide produced male-predominantly is transported to various brain regions.
(A, B) Line drawings of lateral (A) and ventral (B) views (anterior to the left) of the medaka brain showing the approximate levels of sections in panels C and D, respectively. (C) Representative micrographs of coronal brain sections from adult males (upper panels) and females (lower panels) showing the distribution of Gal-immunoreactive cell bodies and axons. Arrowheads indicate Gal-immunoreactive neuronal cell bodies. Scale bars represent 100 μm. (D, E) Representative low (D) and high (E) magnification micrographs of sagittal brain sections (anterior to the left) from adult males (upper panels) and females (lower panels) showing the distribution of Gal-immunoreactive cell bodies and axons. Arrowheads indicate Gal-immunoreactive neuronal cell bodies in pPMp. Scale bars represent 500 μm (D) and 100 μm (E). For abbreviations of brain regions and nuclei, see Supplementary file 1. See also Figure 5—figure supplement 1.
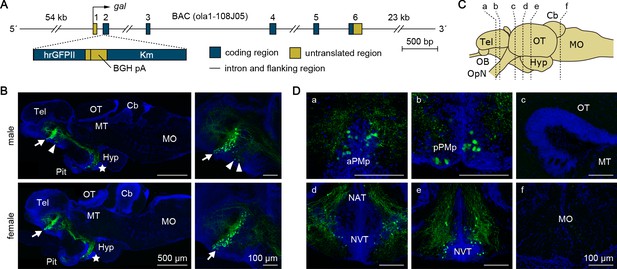
Generation and fluorescence imaging of gal-GFP transgenic medaka.
(A) Structure of the transgene in gal-GFP transgenic medaka. A 7 bp sequence containing the translation initiation site of gal in a medaka BAC clone (clone ID: ola1-108J05) was replaced by a 2136 bp DNA cassette containing the humanized Renilla reniformis GFP II (hrGFPII)-coding sequence, bovine growth hormone polyadenylation signal (BGH pA), and kanamycin resistance gene (Km). This BAC clone contains the whole transcriptional unit of gal together with 54 kb of 5′-flanking and 23 kb of 3′-flanking sequences. (B) Representative micrographs of sagittal brain sections (anterior to the left) from adult male (upper panels) and female (lower panels) gal-GFP transgenic medaka showing the distribution of GFP expression. Left panels show low-magnification images of the whole brain; right panels show high-magnification images of the MPOA. Arrows and arrowheads indicate GFP-labeled neuronal cell bodies in aPMp/PPa and pPMp, respectively; stars indicate those in NAT/NVT/NRL. Scale bars represent 500 μm (left panels) and 100 μm (right panels). (C) Line drawing of a lateral view (anterior to the left) of the medaka brain showing the approximate levels of sections in panel D. (D) Representative micrographs of coronal brain sections from gal-GFP transgenic medaka showing the distribution of GFP expression. Only images of males are presented because there were no obvious sex differences in the distribution of GFP expression. Scale bars represent 100 μm. For abbreviations of brain regions and nuclei, see Supplementary file 1.
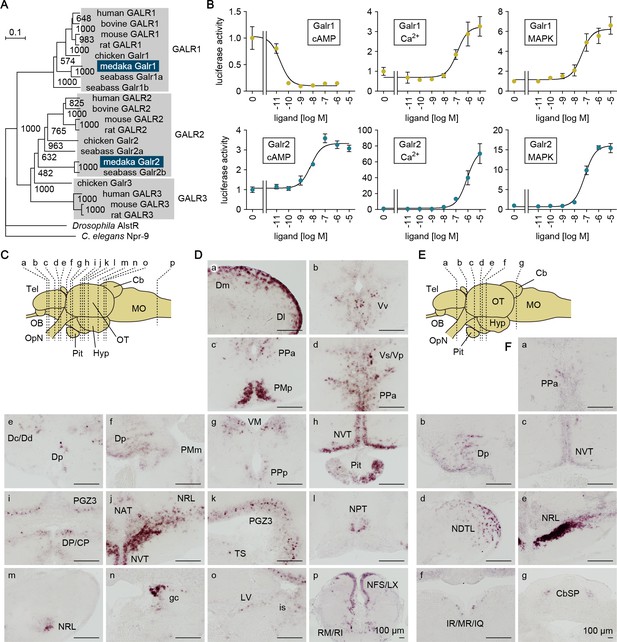
Gal receptors coupled to different signaling pathways are expressed widely in the brain.
(A) Phylogenetic tree showing the relationship of medaka Galr1 and Galr2 to other known GAL receptors. The number at each node indicates bootstrap values for 1000 replicates. Scale bar represents 0.1 substitution per site. AlstR, allatostatin receptor. For species names and GenBank accession numbers, see Supplementary file 2. (B) Intracellular signaling pathways initiated by the activation of medaka Galr1 (upper graphs) and Galr2 (lower graphs). Cells transfected with Galr1 or Galr2 were stimulated with increasing concentrations of Gal and assayed for luciferase activity indicative of intracellular cAMP levels (left panels), Ca2+ levels (middle panels), and MAPK (right panels) (n = 3). x-axis shows the concentration of Gal; y-axis shows the fold change in luciferase activity relative to the basal level, which was measured in the absence of Gal. (C) Line drawing of a lateral view (anterior to the left) of the medaka brain showing the approximate levels of sections in panel D. (D) Distribution of galr1 expression in the brain and pituitary. (E) Line drawing of a lateral view (anterior to the left) of the medaka brain showing the approximate levels of sections in panel F. (F) Distribution of galr2 expression in the brain and pituitary. All images are coronal sections. Only images of males are shown because there were no obvious sex differences in the distribution of expression (n = 5 per sex). Scale bars represent 100 μm. For abbreviations of brain regions and nuclei, see Supplementary file 1. See also Figure 6—figure supplement 1.
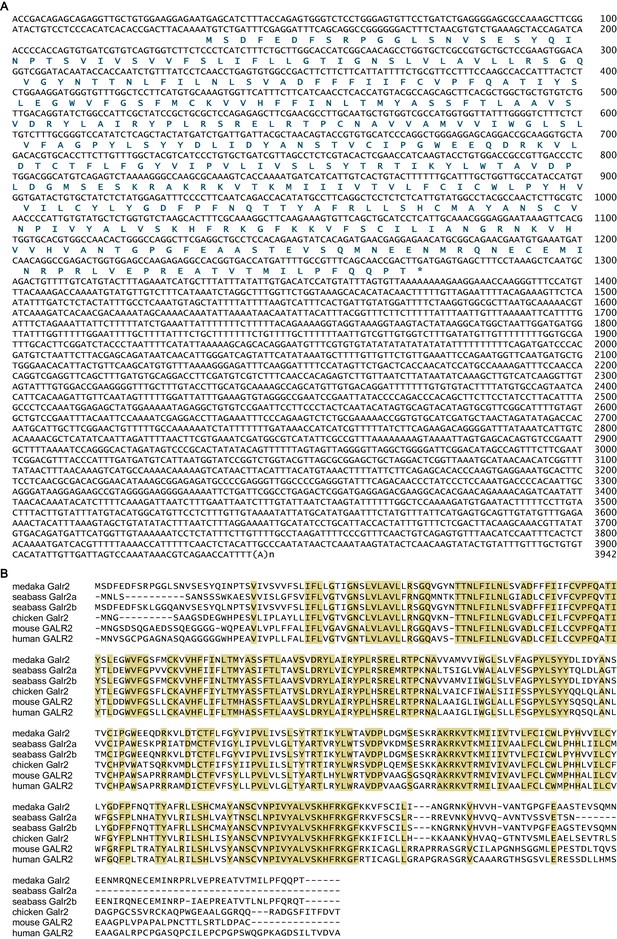
Sequence information for medaka galr2.
(A) Nucleotide and deduced amino acid sequences of the medaka galr2 cDNA. Asterisk indicates stop codon. Nucleotide numbers are shown at the right of each sequence line. (B) Sequence comparison of medaka Galr2 and its orthologs in other vertebrate species. Amino acids identical in all sequences are shaded in beige. For species names and GenBank accession numbers, see Supplementary file 2. Supplementary Data List.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Oryzias latipes) | gal | this paper | GenBank:LC532140 | |
| Gene (O. latipes) | galr2 | this paper | GenBank:LC532141 | |
| Gene (O. latipes) | actb | GenBank | GenBank:NM_001104808 | |
| Strain, strain background (O. latipes) | d-rR | NBRP Medaka | strain ID:MT837 | maintained in a closed colony over 10 years in Okubo lab |
| Genetic reagent (O. latipes) | gal knockout Δ2 line | this paper | N/A | generated and maintained in Okubo lab |
| Genetic reagent (O. latipes) | gal knockout Δ10 line | this paper | N/A | generated and maintained in Okubo lab |
| Genetic reagent (O. latipes) | gal-GFP transgenic | this paper | N/A | generated and maintained in Okubo lab |
| Cell line (Homo sapiens) | HEK293T | Riken BRC Cell Bank | cell number:RCB2202; RRID:CVCL_0063 | |
| Cell line (Escherichia coli) | DY380 | DOI:10.1038/35093556; DOI:10.1006/geno.2000.6451 | N/A | |
| Transfected construct (H. sapiens) | pcDNA3.1/V5-His-TOPO | Thermo Fisher Scientific | cat#:K480001 | |
| Transfected construct (H. sapiens) | pGL4.29 | Promega | cat#:E8471 | |
| Transfected construct (H. sapiens) | pGL4.33 | Promega | cat#:E1340 | |
| Transfected construct (H. sapiens) | pGL4.74 | Promega | cat#:E6921 | |
| Antibody | alkaline phosphatase-conjugated anti-DIG antibody (sheep polyclonal) | Roche Diagnostics | cat#:11093274910; RRID:AB_514497 | (1:500 or 1:2000) |
| Antibody | horseradish peroxidase-conjugated anti-fluorescein antibody (sheep polyclonal) | PerkinElmer | cat#:NEF710001EA; RRID:AB_2737388 | (1:1000) |
| Antibody | anti-GAL antibody (rabbit polyclonal) | Enzo Life Sciences | cat#:BML-GA1161-0025; RRID:AB_2051473 | (1:200 or 1:500) |
| Antibody | Alexa Fluor 555-conjugated goat anti-rabbit IgG (goat polyclonal) | Thermo Fisher Scientific | cat#:A-21428; RRID:AB_2535849 | (1:1000) |
| Antibody | Alexa Fluor 488-conjugated goat anti-rabbit IgG (goat polyclonal) | Thermo Fisher Scientific | cat#:A-11070; RRID:AB_2534114 | (1:1000) |
| Recombinant DNA reagent | full-length cDNA clone for medaka gal | this paper | clone ID:56_B03 | |
| Recombinant DNA reagent | full-length cDNA clone for medaka galr2 | this paper | clone ID:39_L19 | |
| Recombinant DNA reagent | pGEM-Teasy vector | Promega | cat#:A1360 | |
| Recombinant DNA reagent | pCS2+hSpCas9 plasmid | Addgene | RRID:Addgene_51815 | |
| Recombinant DNA reagent | medaka bacterial artificial chromosome (BAC) clone containing the gal locus | NBRP Medaka | clone ID:108_J05 | |
| Recombinant DNA reagent | phrGFP II-1 mammalian expression vector | Agilent Technologies | cat#:240143 | |
| Sequence-based reagent | CRISPR RNA (crRNA) for medaka gal | Fasmac | N/A | GAGCATCGGGCTGGTTATCGCGG |
| Sequence-based reagent | trans-activating CRISPR RNA (tracrRNA) | Fasmac | cat#:GE-002 | |
| Peptide, recombinant protein | medaka Gal peptide | Scrum | this paper | GWTLNSAGYLLGPHGIDGHRTLGDKQGLA-NH2 |
| Commercial assay or kit | Isogen Poly(A)+Isolation Pack | Fujifilm Wako Pure Chemical Corporation | cat#:314–05651 | |
| Commercial assay or kit | nylon membrane | Roche Diagnostics | cat#:11209272001 | |
| Commercial assay or kit | DIG RNA Labeling Mix | Roche Diagnostics | cat#:11277073910 | |
| Commercial assay or kit | T7 RNA polymerase | Roche Diagnostics | cat#:10881775001 | |
| Commercial assay or kit | DIG Easy Hyb | Roche Diagnostics | cat#:11603558001 | |
| Commercial assay or kit | CDP-Star | Roche Diagnostics | cat#:12041677001 | |
| Commercial assay or kit | RNeasy Lipid Tissue Mini Kit | Qiagen | cat#:74804 | |
| Commercial assay or kit | RNeasy Plus Universal Mini Kit | Qiagen | cat#:73404 | |
| Commercial assay or kit | Omniscript RT Kit | Qiagen | cat#:205111 | |
| Commercial assay or kit | SuperScript VILO cDNA Synthesis Kit | Thermo Fisher Scientific | cat#:11754050 | |
| Commercial assay or kit | Power SYBR Green PCR Master Mix | Thermo Fisher Scientific | cat#:4367659 | |
| Commercial assay or kit | LightCycler 480 SYBR Green I Master | Roche Diagnostics | cat#:04887352001 | |
| Commercial assay or kit | TSA Plus Fluorescein System | PerkinElmer | cat#:NEL741001KT | |
| Commercial assay or kit | mMessage mMachine SP6 Kit | Thermo Fisher Scientific | cat#:AM1340 | |
| Commercial assay or kit | Dual-Luciferase Reporter Assay System | Promega | cat#:E1910 | |
| Chemical compound, drug | methyltestosterone | Fujifilm Wako Pure Chemical Corporation | cat#:136–09931 | |
| Chemical compound, drug | estradiol-17β (E2) | Fujifilm Wako Pure Chemical Corporation | cat#:058–04043 | |
| Chemical compound, drug | 11-ketotestosterone (KT) | Cosmo Bio | cat#:117 ST | |
| Chemical compound, drug | tricaine methane sulfonate | Sigma-Aldrich | cat#:E10521 | |
| Chemical compound, drug | 5-bromo-4-chloro-3-indolyl phosphate | Roche Diagnostics | cat#:11383221001 | |
| Chemical compound, drug | nitro blue tetrazolium | Roche Diagnostics | cat#:11383213001 | |
| Chemical compound, drug | agarose, type IX-A | Sigma-Aldrich | cat#:A2576 | |
| Chemical compound, drug | Fast Red | Roche Diagnostics | cat#:11496549001 | |
| Chemical compound, drug | 4′,6-diamidino-2-phenylindole (DAPI) | Thermo Fisher Scientific | cat#:D1306 | |
| Chemical compound, drug | blocking reagent | Roche Diagnostics | cat#:11096176001 | |
| Chemical compound, drug | Lipofectamine LTX | Thermo Fisher Scientific | cat#:15338100 | |
| Software, algorithm | InterPro | https://www.ebi.ac.uk/interpro/ | RRID:SCR_006695 | |
| Software, algorithm | SignalP | http://www.cbs.dtu.dk/services/SignalP/ | RRID:SCR_015644 | |
| Software, algorithm | ClustalW | http://clustalw.ddbj.nig.ac.jp/index.php | RRID:SCR_017277 | |
| Software, algorithm | GraphPad Prism | GraphPad Software | RRID:SCR_002798 |
Additional files
-
Supplementary file 1
Abbreviations of medaka brain regions and nuclei.
- https://cdn.elifesciences.org/articles/59470/elife-59470-supp1-v1.docx
-
Supplementary file 2
Species names and GenBank accession numbers of the protein sequences used in this study.
- https://cdn.elifesciences.org/articles/59470/elife-59470-supp2-v1.docx
-
Supplementary file 3
Primers used in this study.
- https://cdn.elifesciences.org/articles/59470/elife-59470-supp3-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/59470/elife-59470-transrepform-v1.docx