Discovery of a molecular glue promoting CDK12-DDB1 interaction to trigger cyclin K degradation
Figures
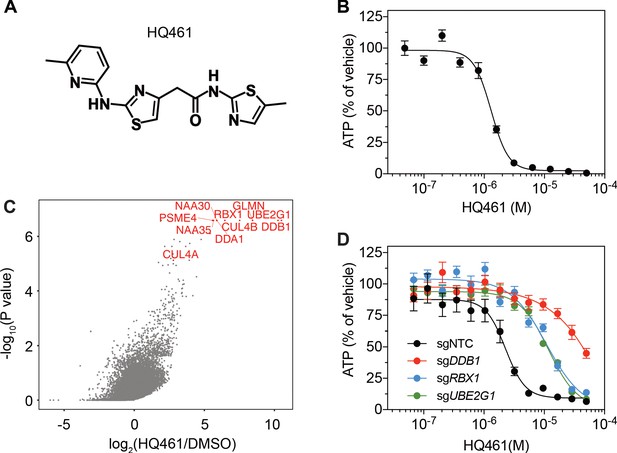
DDB1-CUL4-RBX1 mediates HQ461’s cytotoxicity.
(A) Chemical structure of HQ461. (B) Measurement of the HQ461 IC50 on the viability of A549 cells (IC50 = 1.3 µM, 95% confidence interval (CI): 1.0 µM-1.6 µM). Error bars represent standard errors of mean (SEM) from three biological replicates. (C) MAGeCK analysis of pooled genome-wide CRISPR-Cas9 sgRNA screening of HQ461 resistance in A549 cells. (D) Measurement of the HQ461 IC50 on the viability of A549 cells expressing non-targeting control (NTC, IC50 = 2.3 µM, 95% CI: 1.9 µM-2.8 µM) or sgRNAs targeting DDB1 (IC50 >28.8 µM), RBX1 (IC50 = 10.9 µM, 95% CI: 6.5 µM-101 µM), or UBE2G1 (IC50 = 11.9 µM, 95% CI: 9.8 µM-15.5 µM). Error bars represent SEM from three biological replicates.
-
Figure 1—source data 1
Measurement of the HQ461 IC50 on the viability of A549 cells (Source data for Figure 1B).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig1-data1-v3.xlsx
-
Figure 1—source data 2
MAGeCK analysis of pooled genome-wide CRISPR-Cas9 sgRNA screening of HQ461 resistance in A549 cells (Source data for Figure 1C).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig1-data2-v3.xlsx
-
Figure 1—source data 3
Measurement of the HQ461 IC50 on the viability of A549 cells expressing non-targeting control or sgRNAs targeting DDB1, RBX1, or UBE2G1 (Source data for Figure 1D).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig1-data3-v3.xlsx
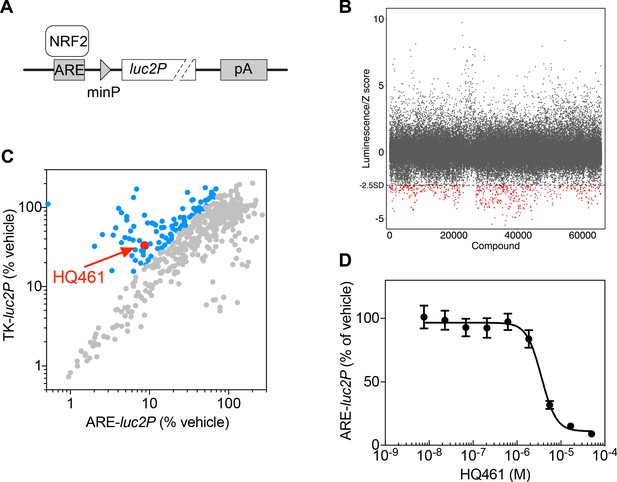
High-throughput chemical screening of NRF2 inhibitors.
(A) ARE-luc2P reporter used for high-throughput screening. NRF2 binds to anti-oxidant response elements (ARE) to drive luciferase (luc2P) expression. (B) Primary screen results. The ARE-luc2P reporter cell line was treated for 12 hr with library compounds at 10 µM. Each screening plate was individually Z-score normalized, and all the Z-scores were aggregated. The dotted line represents the cutoff (Z-score <−2.5). Red dots mark hits from the primary screen. (C) Counter screen results. Hits from the primary screen were tested in both ARE-luc2P and TK-luc2P reporter cell lines at 10 µM for 12 hr. Blue dots represent compounds selectively inhibiting ARE-luc2P relative to TK-luc2P. (D) Measurement of the HQ461 IC50 on ARE-luc2P activity (IC50 = 3.6 µM, 95% CI: 2.9 µM-4.5 µM). Error bars represent standard errors of the mean (SEM) from three biological replicates.
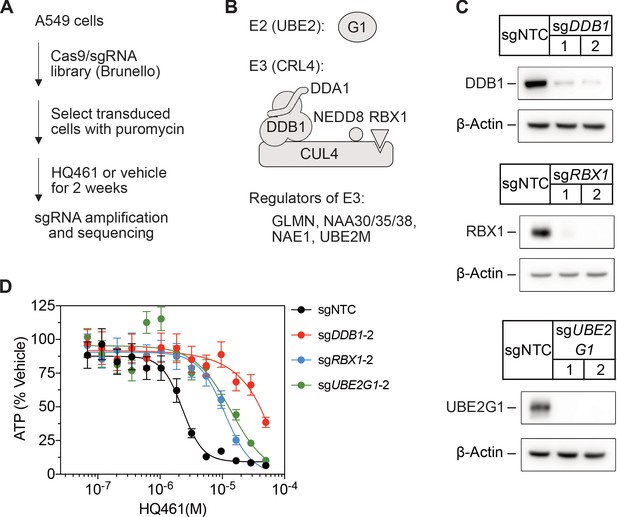
CRISPR-Cas9 screening for HQ461 resistance.
(A) Genome-wide CRISPR-Cas9 screening procedures. (B) Classification of top-ranking genes into functional categories in the ubiquitin-proteasome system. (C) Verification of successful depletion of DDB1, RBX1, and UBE2G1 by individual sgRNAs. (D) Measurement of the HQ461 IC50 on the viability of A549 cells expressing sgRNAs targeting DDB1 (IC50 >28.8 µM), RBX1 (IC50 = 11.0 µM, 95% CI: 9.1 µM-14.2 µM), or UBE2G1 (IC50 = 13.6 µM, 95% CI: 5.0 µM-50 µM) that are independent of the sgRNAs used for the experiments presented in Figure 1D. Error bars represent SEM from three biological replicates.
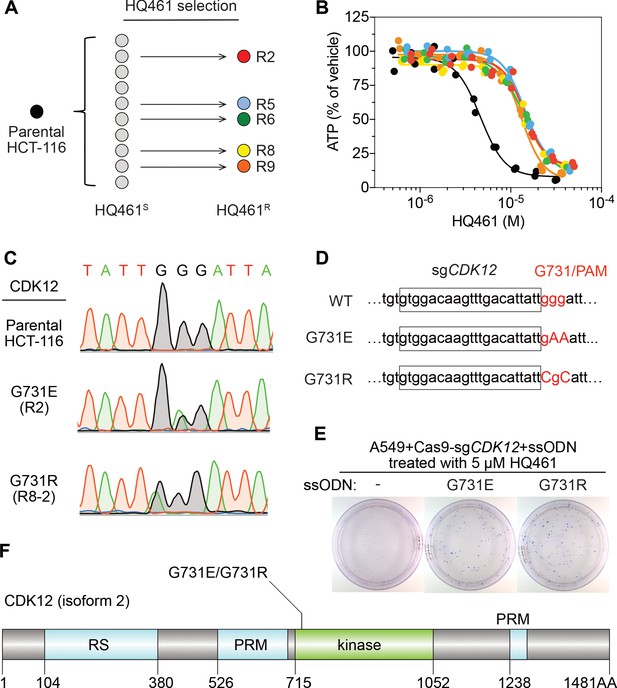
Mutations in CDK12 confer resistance to HQ461.
(A) Strategy for isolation of independent HQ461-resistant HCT-116 clones. (B) Measurement of the HQ461 IC50 on the viability of parental HCT-116 (IC50 = 4.6 µM, 95% CI: 4.1 µM-5.3 µM) and five HQ461-resistant HCT-116 clones (color coded in (A), IC50 ranging from 13.0 µM to 14.3 µM). Each point represents the average of two biological replicates. (C) Sanger sequencing verification of CDK12 mutations found in HQ461-resistant HCT-116 clones. (D) The edited genomic sequence of CDK12, highlighting the PAM motif, the target sequence (boxed), and G731E or G731R mutations. (E) CDK12 G731E or G731R knock-in in A549 cells gives rise to HQ461 resistance, visualized by crystal violet staining. Mock transfection (omitting the repair template) results in no HQ461 resistance. (F) The domain structure of CDK12 (isoform two is shown here and used in this study). G731 is located within the central kinase domain of CDK12.
-
Figure 2—source data 1
Measurement of the HQ461 IC50 on the viability of parental HCT-116 and five HQ461-resistant HCT-116 clones (source data for Figure 2B).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig2-data1-v3.xlsx
-
Figure 2—source data 2
Exome-sequencing of HQ461S versus HQ461R HCT-116 (source data for Figure 2C).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig2-data2-v3.xlsx
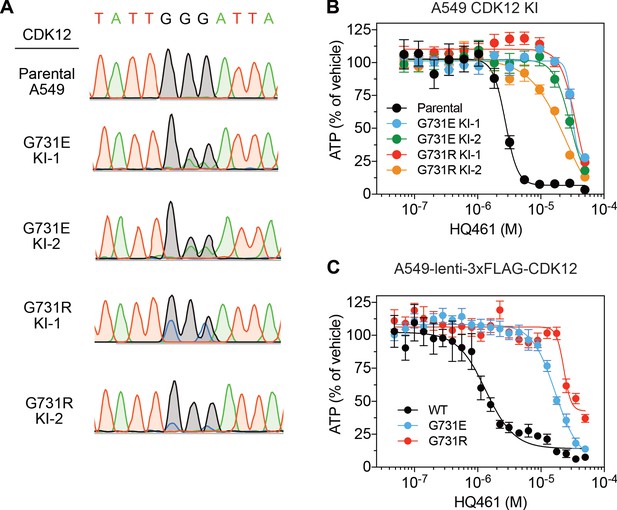
The CDK12 G731E and G731R mutations confer HQ461 resistance.
(A) Sanger sequencing verification of GGG to GAA and GGG to CGC mutations introduced by CRISPR-Cas9-mediated homology-dependent repair in A549 cells. (B) Measurement of the HQ461 IC50 on the viability of CDK12 G731E or G731R knock-in A549 cells (IC50 ranging from 25.2 µM to 36.5 µM). Error bars represent SEM from three biological replicates. (C) Measurement of the HQ461 IC50 on the viability of A549 cells stably expressing wild-type (IC50 = 1.3 µM, 95% CI: 1.1 µM-1.7 µM) or G731E/R mutant CDK12 (G731E IC50 = 16.9 µM, 95% CI: 14.2 µM-20.2 µM; G731R IC50 = 22.9 µM, 95% CI: 19.7 µM-26.8 µM). Error bars represent SEM from three biological replicates.
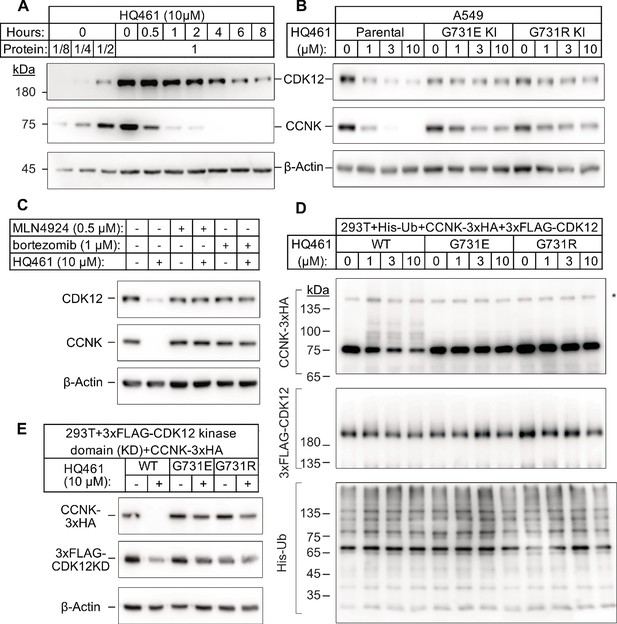
HQ461 promotes CCNK polyubiquitination and degradation through DDB1-CUL4-RBX1.
(A) Western blotting of CDK12 and CCNK from A549 cells treated with HQ461. Dilutions of untreated sample (0 hr) were included to allow quantitative assessment of the reduction of protein levels. (B) Effect of CDK12 G731E and G731R mutations on CCNK and CDK12 protein levels in A549 cells treated with HQ461. (C) Effect of bortezomib and MLN4924 on CCNK and CDK12 protein levels in A549 cells treated with HQ461. (D) HQ461 triggers in vivo polyubiquitination of CCNK in complex with wild-type CDK12 but not CDK12 G731E or G731R. The asterisk indicates a non-specific band. (E) Wild-type CDK12 kinase domain but not G731E or G731R mutant is sufficient for mediating HQ461-dependent destabilization of CCNK.
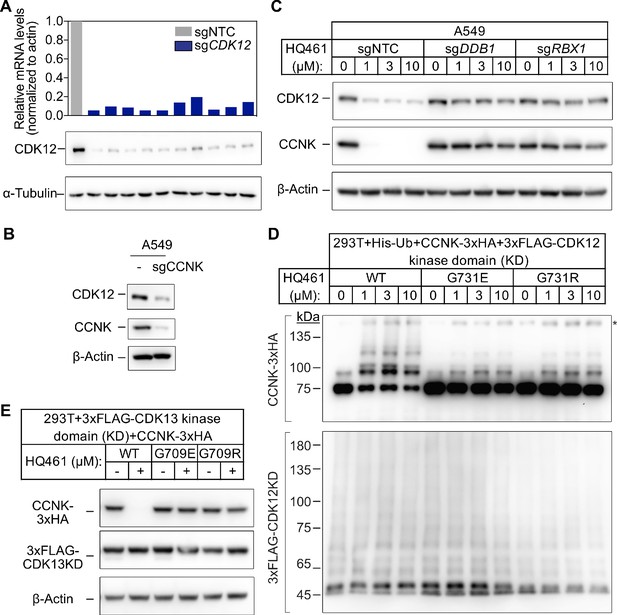
Both CDK12 and CDK13 support HQ461-induced CCNK degradation.
(A) Measurements of CDK12 mRNA and protein levels in A549 cells following CDK12 CRISPRi. (B) Effect of CCNK depletion on CDK12 protein level in A549 cells. (C) Effect of DDB1 and RBX1 depletion on CCNK and CDK12 protein levels in A549 cells treated with HQ461. (D) HQ461 triggers in vivo polyubiquitination of CCNK in complex with the wild-type CDK12 kinase domain but not with G731E or G731R mutant kinase domains. The asterisk indicates a non-specific band. (E) Wild-type CDK13 kinase domain but not G709E or G709R mutant kinase domain is sufficient for mediating HQ461-dependent degradation of CCNK.
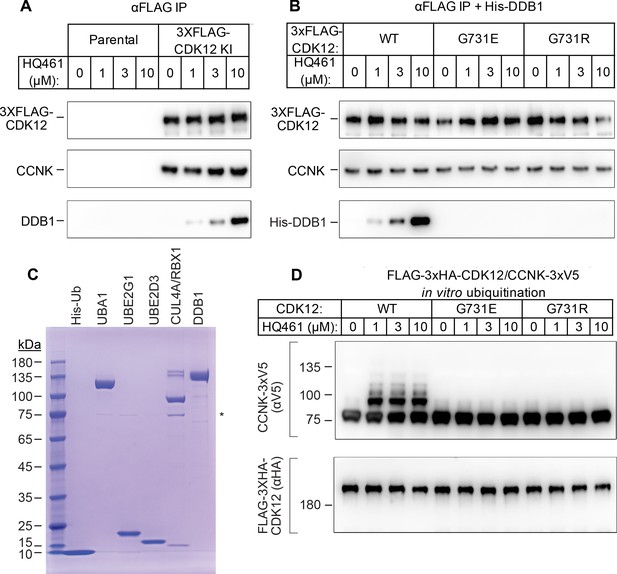
HQ461 functions as a molecular glue between CDK12 and DDB1 to promote CCNK polyubiquitination.
(A) Coimmunoprecipitation of DDB1 with 3xFLAG-CDK12/CCNK from A549 lysates treated with HQ461. (B) Coimmunoprecipitation of His-DDB1 with 3xFLAG-CDK12/CCNK WT, G731E, or G731R from A549 lysates treated with HQ461. (C) Coomassie blue staining of recombinant proteins used for an in vitro ubiquitination assay. The asterisk indicates contaminating HSP70. (D) In vitro ubiquitination of CDK12/CCNK with recombinant ubiquitin, E1 (UBA1), E2 (UBE2G1 and UBE2D3), and E3 (DDB1-CUL4-RBX1).
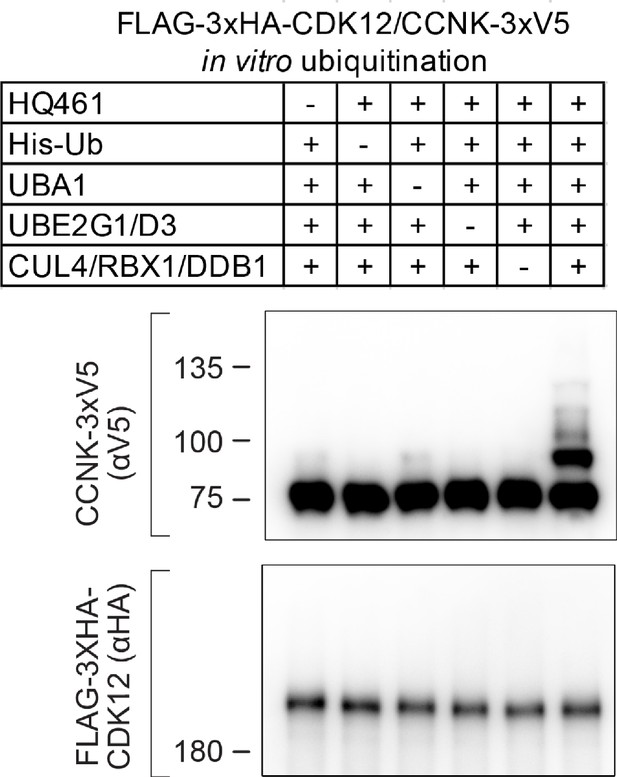
In vitro polyubiquitination of CDK12/CCNK.
The essentiality of each component for the in vitro polyubiquitination of CDK12/CCNK is demonstrated with the ‘all-minus-one’ experiment.
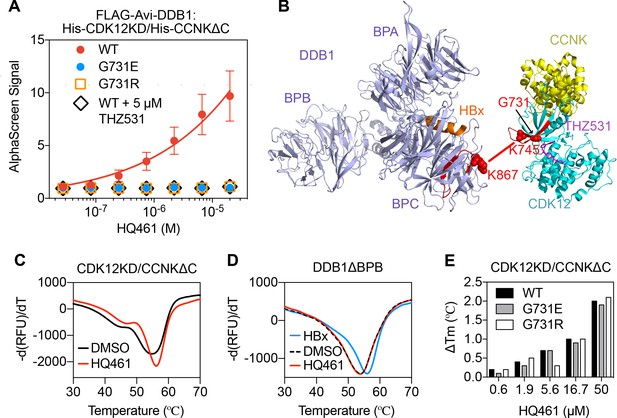
HQ461 binds to CDK12 to recruit DDB1.
(A) Detection of HQ461-dependent interaction between FLAG-Avi-DDB1 and His-CDK12KD/His-CCNKΔC examined in an AlphaScreen assay. Error bars represent SEM from three technical replicates. (B) CXMS analysis identified one inter-protein cross-link between the DDB1 (PDB code: 3i7h) lysine 867 and CDK12KD/CCNKΔC (PDB code: 5acb) lysine 745. (C) DSF analysis of CDK12KD/CCNKΔC thermal unfolding in the presence of 50 µM HQ461 or vehicle DMSO. (D) DSF analysis of DDB1ΔBPB thermal unfolding in the presence of DMSO, 50 µM HQ461 or HBx peptide. (E) Nano DSF analysis of CDK12KD/CCNKΔC (WT or G731E/R) thermal unfolding in the presence of increasing concentrations of HQ461. Δ Tm is calculated by subtracting the Tm of DMSO control.
-
Figure 5—source data 1
Detection of HQ461-dependent interaction between FLAG-Avi-DDB1 and His-CDK12KD/His-CCNKΔC examined in an AlphaScreen assay (source data for Figure 5A).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig5-data1-v3.xlsx
-
Figure 5—source data 2
CXMS analysis of DDB1 and CDK12KD/CCNKΔC in the presence of HQ461 (source data for Figure 5B).
- https://cdn.elifesciences.org/articles/59994/elife-59994-fig5-data2-v3.xlsx
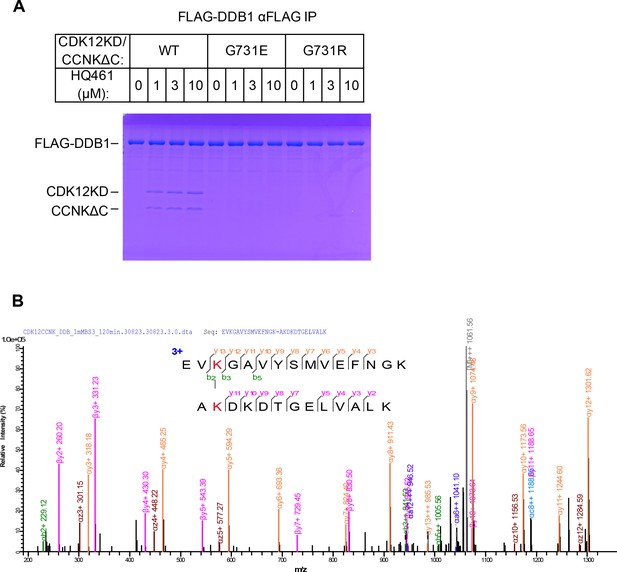
Biochemical characterization of HQ461’s molecular-glue activity.
(A) HQ461-dependent formation of a FLAG-DDB1/CDK12KD/CCNKΔC complex visualized by SDS-PAGE followed by coomassie blue staining. (B) Annotated MS/MS spectrum of the linked peptide pair from CDK12 and DDB1.
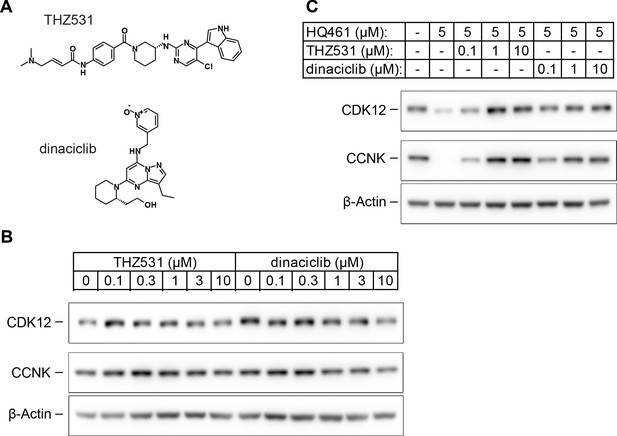
HQ461 binds to the ATP-binding pocket of CDK12.
(A) Chemical structures of THZ531 and dinaciclib. (B) Effect of THZ531 and dinaciclib on CCNK and CDK12 protein levels in A549 cells. (C) Effect of THZ531 and dinaciclib on CCNK and CDK12 protein levels in A549 cells treated with HQ461.
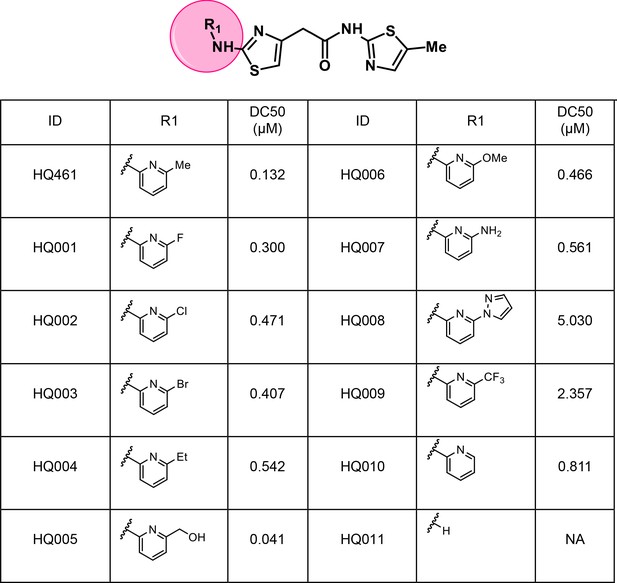
Structure-activity relationship of HQ461 analogs.
Half maximal CCNK-degradation concentrations are reported for each analog. CCNK degradation was measured using a CCNK-luc reporter.
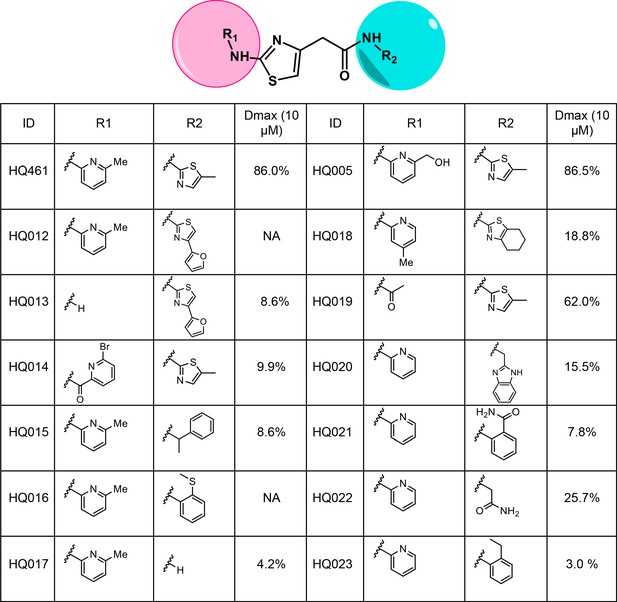
Structure-activity relationship of HQ461 analogs.
Dmax represents the percentage of CCNK degradation with 10 µM of test compound treatment relative to DMSO control. CCNK degradation was measured using a CCNK-luc reporter.
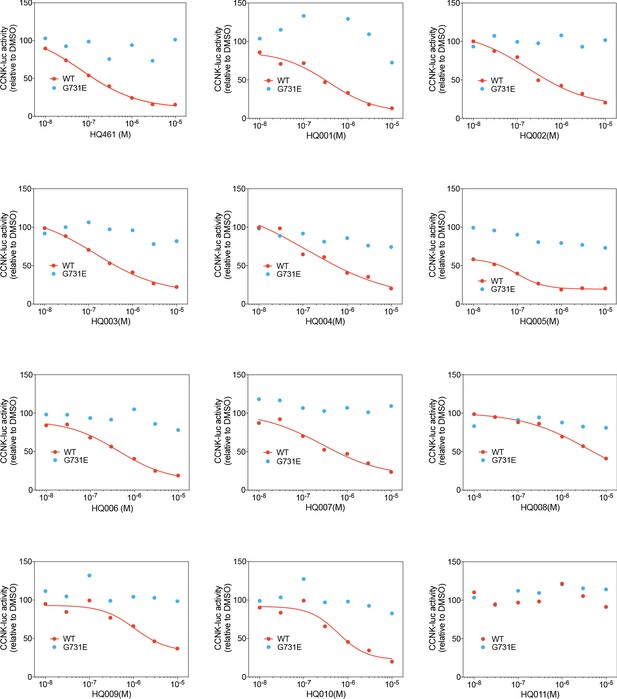
Dose response curves of HQ461 analogs in reducing CCNK-luc activity.
The CCNK-luc reporter co-transfected with CDK12KD with the G731E mutation verifies that CCNK-luc degradation is on-target.
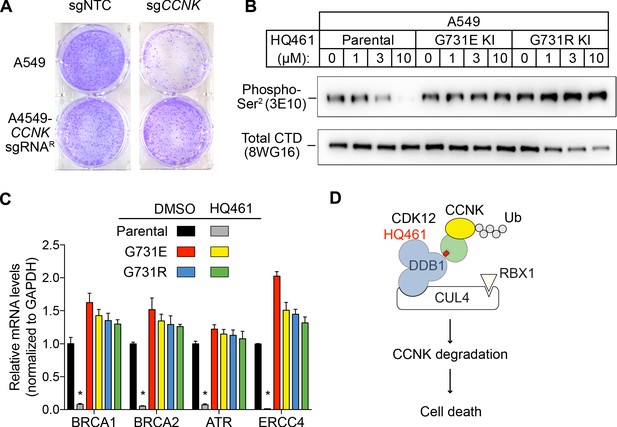
HQ461-mediated CCNK degradation causes cell death and impairs CDK12 function.
(A) Depletion of CCNK by CRISPR reduces colony formation in parental A549 cells but not in A549 cells expressing an sgRNA-resistant CCNK cDNA. (B) Western blotting to examine levels of POLII CTD Serine 2 phosphorylation and total POLII CTD in parental A549 cells or A549 CDK12 G731E or G731R knock-in cells following HQ461 treatment. (C) RT-qPCR to examine levels of BRCA1, BRCA2, ATR, and ERCC4 mRNAs in parental A549 cells or A549 CDK12 G731E or G731R knock-in cells following 10 µM HQ461 treatment for 16 hr. Error bars represent standard deviations (SD) from three technical replicates. * represents p<0.05 (T-test, two tail, paired). (D) A model of HQ461’s mechanism of action.
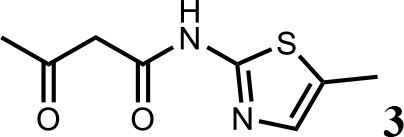
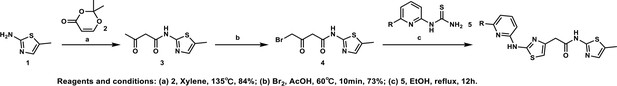
Synthesis of N-(5-methylthiazol-2-yl)−3-oxobutanamide (3).
To a solution of 5-methylthiazol-2-amine (1) (0.5 g; 4.39 mmol) in xylene (10 ml) was added 2, 2-dimethyl-4H-1, 3-dioxin-4-one (2) (685 mg; 4.83 mmol). The mixture was stirred for 12 hr at 135°C, then cooled to room temperature. The solution was filtered and washed with petroleum ether, after filtration, the solid was used directly without purification (3) (0.73 g; 3.69 mmol, 84%). 1H NMR (400 MHz, DMSO-d6) δ 11.95 (s, 1H), 7.13 (s, 1H), 3.66 (s, 2H), 2.34 (s, 3H), 2.19 (s, 3H); 13C NMR (101 MHz, DMSO-d6) δ 202.68, 165.26, 156.30, 135.18, 126.70, 51.22, 30.70, 11.53; HRMS (m/z): [M+H]+ calculated for C8H11N2O2S, 199.0530; found, 199.0548.
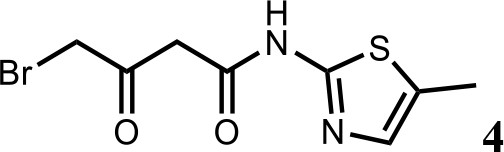
Synthesis of 4-bromo-N-(5-methylthiazol-2-yl)−3-oxobutanamide (4).
To a solution of N-(5-methylthiazol-2-yl)−3-oxobutanamide (3) (2.0 g; 10.1 mmol) in AcOH (30 ml) was added Br2 (2.0 g; 11.1 mmol) at 60°C. The mixture was stirred for 10 min at 60°C, then cooled to room temperature. The reaction mixture was quenched with saturated aqueous Na2SO3 solution, extracted with EtOAc and the organic layer dried over Na2SO4, filtered and evaporated. The product was purified by column chromatography on silica gel to obtain 4-bromo-N-(5-methylthiazol-2-yl)−3-oxobutan amide (4) (2.0 g; 7.25 mmol, 73%). 1H NMR (400 MHz, DMSO-d6) δ 12.02 (s, 1H), 7.14 (s, 1H), 4.48 (s, 2H), 3.83 (s, 2H), 2.34 (s, 3H). 13C NMR (101 MHz, DMSO-d6) δ 196.04, 164.67, 156.24, 135.15, 126.89, 47.91, 37.59, 11.55; HRMS (m/z): [M+H]+ calculated for C8H10BrN2O2S, 276.9641; found, 276.9659.
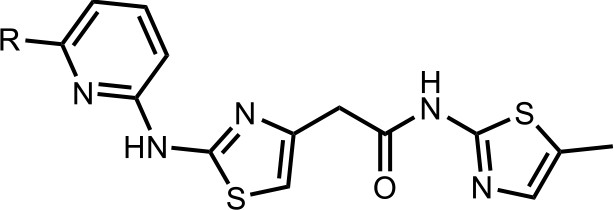
Synthesis of the HQ461 and analogues-A.
To a solution of above 4 (10 mg; 0.036 mmol) in EtOH (2 ml) was added the pyridinyl thiourea 5 (0.054 mmol). The mixture was stirred for 12 hr at 70°C, then cooled to room temperature. The mixture was evaporated to dryness and purified by silica gel chromatography to get HQ461 analogues as HBr salt.
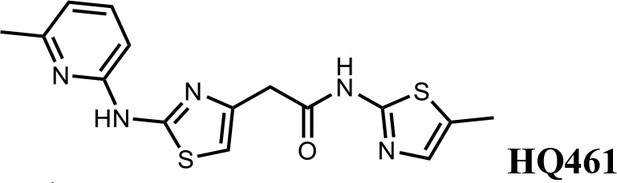
2-(2-((6-methylpyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ461).
Pale yellow solid; Yield 52%; 1H NMR (400 MHz, DMSO-d6) δ 12.12 (s, 1H), 11.67 (s, 1H), 7.66 (t, J = 7.7 Hz, 1H), 7.14 (d, J = 1.2 Hz, 1H), 6.88 (t, J = 10.2 Hz, 3H), 3.85 (s, 2H), 2.47 (s, 3H), 2.33 (s, 3H); 13C NMR (101 MHz, DMSO-d6) δ 167.50, 161.13, 156.59, 154.46, 149.70, 140.66, 138.69, 134.87, 126.86, 117.62, 110.42, 109.70, 36.39, 22.99, 11.61; HRMS (m/z): [M+H]+ calculated for C15H16N5OS2, 346.0791; found, 346.0847.
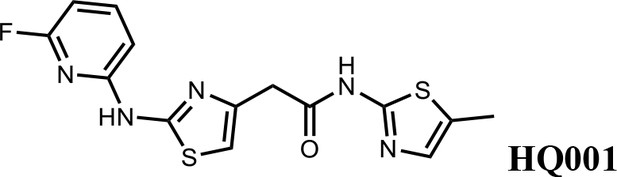
2-(2-((6-fluoropyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ001).
Pale yellow solid; Yield 50%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.53 (dd, J = 15.9, 7.9 Hz, 1H), 6.87 (s, 1H), 6.60 (d, J = 7.7 Hz, 1H), 6.49 (s, 1H), 6.28 (d, J = 7.8 Hz, 1H), 3.61 (s, 2H), 2.20 (s, 3H). 19F NMR (376 MHz, CDCl3: CD3OD = 4:1) δ −66.70; HRMS (m/z): [M+H]+ calculated for C14H13FN5OS2, 350.0540; found, 350.0564.
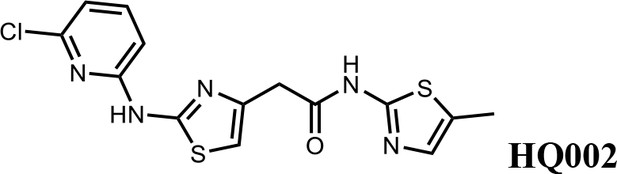
2-(2-((6-chloropyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ002).
Pale yellow solid; Yield 35%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.52 (dt, J = 11.0, 7.6 Hz, 1H), 7.01 (s, 1H), 6.89–6.82 (t, J = 7.2 Hz, 1H), 6.78 (dd, J = 8.2, 3.2 Hz, 1H), 6.58 (s, 1H), 3.76 (s, 2H), 2.35 (s, 3H). HRMS (m/z): [M+H]+ calculated for C14H13ClN5OS2, 366.0245; found, 366.0265.
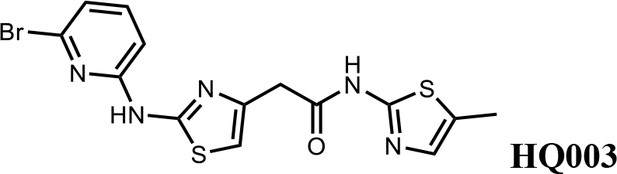
2-(2-((6-bromopyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ003).
Pale yellow solid; Yield 53%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.41 (t, J = 7.9 Hz, 1H), 6.99 (m, 2H), 6.79 (d, J = 8.3 Hz, 1H), 6.58 (s, 1H), 3.74 (s, 2H), 2.32 (s, 3H); HRMS (m/z): [M+H]+ calculated for C14H13BrN5OS2, 409.9739; found, 409.9784.
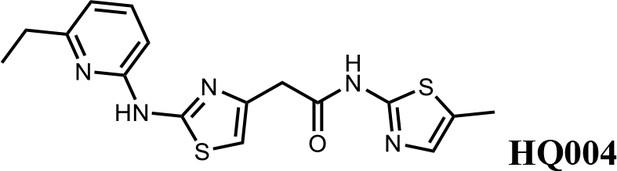
2-(2-((6-ethylpyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ004).
Pale yellow solid; Yield 53%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.52 (t, J = 7.8 Hz, 1H), 7.06 (s, 1H), 6.76 (d, J = 7.6 Hz, 1H), 6.64 (d, J = 7.8 Hz, 1H), 6.59 (s, 1H), 3.79 (s, 2H), 2.83 (q, J = 7.2, 14.8 Hz, 2H), 2.39 (s, 3H), 1.38 (t, J = 7.2, 3H); HRMS (m/z): [M+H]+ calculated for C16H18N5OS2, 360.0947; found, 360.0967.
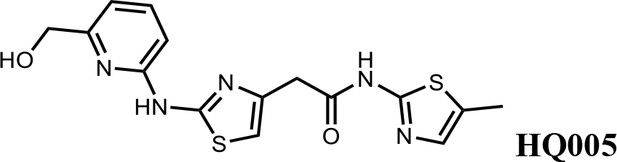
2-(2-((6-(hydroxymethyl)pyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ005).
Pale yellow solid; Yield 33%; 1H NMR (400 MHz, CDCl3) δ 11.93 (s, 1H), 9.83 (s, 1H), 7.61 (t, J = 7.8 Hz, 1H), 7.08 (s, 1H), 6.86 (d, J = 7.6 Hz, 1H), 6.72 (d, J = 7.7 Hz, 1H), 6.63 (s, 1H), 4.80 (s, 2H), 3.80 (s, 2H), 2.40 (s, 3H); HRMS (m/z): [M+H]+ calculated for C15H16N5O2S2, 362.0740; found, 362.0768.
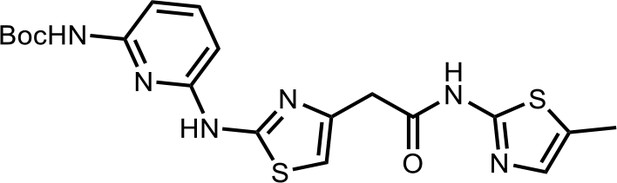
Tert-butyl(6-((4-(2-((5-methylthiazol-2-yl)amino)−2-oxoethyl)thiazol-2-yl)amino)pyridin-2-yl)carbamate (HQ007-01).
White solid; Yield 35%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.60 (t, J = 7.6 Hz, 1H), 7.50 (d, J = 7.7 Hz, 1H), 7.06 (s, 1H), 6.56 (s, 1H), 6.51 (d, J = 7.9 Hz, 1H), 3.79 (s, 2H), 2.39 (s, 3H), 1.549 (s, 9H); HRMS (m/z): [M+H]+ calculated for C19H23N6O3S2, 447.1268; found, 447.1295.
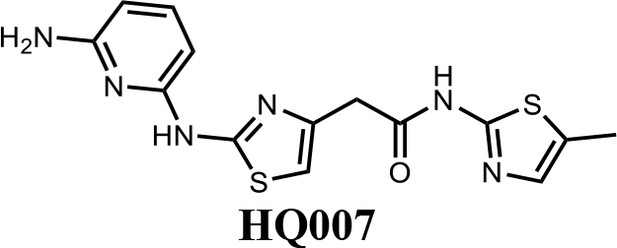
2-(2-((6-aminopyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ007).
To a solution of HQ007-01 (3 mg) in dichloromethane (1 ml) was added trifluoroacetic acid (0.5 ml). The mixture was stirred for 2 hr at room temperature. The mixture was evaporated to dryness and purified by silica gel chromatography to get 2-(2-((6-aminopyridin-2-yl) amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide as TFA salt (2.4 mg, 80%). 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.42 (t, J = 8.3 Hz, 1H), 6.83 (s, 1H), 6.66 (s, 1H), 6.13 (d, J = 7.9 Hz, 1H), 6.06 (d, J = 8.1 Hz, 1H), 3.65 (s, 2H), 2.13 (s, 3H); HRMS (m/z): [M+H]+ calculated for C14H15N6OS2, 347.0743; found, 347.0777.
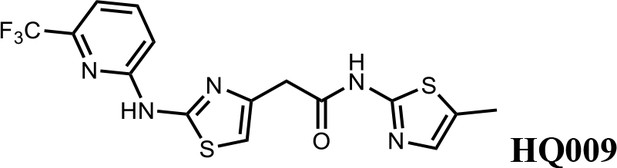
N-(5-methylthiazol-2-yl)−2-(2-((6-(trifluoromethyl)pyridin-2-yl)amino)thiazol-4-yl)acetamide (HQ009).
White solid; Yield 52%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 7.71 (t, J = 8.1 Hz, 1H), 7.18 (d, J = 7.6 Hz, 1H), 7.03 (d, J = 8.1 Hz, 1H), 6.99 (s, 1H), 6.59 (s, 1H), 3.75 (s, 2H), 2.33 (s, 3H). 19F NMR (376 MHz, CDCl3: CD3OD = 4:1) δ −68.59; HRMS (m/z): [M+Na]+ calculated for C15H12F3N5OS2Na, 422.0333; found, 422.0359.
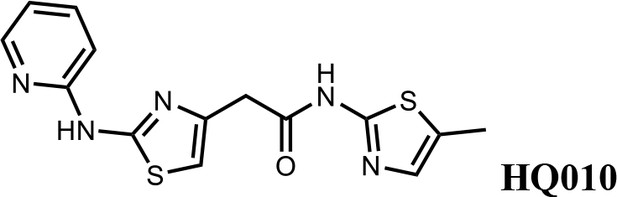
N-(5-methylthiazol-2-yl)−2-(2-(pyridin-2-ylamino)thiazol-4-yl)acetamide (HQ010).
Pale yellow solid; Yield 53%; 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 8.28 (d, J = 4.6 Hz, 1H), 7.57 (t, J = 7.4 Hz, 1H), 6.99 (s, 1H), 6.85 (m, 2H), 6.51 (s, 1H), 3.73 (s, 2H), 2.33 (s, 3H); HRMS (m/z): [M+H]+ calculated for C14H14N5OS2, 332.0634; found, 332.0663.
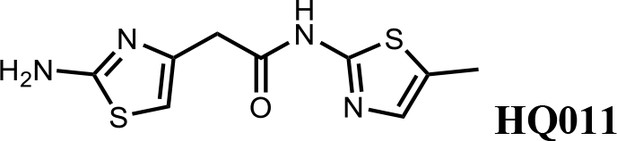
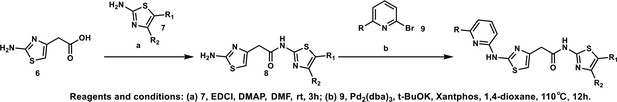
Synthesis of 2-(2-aminothiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ011).
To a mixture of 2-(2-aminothiazol-4-yl)acetic acid (6) (1.00 g; 6.33 mmol) and 5-methylthiazol-2-amine (1) (722 mg; 6.33 mmol) in dimethylformamide (15 ml) was added 1-Ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (1.82 g; 9.50 mmol) and dimethylaminopyridine (154 mg; 1.27 mmol) under argon atmosphere. The mixture stirred at room temperature for 3 hr. The reaction mixture was quenched with water, extracted with EtOAc and the organic layer dried over Na2SO4, filtered and evaporated. The product was purified by column chromatography on silica gel to obtain 2-(2-aminothiazol-4-yl)-N-(5-methylthiazol-2-yl) acetamide (7) (1.15 g; 4.53 mmol, 72%). 1H NMR (400 MHz, DMSO-d6) δ 11.97 (s, 1H), 7.12 (s,, 1H), 6.95 (s, 2H), 6.32 (s, 1H), 3.56 (s, 2H), 2.33 (s, 3H); 13C NMR (101 MHz, DMSO-d6) δ 168.82, 168.27, 156.59, 145.33, 135.17, 126.54, 103.53, 38.52, 11.57; HRMS (m/z): [M+Na]+ calculated for C9H10N4OS2Na, 277.0194; found, 277.0209.
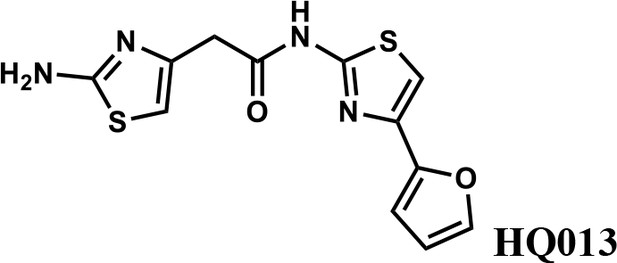
2-(2-aminothiazol-4-yl)-N-(4-(furan-2-yl)thiazol-2-yl)acetamide (HQ013).
To a mixture of 2-(2-aminothiazol-4-yl)acetic acid (6) (100 mg; 0.63 mmol) and 4-(furan-2-yl)thiazol-2-amine (105 mg; 0.63 mmol) in dimethylformamide (5 ml) was added 1-Ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (182 mg; 0.95 mmol) and dimethylaminopyridine (15 mg; 0.13 mmol) under argon atmosphere. The mixture stirred at room temperature for 3 hr. The reaction mixture was quenched with water, extracted with EtOAc and the organic layer dried over Na2SO4, filtered and evaporated. The product was purified by column chromatography on silica gel to obtain 2-(2-aminothiazol-4-yl)-N-(4-(furan-2-yl)thiazol-2-yl)acetamide (HQ013) (124 mg; 0.41 mmol, 65%). 1H NMR (400 MHz, DMSO-d6) δ 12.43 (s, 1H), 7.72 (dd, J = 1.8, 0.8 Hz, 1H), 7.32 (s, 1H), 6.93 (s, 2H), 6.67 (d, J = 2.7 Hz, 1H), 6.58 (dd, J = 3.3, 1.8 Hz, 1H), 6.35 (s, 1H), 3.60 (s, 2H); HRMS (m/z): [M+H]+ calculated for C12H11N4O2S2, 307.0318; found, 307.0337.
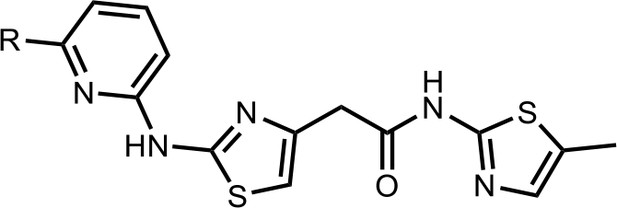
Synthesis of the HQ461 and analogues-B.
A mixture of HQ011 (20 mg, 0.08 mmol), 9 (0.096 mmol), Xantphos (5.6 mg, 0.0096 mmol) Tris(dibenzylideneacetone)dipalladium (8.8 mg, 0.0096 mmol), tBuOK (21 mg, 0.19 mmol) in 1,4-dioxane(2.0 ml) under argon atmosphere was stirred at 110°C for 12 hr. The mixture was filtered and concentrated in vacuum and the crude product purified by column chromatography on silica gel.
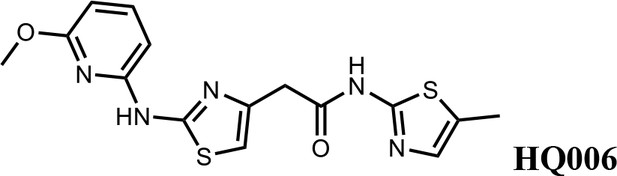
2-(2-((6-methoxypyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ006).
Pale yellow solid; Yield 33%; 1H NMR (400 MHz, CDCl3) δ 11.60 (s, 1H), 9.36 (s, 1H), 7.51 (t, J = 7.9 Hz, 1H), 7.09 (s, 1H), 6.62 (s, 1H), 6.35 (dd, J = 7.9, 3.3 Hz, 2H), 4.08 (s, 3H), 3.79 (s, 2H), 2.40 (s, 3H); HRMS (m/z): [M+H]+ calculated for C15H16N5O2S2, 362.0740; found, 362.0768.
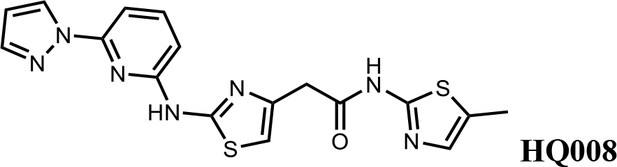
2-(2-((6-(1H-pyrazol-1-yl)pyridin-2-yl)amino)thiazol-4-yl)-N-(5-methylthiazol-2-yl)acetamide (HQ008).
Pale yellow solid; Yield 25%; 1H NMR (400 MHz, DMSO-d6) δ 12.09 (s, 1H), 11.63 (s, 1H), 8.75 (d, J = 2.2 Hz, 1H), 7.91–7.81 (m, 2H), 7.42 (d, J = 7.9 Hz, 1H), 7.14 (s, 1H), 6.94–6.88 (m, 2H), 6.64 (s, 1H), 3.80 (s, 2H), 2.33 (s, 3H); HRMS (m/z): [M+H]+ calculated for C17H16N7OS2, 398.0852; found, 398.0882.
N-(4-(furan-2-yl)thiazol-2-yl)−2-(2-((6-methylpyridin-2-yl)amino)thiazol-4-yl)acetamide (HQ012).
A mixture of HQ013 (20 mg, 0.08 mmol), 2-bromo-6-methylpyridine (16.4 mg, 0.096 mmol), Xantphos (5.6 mg, 0.0096 mmol), Tris(dibenzylideneacetone)dipalladium (8.8 mg, 0.0096 mmol), t-BuOK (21 mg, 0.19 mmol) in 1,4-dioxane (2.0 ml) under argon atmosphere was stirred at 110°C for 12 hr. The mixture was filtered and concentrated in vacuum and the crude product purified by column chromatography on silica gel to obtain N-(4-(furan-2-yl)thiazol-2-yl)−2-(2-((6-methylpyridin-2-yl)amino)thiazol-4-yl) acetamide (HQ012) (9.8 mg; 0.026 mmol, 21%). 1H NMR (400 MHz, CDCl3) δ 11.99 (s, 1H), 9.40 (s, 1H), 7.54–7.47 (m, 1H), 7.43 (d, J = 1.8 Hz, 1H), 7.08 (s, 1H), 6.76 (d, J = 7.4 Hz, 1H), 6.69 (d, J = 3.3 Hz, 1H), 6.64 (d, J = 8.1 Hz, 1H), 6.57 (s, 1H), 6.48–6.43 (m, 1H), 3.82 (s, 2H), 2.55 (d, J = 2.9 Hz, 3H); HRMS (m/z): [M+H]+ calculated for C18H16N5O2S2, 398.0740; found, 398.0784.
Synthesis of 6-bromo-N-(4-(2-((5-methylthiazol-2-yl)amino)−2-oxoethyl)thiazol-2-yl) Picolinamide (HQ014).
A mixture of HQ011 (25 mg, 0.10 mmol), 6-bromopicolinic acid (20 mg, 0.10 mmol), 1-Ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (28 mg, 0.15 mmol) dimethylaminopyridine (1.22 mg, 0.01 mmol) in dimethylformamide (2.0 ml) under argon atmosphere was stirred at room temperature for 12 hr. The reaction mixture was quenched with water, extracted with EtOAc and the organic layer dried over Na2SO4, filtered and evaporated. The product was purified by column chromatography on silica gel to obtain HQ014 (28 mg; 0.064 mmol, 83%). 1H NMR (400 MHz, CDCl3: CD3OD = 4:1) δ 8.18 (dd, J = 7.5, 0.6 Hz, 1H), 7.76 (t, J = 7.7 Hz, 1H), 7.71–7.66 (m, 1H), 6.99 (s, 1H), 6.87 (s, 1H), 3.82 (s, 2H), 2.32 (s, 3H); HRMS (m/z): [M+Na]+ calculated for C15H12BrN5O2S2Na, 459.9513; found, 459.9541.

Enrichment of polyubiquitinated proteins by FLAG-TUBE.
293F cells were transiently transfected with plasmid expressing FLAG-TUBE. Twenty-four hours later, transfected cells were treated with 10μM HQ461 for 0, 1, 2, 4 or 8 hours.FLAG-TUBE and its interacting proteins were immunoprecipitated by anti-FLAG antibody.
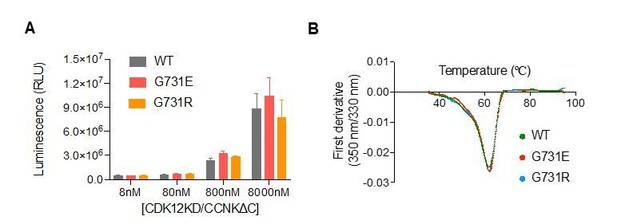
CDK12G731E/R mutations do not affect CDK12 kinase activity or protein stability.
(A) CDK12KD/CCNKΔC (WT or G731E/R mutants) kinase activity was perform in 10μL reaction volume containing 45nM substrate (GST-CTD) and a reaction buffer of 20mM HEPES, pH7.9, 20mM MgCl2, 2mM DTT, 1mM ATP. The reaction was initiated by adding 8nM, 80nM, 800nM, or 8μM CDK12KD/CCNKΔC (WT or G731E/R mutants) and incubated at 30°C for 1hour followed by the addition of 10μL ADP-GloTM Reagent (Promega, V6930) to terminate kinase reaction and remove unreacted ATP. After incubation at room temperature for 40 minutes, the mixtures were transferred from PCR tubes to a 384-well plate and mixed with 20μL Kinase Detection Reagent (Promega, V6930). The mixtures were allowed to sit at room temperature for 40 minutes before reading luminescence by EnVision multimode plate reader (Perkin Elmer). Error bars represent SD from three replicates. (B) Thermal denaturation of CDK12KD/CCNKΔC (WT or G731E/R mutants) measured by nano DSF.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | A549 | Dr. Deepak Nijhawan’s lab at University of Texas Southwestern Medical Center | Male | |
| Cell line (Homo sapiens) | HCT116 | Dr. Deepak Nijhawan’s lab at University of Texas Southwestern Medical Center | Male | |
| Cell line (Homo sapiens) | HEK293T | Dr. Deepak Nijhawan’s lab at University of Texas Southwestern Medical Center | Female | |
| Cell line (Spodoptera frugiperda) | SF9 | Dr. Sanduo Zheng from National Institute of Biological Sciences, Beijing | ||
| Cell line (Trichoplusia ni) | High Five | Dr. Sanduo Zheng from National Institute of Biological Sciences, Beijing | ||
| Cell line (Homo sapiens) | Free style 293 F | Dr. Linfeng Sun at China University of Science and Technology | Female | |
| Antibody | Anti-NRF2 (Rabbit monoclonal) | Abcam (ab62352) | RRID:AB_944418 | WB (1:4000) |
| Antibody | Anti-β-Actin-HRP (Mouse monoclonal) | Huaxingbio (HX18271) | WB (1:10000) | |
| Antibody | Anti-α-Tubulin-HRP (Rabbit polyclonal) | MBL Life Science (PM054-7) | RRID:AB_10695326 | WB (1:10000) |
| Antibody | Anti-DDB1 (Rabbit monoclonal) | Abcam (ab109027) | RRID:AB_10859111 | WB (1:10000) |
| Antibody | Anti-RBX1 (Rabbit polyclonal) | Proteintech(14895–1-AP) | RRID:AB_2179719 | WB (1:5000) |
| Antibody | Anti-UBE2G1 (Rabbit polyclonal) | Proteintech(12012–1-AP) | RRID:AB_10665812 | WB (1:2000) |
| Antibody | Anti-CDK12 (Rabbit polyclonal) | Cell Signaling Technology (11973S) | RRID:AB_2715688 | WB (1:4000) |
| Antibody | Anti-CCNK (Rabbit polyclonal) | Bethyl lab (A301-939A-T) | RRID:AB_2780226 | WB (1:4000) |
| Antibody | Anti-FLAG-HRP (Mouse monoclonal) | Sigma-Aldrich (A8592) | RRID:AB_439702 | WB (1:10000) |
| Antibody | Anti-HA (Mouse monoclonal) | BioLegend (901533) | RRID:AB_2801249 | WB (1:4000) |
| Antibody | Anti-RNA polymerase II subunit B1 (phospho CTD Ser-2) (Rat monoclonal) | Sigma-Aldrich (041571) | RRID:AB_10627998 | WB (1:5000) |
| Antibody | Anti-RNA polymerase II CTD repeat YSPTSPS (Mouse monoclonal) | Abcam (ab817) | RRID:AB_306327 | WB (1:5000) |
| Antibody | Anti-rabbit IgG (Goat polyclonal) | Cell Signaling Technology (7074S) | RRID:AB_2099233 | WB (1:10000) |
| Antibody | Anti-mouse IgG (Goat polyclonal) | Zsbio (ZB-2305) | RRID:AB_2747415 | WB (1:10000) |
| Antibody | Anti-rat IgG (Goat polyclonal) | Sino Biological (SSA005) | WB (1:10000) | |
| Recombinant DNA reagent | Lenti-EF1α−3xFLAG-CDK12-P2A-BSD WT/GE/GR | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pCDNA3.1-1XFLAG-CDK12 WT/GE/GR | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pCDNA3.1-1XFLAG-CDK12-kinase domain (715–1052) WT/GE/GR | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pCDNA3.1-3XFLAG-CDK12/13-kinase domain (715–1052/693-1030) WT/GE/GR | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pcDNA3.1-CCNK-3xHA | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pET28a-6xHis-Ub | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pCMV-8×His-Ub | Dr. William Kaelin at Dana Farber Cancer Institute of Harvard | ||
| Recombinant DNA reagent | pPB-CAG-1xFLAG-UBA1 | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pPB-CAG-1xFLAG-UBE2D3 | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pPB-CAG-1xFLAG-CUL4A | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pPB-CAG-RBX1 | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pPB-CAG-1xFLAG-Avi-DDB1 | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pFastBac-Strep-8×His-DDB1ΔBPB | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pFastBac-6×His-CDK12KD | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pFastBac-6×His-CCNKΔC | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pFastBac-CAK1 | This paper | Described in Materials and methods; available upon request | |
| Recombinant DNA reagent | pCDNA3.1-CCNK-luc | This paper | Described in Materials and methods; available upon request | |
| Peptide, recombinant protein | HBx peptide | ChinaPetides | ILPKVLHKRTLGLS | |
| Commercial assay or kit | CellTiter-Glo | Promega | G7570 | |
| Commercial assay or kit | AlphaScreen Histidine (Nickel Chelate) Detection Kit | Perkin Elmer | 6760619C | |
| Commercial assay or kit | AlphaScreen Streptavidin Donor beads | Perkin Elmer | 6760619C | |
| Chemical compound, drug | Bortezomib | Targetmol | T2399 | CAS No. 179324-69-7 |
| Chemical compound, drug | THZ531 | Targetmol | T4293 | CAS No. 1702809-17-3 |
| Chemical compound, drug | Dinaciclib | Targetmol | T1912 | CAS No. 779353-01-4 |
| Chemical compound, drug | MLN4924 | Selleckchem | S7109 | CAS No. 905579-51-3 |
| Software, algorithm | BWA | Li, 2013 | v0.7.17 | |
| Software, algorithm | samtools | Li et al., 2009 | v1.10 | |
| Software, algorithm | GATK | DePristo et al., 2011; McKenna et al., 2010; Van der Auwera et al., 2013 | v4.1.7 | |
| Software, algorithm | SnpSift | Cingolani et al., 2012a | v4.3t | |
| Software, algorithm | MAGeCK | Li et al., 2014 | v0.5.9.2 | |
| Software, algorithm | ggplot2 | Wickham, 2016 | ||
| Software, algorithm | GraphPad Prism | v8.0.2 | ||
| Software, algorithm | pLink 2 | Chen et al., 2019 |