Effects of domestication on the gut microbiota parallel those of human industrialization
Figures
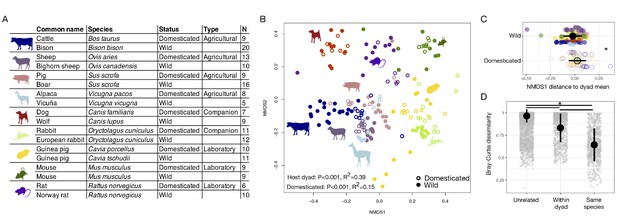
The mammalian gut microbiota carries a global signature of domestication.
(A) Sampling scheme for cross-species study. (B) Nonmetric multidimensional scaling (NMDS) ordination of Bray–Curtis dissimilarities illustrates a significant signal (p<0.001, R2 = 0.15, F = 6.081, permutational MANOVA) of domestication (closed versus open circles; N = 82 domesticated and 99 wild) and clustering by host dyad (color; N = 5–20 individuals per species). (C) Distance to dyad (color) mean along Bray–Curtis ordination NMDS axis 1 differs by domestication status (p=0.006, Mann–Whitney U test). (D) Bray–Curtis dissimilarity between individuals is lowest among conspecifics, but wild–domesticated pairs also have lower dissimilarity than unrelated pairs (p<0.001, bootstrapped Mann-Whitney U tests). * indicates p<0.05. Large circles are means; bars show standard deviations.
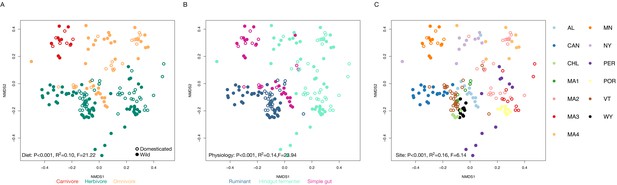
Alternative factors associated with gut microbiota variation among wild and domesticated mammals.
Diet type (A), digestive physiology (B), and collection locale (C) are significantly associated (permutational MANOVAs) with variation in microbial community composition among wild and domesticated mammals, visualized here with nonmetric multidimensional scaling (NMDS) ordination of Bray–Curtis dissimilarity. Site abbreviations are as listed in 'Materials and methods'.
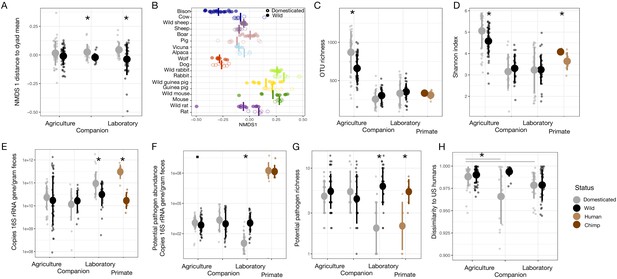
Microbial parameter comparisons between wild and domesticated mammals and between chimpanzees and humans.
Bray–Curtis nonmetric multidimensional scaling (NMDS) ordination shifts for domestication groups (A) and dyads (B), OTU richness (C), Shannon index (D), bacterial density (E), potential human pathogen (see Kembel et al., 2012; Reese et al., 2016) abundance (F), and potential pathogen richness (G) varied by domestication status for at least one domestication type (agriculture, companion, or laboratory) in our cross-species dataset. Trends often mirrored those seen in comparing US humans to wild chimpanzees. Among domesticated animals, dissimilarity from US humans (H) was greater for agricultural animals than for companion or laboratory animals. However, among wild animals, dissimilarity from US humans did not differ by type. * indicates p<0.05, • indicates p<0.10, Mann–Whitney U test. Large circles are means; bars show standard deviations.
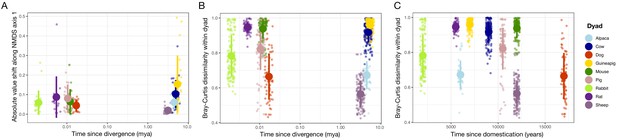
Relatedness was correlated with ordination shifts but not dissimilarity within a dyad.
(A) The absolute value of shifts along nonmetric multidimensional scaling (NMDS) axis 1 for individuals relative to their dyad average was higher for dyads with a greater time since divergence (p=0.012, rho = 0.19, Spearman correlation). (B, C) Average Bray–Curtis dissimilarity between members of wild–domestic dyads was not correlated with time since divergence (p=0.380; B) or time since domestication (p=0.854; C). Large circles are means; bars show standard deviations.
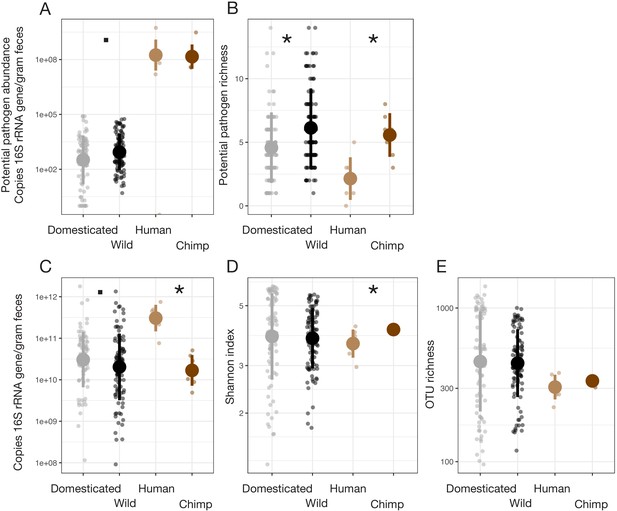
Domestication explained few global gut microbial community characteristics in our cross-species dataset.
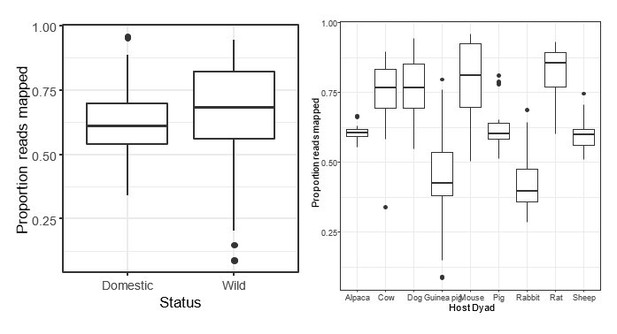
Potential pathogen richness (B) but not abundance (A) was significantly higher in wild animals. (C–E) Microbial density (quantified as copies of the 16S rRNA gene per gram feces [C]) and alpha-diversity metrics (Shannon index [D] and OTU richness [E]) did not vary consistently between wild and domesticated animals. Generally, greater differences were observed in comparisons between wild chimpanzees and a US human population. * indicates p<0.05, • indicates p<0.10, Mann–Whitney U test. Large circles are means; bars show standard deviations.
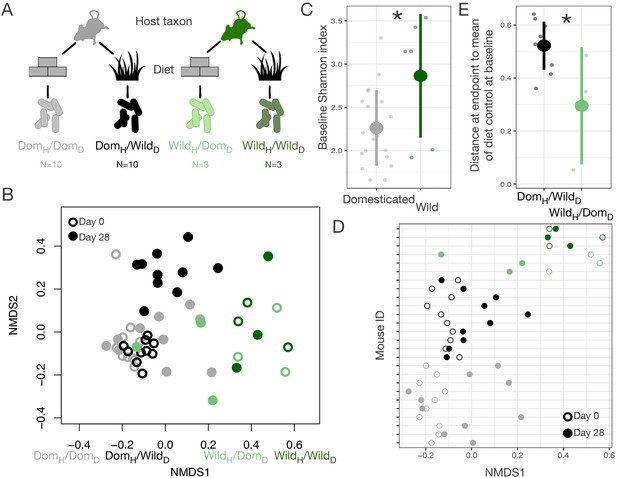
Gut microbial differences between wild and domesticated mice can be partially overcome by diet swap.
(A) Design scheme for fully factorial host taxon by diet mouse experiment (N = 10 laboratory mice or three wild mice per diet group). (B) Nonmetric multidimensional scaling (NMDS) ordination of Bray–Curtis dissimilarities showing changes for mice from day 0 (open circle) to day 28 (filled circle) by experimental groups (color). Composition varied by host taxon (p<0.001, R2 = 0.173, F = 64.255, permutational MANOVA), diet (p<0.001, R2 = 0.042, F = 15.427), and a host taxon by diet interaction (p<0.001, R2 = 0.020, F = 7.557). (C) Shannon index differed between host taxa on day 0 (p=0.011, Mann–Whitney U test). (D) Animals on reciprocal diets (DomH/WildD and WildH/DomD) but not control diets tended to move in opposite directions along Bray–Curtis ordination NMDS axis 1 from day 0 to day 28 (p=0.048 and p=0.25, respectively, one-sample Wilcoxon test). (E) At the end of the experiment, distance to the mean of the diet control at baseline (DomH/DomD and WildH/WildD) was lower for wild mice than for laboratory mice (p=0.048, Mann–Whitney U test). * indicates p<0.05, Mann–Whitney U test. Large circles are means; bars show standard deviations.
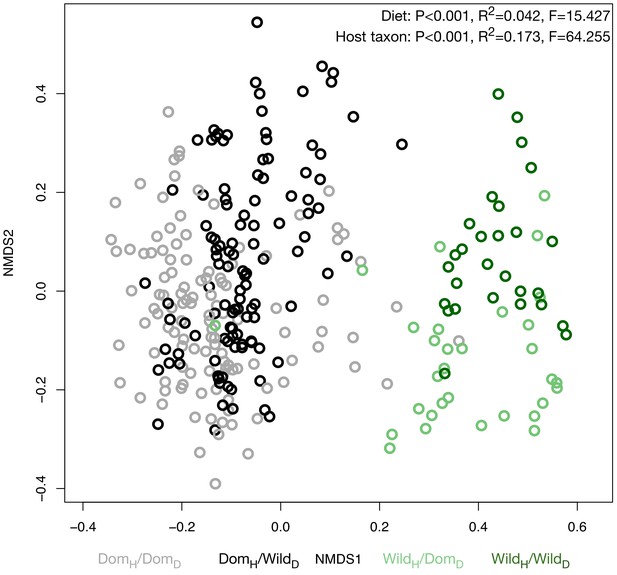
Microbiota composition at all time points during the mouse diet swap experiment.
Nonmetric multidimensional scaling (NMDS) ordination of all time points illustrates significant effects (permutational MANOVA tests) of host taxon and diet on Bray–Curtis dissimilarity.
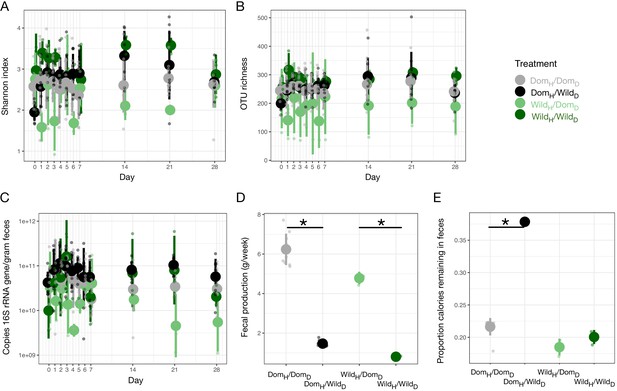
Select microbial and host metabolic parameters under mouse diet swap.
(A) Shannon index by experimental groups over time.(B) OTU richness by experimental groups over time. (C) Microbial load by experimental groups over time. (D) Total fecal production over 1 week differs between experimental groups. (E) Calories remaining in feces as a function of total calories consumed varies by diet in DomH mice. * in (D) and (E) indicates p<0.05, Mann–Whitney U test. Large circles are means; bars show standard deviations.
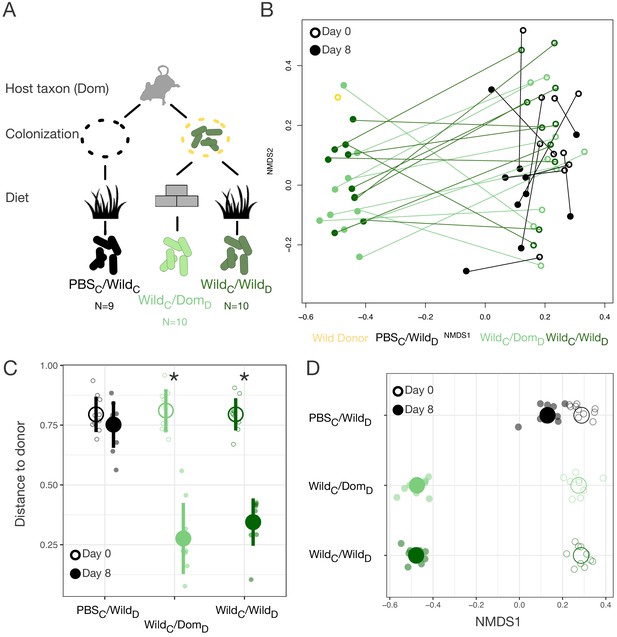
Laboratory mice can be re-wilded through colonization with a wild gut microbial community.
(A) Design scheme for colonization/diet mouse experiment (N = 9–10 mice per treatment group). (B) Nonmetric multidimensional scaling (NMDS) ordination of Bray–Curtis dissimilarities showing changes for mice from day 0 (open circles) to day 8 (filled circles) by experimental groups (color). (C, D) At the end of the experiment (filled circle), distance to the wild community donor decreased most in animals colonized with wild communities (p=0.004 WildC/DomD and p=0.002 WildC/WildD, Mann–Whitney U test; C), but all experimental groups exhibited change along Bray–Curtis ordination NMDS axis 1 (p=0.002 PBSC/WildD, p=0.004 WildC/DomD, and p=0.002 WildC/WildD, one-sample Wilcoxon tests; D) during the course of the experiment. * in (C) indicates p<0.05, Mann–Whitney U test comparing day 0 to day 8 for each experimental group. Large circles are means; bars in (C) show standard deviations.
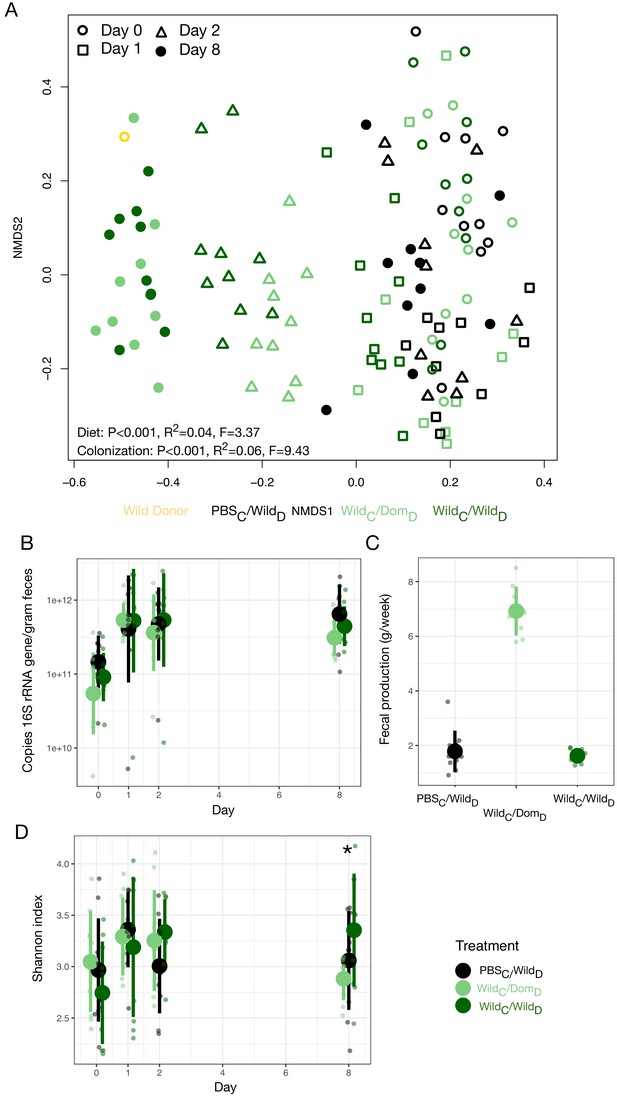
Select microbial and host metabolic parameters under wild colonization treatment.
(A) Nonmetric multidimensional scaling (NMDS) ordination of all time points illustrates significant effects (permutational MANOVA tests) of colonization and diet treatment on Bray–Curtis dissimilarity.(B) Microbial load by experimental groups over time. (C) Total fecal production over 1 week differs between experimental groups. (D) Shannon index by experimental groups over time. * indicates p<0.05, Kruskal–Wallis test. Large circles are means; bars show standard deviations.
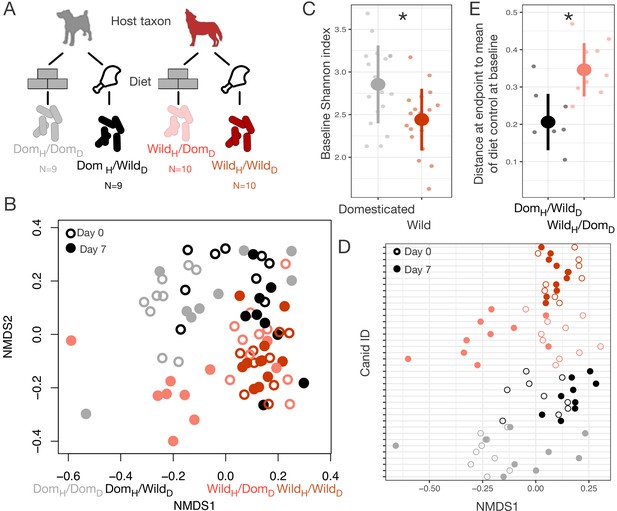
Microbial differences between wild and domesticated canids can be partially overcome by diet shifts.
(A) Design scheme for fully factorial host taxon by diet canid experiment (N = 9 dogs or N = 10 wolves per diet group). (B) Nonmetric multidimensional scaling (NMDS) ordination of Bray–Curtis dissimilarities showing changes for canids from day 0 (open circle) to day 7 (filled circle) by experimental groups (color). Composition varied by host taxon (p<0.001, R2 = 0.098, F = 13.70, permutational MANOVA), diet (p<0.001, R2 = 0.058, F = 8.15), and a host taxon by diet interaction (p<0.001, R2 = 0.028, F = 3.93). (C) Shannon index differed between dogs and wolves on day 0 (p=0.003, Mann–Whitney U test). (D) Canids on reciprocal diets (DomH/WildD and WildH/DomD) but not control diets moved in opposite directions along Bray–Curtis ordination NMDS axis 1 over time (p=0.004 and 0.002, respectively, one-sample Wilcoxon tests). (E) At the end of the experiment, distance to the mean of diet controls at baseline (DomH/DomD and WildH/WildD) was lower for dogs than for wolves on reciprocal diets (p=0.001, Mann–Whitney U test). * indicates p<0.05, Mann–Whitney U test. Large circles are means; bars show standard deviations.
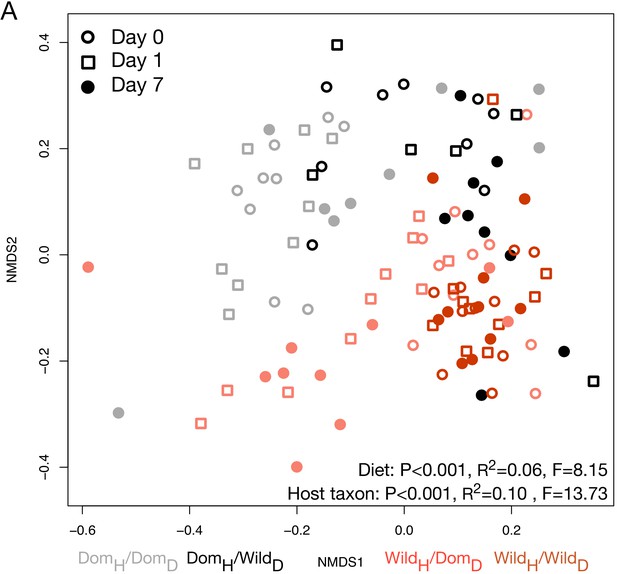
Microbiota composition at all time points during the canid diet swap experiment.
Nonmetric multidimensional scaling (NMDS) ordination of all time points illustrates significant effects (permutational MANOVA tests) of host taxon and diet on Bray–Curtis dissimilarity even after a single day.
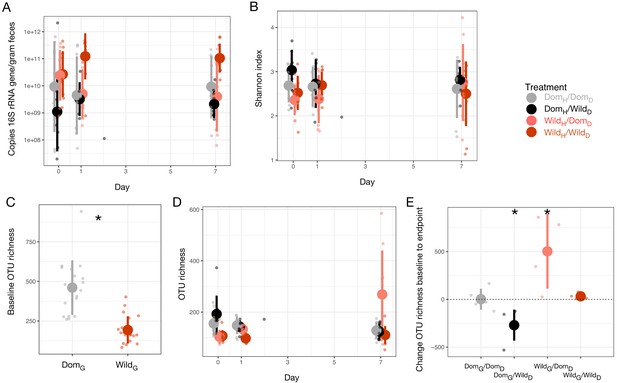
Select microbial parameters under canid diet swap.
(A) Microbial loads of experimental groups over time.(B) Shannon index by experimental groups over time. (C) OTU richness differed between genotypes on day 0. * indicates p<0.05, Mann–Whitney U test. (D) OTU richness by experimental groups over time. (E) Change in OTU richness from day 0 to day 7 differed significantly from 0 in opposite directions for animals on altered diets (DomG/WildD and WildG/DomD). * indicates p<0.05, one-sample Wilcoxon test, and dashed line indicates a shift of 0. Large circles are means; bars show standard deviations.
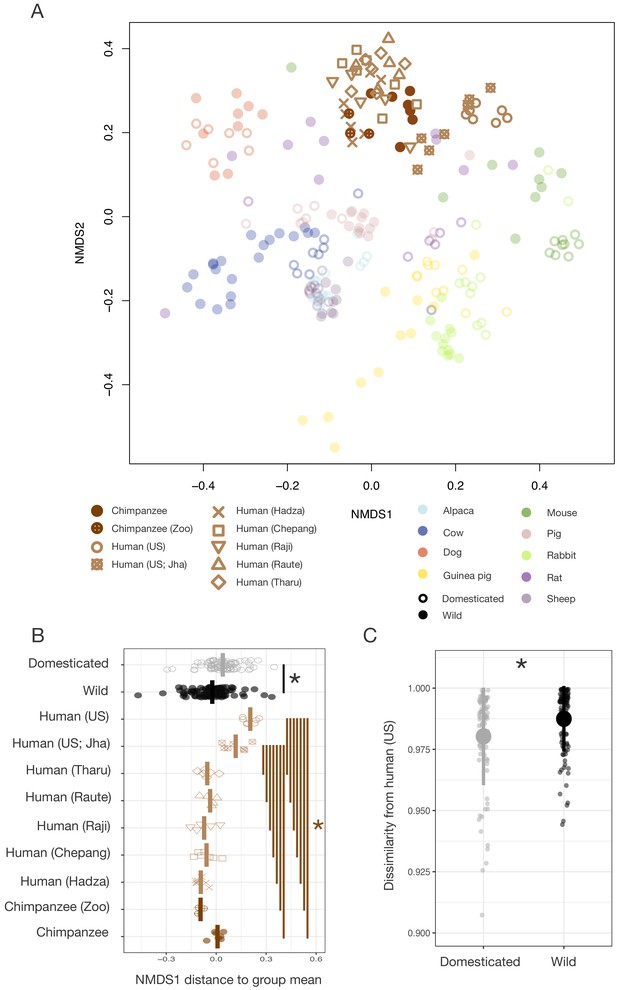
Differences in gut microbial communities between industrialized humans and wild chimpanzees parallel those observed between domesticated and wild mammals.
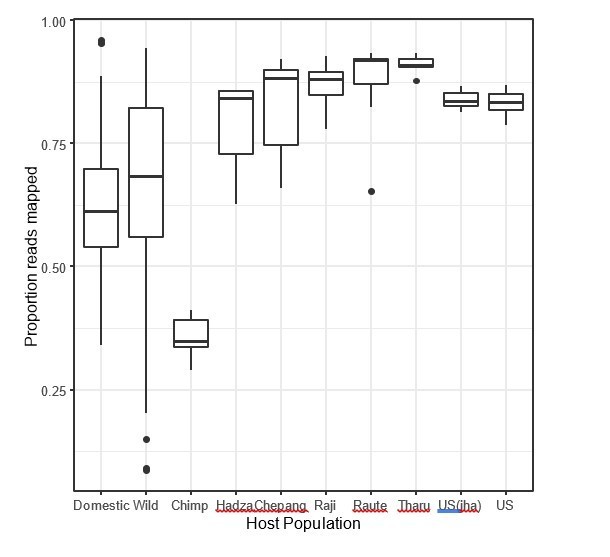
(A) Nonmetric multidimensional scaling (NMDS) ordination of Bray–Curtis dissimilarities in the gut microbiota illustrates that industrialized human populations (US and US Jha) exhibit similar trends relative to wild chimpanzees as domesticated animals do to wild animals, but that non-industrialized human populations (Hadza, Chepang, Raji, Raute, and Tharu) do not (N = 5–7 individuals per primate population and 5–20 individuals per other animal species). (B) Distance along the first Bray–Curtis ordination NMDS axis relative to group mean differs in the same direction for the two industrialized human populations relative to wild chimpanzees or non-industrialized human populations as for domesticated animals relative to wild animals (p<0.05, Mann–Whitney U tests, N = 7–99). (C) The gut microbial communities of wild animals are more dissimilar to those of industrialized humans than are those of domesticated animals (p<0.001, bootstrapped Mann–Whitney U test, N = 82 domesticated and 99 wild). * indicates p<0.05, Mann–Whitney U test. Large shapes are means; bars in (C) show standard deviations.
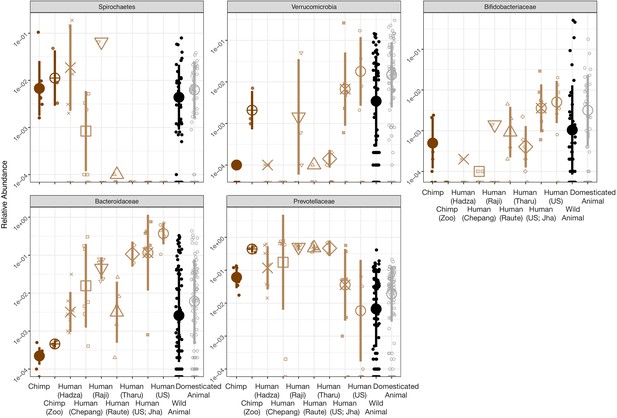
Trends in gut microbial taxa previously linked to differing human lifestyles.
Abundance of gut microbial taxa previously identified as distinguishing among human lifestyles (Smits et al., 2017) trended in similar directions between domesticated and wild animals as between industrialized humans and wild chimpanzees, except for the bacterial families Prevotellaceae and Spirochaetes. Human populations represented include the Hadza (Tanzanian hunter-gatherers), Chepang (Nepalese foragers), Raji (Nepalese foragers transitioning to subsistence farming), Raute (Nepalese foragers transitioning to subsistence farming), Tharu (Nepalese subsistence farmers), and Americans drawn from Jha et al., 2018 as well as Americans sampled and sequenced by our team. Large shapes are means; bars show standard deviations.
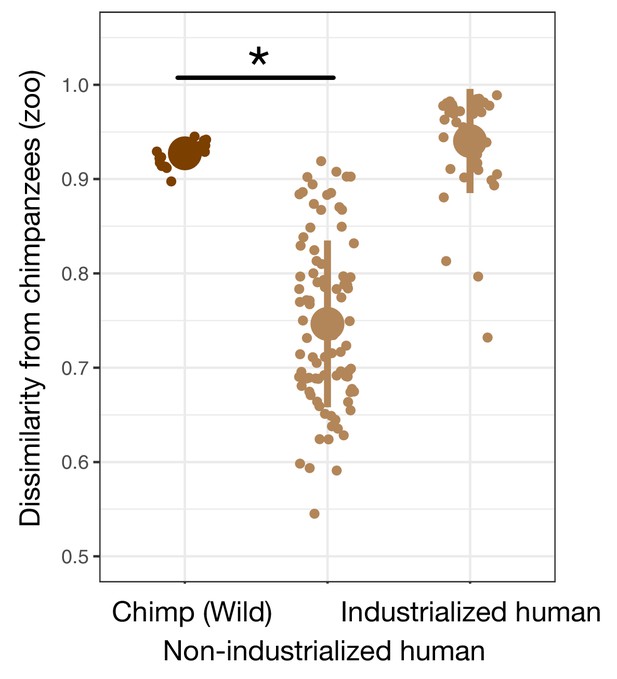
Dissimilarity in the gut microbiota among chimpanzees and humans.
Bray–Curtis dissimilarity among samples taken from captive zoo chimpanzees, wild chimpanzees, and human populations shows that the gut microbial communities of captive chimpanzees are more similar to those of non-industrialized humans, but not industrialized humans, than to those of wild chimpanzees. * indicates p<0.05, Bonferroni-corrected Mann–Whitney U test comparing to wild chimpanzees; all groups were statistically greater than zoo chimpanzees’ dissimilarity to other zoo chimpanzees. Large shapes are means; bars show standard deviations.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Biological sample (Bos taurus) | Feces | This paper | N = 9, sex unknown | |
| Biological sample (Bison bison) | Feces | This paper | N = 20, sex unknown | |
| Biological sample (Ovis aries) | Feces | This paper | N = 13, twelve females | |
| Biological sample (Ovis canadensis) | Feces | This paper | N = 10, sex unknown | |
| Biological sample (Sus scrofa domesticus) | Feces | This paper | N = 9, sex unknown | |
| Biological sample (Sus scrofa) | Feces | This paper | N = 16, five females | |
| Biological sample (Vicugna pacos) | Feces | This paper | N = 8, sex unknown | |
| Biological sample (Vicugna vicugna) | Feces | This paper | N = 5, two females | |
| Biological sample (Canis lupus familiaris) | Feces | This paper | Comparative: N = 7, four females Experiment: N = 9, sex unknown | |
| Biological sample (Canis lupus) | Feces | This paper | Comparative: N = 9, sex unknown Experiment: N = 10, sex unknown | |
| Biological sample (Oryctolagus cuniculus) | Feces | This paper | Domesticated: N = 11, four femalesWild: N = 12, sex unknown | |
| Biological sample (Cavia porcellus) | Feces | This paper | N = 10, zero female | |
| Biological sample (Cavia tschudii) | Feces | This paper | N = 11, sex unknown | |
| Biological sample(Mus musculus) | Feces | This paper | Comparative: N = 9 (domesticated), zero female N = 9 (wild), sex unknown Experiments: N = 49 (domesticated), zero female N = 6 (wild), sex unknown | |
| Biological sample (Rattus norvegicus) | Feces | This paper | Domesticated: N = 6, sex unknown | |
| Biological sample (Rattus norvegicus) | Intestinal sample | This paper | Wild: N = 10, three females | |
| Biological sample (Pan troglodytes) | Feces | This paper | Wild: N = 7, seven females Captive: N = 3, two females | |
| Biological sample (Homo sapiens) | Feces | This paper | N = 7, five females | |
| Sequence-based reagent | 515F | Caporaso et al., 2011 | PCR primers | GTGCCAGCMGCCGCGGTAA |
| Sequenced-based reagent | 806R | Caporaso et al., 2012 | PCR primers | GGACTACNVGGGTWTCTAAT |
| Software, algorithm | R | R Core Team | Version 3.3 | |
| Software, algorithm | QIIME | Caporaso et al., 2010 | Version 1.8 | |
| Software, algorithm | vegan | Oksanen et al., 2017 | ||
| Software, algorithm | lme4 | Bates et al., 2015 | ||
| Software, algorithm | TimeTree | Kumar et al., 2017 | http://timetree.org | |
| Software, algorithm | boot | Canty and Ripley, 2020 | Version 1.3-25 |
Additional files
-
Supplementary file 1
Beta-diversity and nonmetric multidimensional scaling (NMDS) shift analyses were generally robust to the distance metric used and to subsetting the dataset.
- https://cdn.elifesciences.org/articles/60197/elife-60197-supp1-v1.xlsx
-
Supplementary file 2
Expanded sampling metadata.
- https://cdn.elifesciences.org/articles/60197/elife-60197-supp2-v1.xlsx
-
Supplementary file 3
Nutritional information for experimental diets.
- https://cdn.elifesciences.org/articles/60197/elife-60197-supp3-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/60197/elife-60197-transrepform-v1.docx