Thymic stromal lymphopoietin limits primary and recall CD8+ T-cell anti-viral responses
Figures
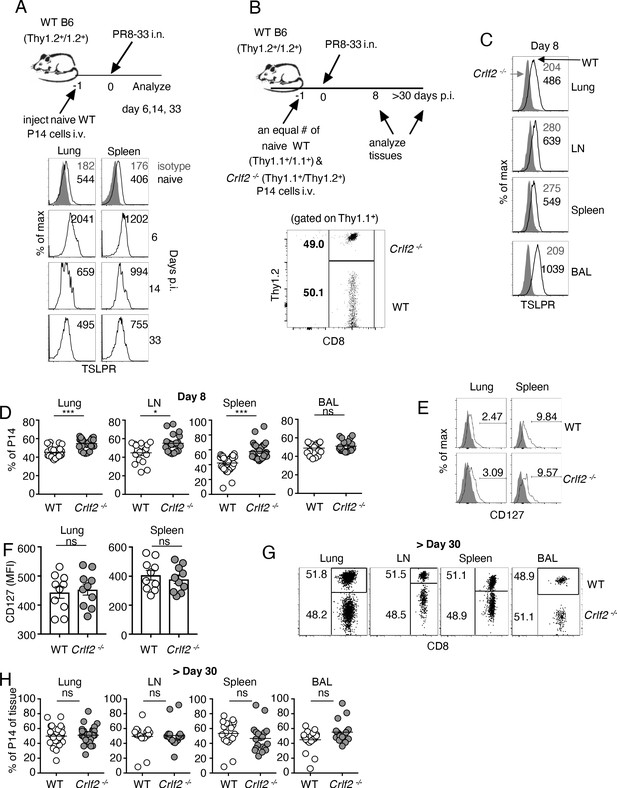
TSLP acts directly on CD8+ T cells during primary influenza infection.
(A) TSLPR expression on influenza-specific CD8+ T cells (P14 tg) during primary influenza infection. Top panel, experimental design. Bottom panel, flow cytometric analysis. Naïve cells were gated on CD44lo cells. (B–F) (B) Top panel, experimental design for C-H, where 2.5 × 104 of WT (Thy1.1+/1.1+) and Crlf2-/- (Thy1.1+/1.2+) P14 T cells were co-transferred into naïve WT C57BL/6 mice (Thy1.2+/1.2+), except in one experiment the markers were reversed, with the WT cells Thy1.1+/1.2+ and the Crlf2-/- P14 T cells were Thy1.1+/Thy1.1+ (see also Figure 1—figure supplement 1 and G). Bottom panel, Similar numbers of WT P14 cells and Crlf2-/- P14 cells were present. On the following day, the mice were infected intranasally (i.n.) with 103 EID50 of PR8-33. Mice were analyzed at Day 8 p.i. (C–F) or at a memory time point (>day 30 p.i.) (G and H). (C) TSLPR expression on influenza-specific P14 CD8+ T cells in the tissues on day 8 p.i. (D) Proportion of WT and Crlf2-/- T cells at day 8 p.i. in the tissues (shown are combined data from three independent experiments). (E and F) The expression of CD127 on WT and Crlf2-/- P14 cells in lungs and spleen. Shown are a representative flow cytometry plot (E) and summary of MFI data for CD127 expression (F). (n = 10). Data are mean ± SEM. (G and H) The proportion of WT and Crlf2-/- P14 cells of transferred cells in BAL, lungs, LN, and spleen at a memory time point, shown as a representative flow cytometry plot (G) and combined data from three independent experiments (H). ns = not significant; *p<0.05; ***p<0.005, using a two-tailed paired students t-test. Data shown are representative of at least two independent experiments.

Thy1.1/Thy1.1 versus Thy1.1/Thy1.2 genetic background differences do not explain the different number of WT versus Crlf2-/- P14 T cells after influenza infection.
2.5 × 104 WT and Crlf2-/- P14 T cells were co-transferred into naive WT C57BL/6 mice (Thy1.2+/1.2+), and on the following day the mice were infected intranasally (i.n.) with PR8-33. In (A), the WT P14 T cells were Thy1.1+/1.1+ and the Crlf2-/- cells were Thy1.1+/1.2+. In (B), the WT P14 T cells were Thy1.1+/1.2+ and the Crlf2-/- cells were Thy1.1+/1.1+. The proportion of WT and Crlf2-/- T cells at day 8 p.i. in the tissues are shown. Data are mean ± SEM. ns = not significant; *p<0.05; ***p<0.001; ****p<0.0001 using a two-tailed paired students t-test.
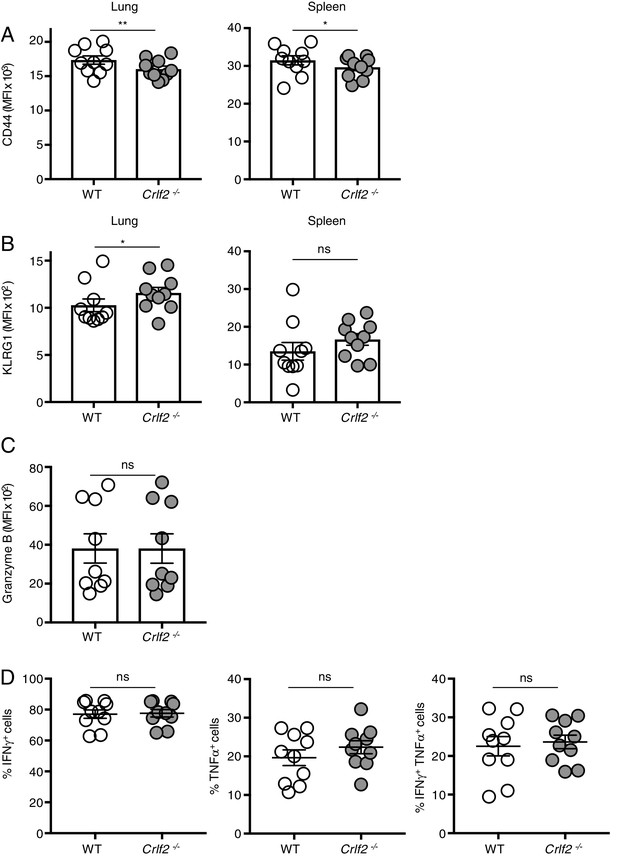
TSLP does not affect the transition to effector cells, cytokine secretion, or granzyme B levels during primary infection.
2.5 × 104 of WT (Thy1.1+/1.1+) and Crlf2-/- (Thy1.1+/1.2+) P14 T cells were co-transferred into naïve WT C57BL/6 mice (Thy1.2+/1.2+) and on the following day the mice were infected intranasally (i.n.) with 103 EID50 of PR8-33. (A–C). Lung mononuclear cells and splenocytes were isolated at day 8 p.i. CD44 (A), KLRG1 (B), and Granzyme B (C) levels were then assessed by mean florescent intensity (MFI) by flow cytometry (gating on live Thy1.1+ CD8+ T cells and then either WT (Thy1.2-) or Crlf2-/- (Thy1.2+)). (n = 10). Data are mean ± SEM. (D) Cells were re-stimulated with gp33 peptide in the presence of monensin and brefeldin A for 5 hr and production of IFN-γ and TNF-α was assessed by flow cytometry. In (D), shown are the percentage of cells expressing IFN-γ, TNF-α, or both IFN-γ and TNF-α. (n = 10). Data are mean ± SEM. ns = not significant; *p<0.05; **p<0.01 using a two-tailed paired students t-test. Data shown are representative of at least two independent experiments.
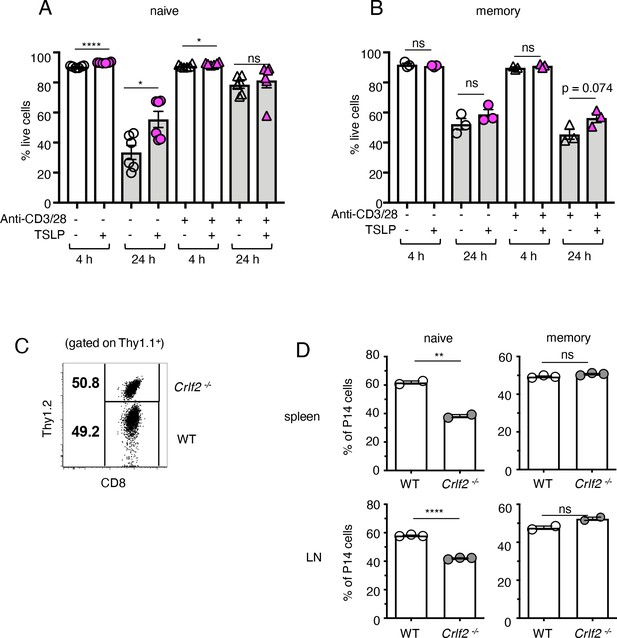
TSLP affects homeostasis of naïve but not memory CD8+ T cells.
(A and B) (A) Naïve P14 CD8+ T cells were purified and plated with either media or TSLP with or without plate-bound anti-CD3 and soluble anti-CD28. Cell viability was assessed by gating on CD8+ T cells and live/dead staining after 4 and 24 hr. (n = 6). Data are mean ± SEM. (B) Memory P14 CD8+ T cells were purified and plated with either media or TSLP with or without soluble anti-CD3 and soluble anti-CD28. Cell viability was assessed by gating on CD8+ T cells and live/dead staining after 4 and 24 hr. (n = 3). Data are mean ± SEM. (C) Naïve P14 CD8+ T cells were purified and equal numbers of WT (Thy1.1+/1.1+) and Crlf2-/- (Thy1.1+/1.2+) P14 CD8+ T cells were co-transferred into naïve WT C57BL/6 mice (Thy1.2+/1.2+). Shown are representative flow cytometry plots (gated on live Thy1.1+ CD8+ cells) of the combined naïve WT and Crlf2-/- P14 cells pre-transfer. (D) Percent of naïve and memory WT and Crlf2-/- cells of total transferred P14 T cells on day 9–11 post cell transfer. (n = 2 or 3). Data are mean ± SEM. For A, B and D: Data are mean ± SEM. ns = not significant; *p<0.05; **p<0.01; ****p<0.0001 using a two-tailed paired students t-test. Data shown are representative of at least two independent experiments.
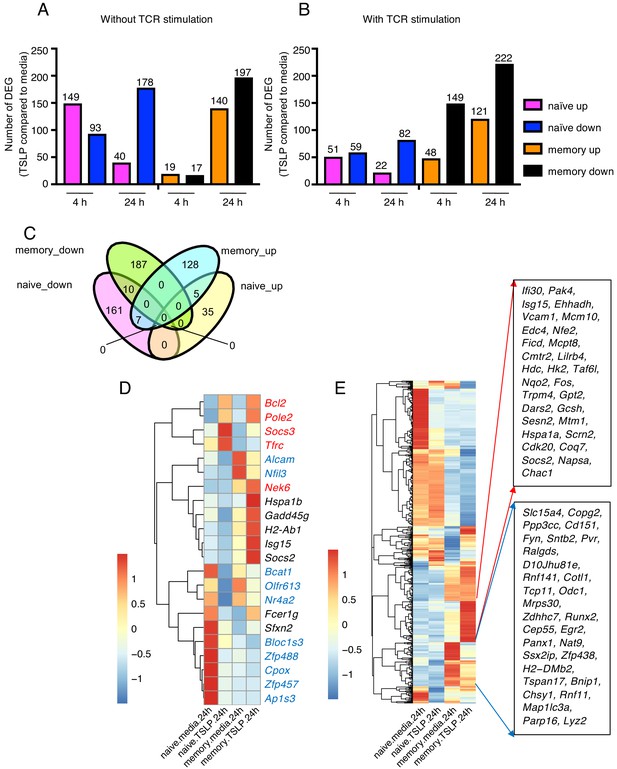
TSLP modulates gene expression on naïve and memory CD8+ T cells in vitro.
RNA-Seq performed on sorted naïve and memory CD8+ T cells after 4 or 24 hr incubation with medium or TSLP with or without TCR stimulation. Shown are the number of differentially expressed genes (FC >1.5, FDR < 0.05). (A and B) Number of genes affected by TSLP in CD8+ T cells not stimulated (A) or stimulated (B) with anti-CD3 + anti-CD28. (C) Venn diagram showing the number of genes whose expression was upregulated or downregulated by TSLP without TCR stimulation at 24 hr of incubation in naïve and memory CD8+ T cells. (D) Genes regulated by TSLP both in naïve and memory CD8+ T cells, color-coded as indicated in the text. (E) Genes differentially regulated by TSLP in naïve vs. memory cells. Highlighted are those that were selectively induced or repressed after stimulation with TSLP for 24 hr.

Gating strategy for sorting memory CD8+ T cells for in vitro stimulation.
Shown is forward versus side scatter (FSC versus SSC) to exclude debris (first plot), then FSC-A and FSC-H analysis as well as SSC-H and SCC-W were performed to exclude doublets (second and third plots). Then dead cells were eliminated (fourth plot). Transferred P14 cells were identified as CD8+ Thy1.1+ cells (fifth plot).
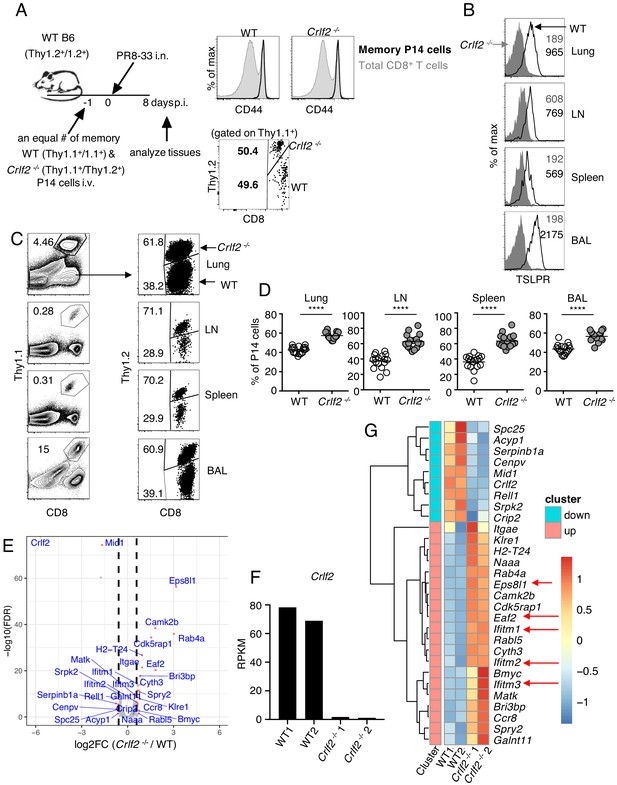
TSLP has a direct inhibitory effect on secondary CD8+ T-cell responses during influenza infection.
(A) Left panel, experimental design for (B–G), where P14 T cells were isolated from P14 WT (Thy1.1+/1.1+) and P14 Crlf2-/- (Thy1.1+/1.2+) mice and then separately injected into naïve WT mice. Each mouse was then infected with influenza PR8-33 i.n. for >30 days, and WT P14 cells and Crlf2-/- P14 cells were separately isolated. These cells were all CD44hi (right upper panel); 104 cells from each type of mouse were then co-transferred into recipient naïve WT C57BL/6 mice (Thy1.2+/1.2+) on day −1, and similar numbers of WT P14 cells and Crlf2-/- P14 cells were present (right lower panel). On the following day (day 0), each mouse was infected with 103 EID50 of PR8-33 i.n. (B–D) Mice were analyzed at day 8 p.i. (B) The expression of TSLPR on CD8+ T cells in BAL fluid, lungs, lymph node, and spleen. (C) Representative flow cytometry plots showing the percent of transferred P14 cells (gated on live lymphocytes) and proportion of WT and Crlf2-/- cells within the transferred population at day 8 p.i. with influenza in the tissues. (D) Percentage of P14 WT and Crlf2-/- T cells at day 8 p.i. with influenza in BAL fluid, lungs, LN, and spleen (combined data from three independent experiments are shown). (E–G) RNA-Seq was performed on cells from WT versus Crlf2-/- mice. (E) Differentially expressed genes are shown. The dashed lines correspond to log2(1.5)=0.585. (F) Expression of Crlf2 in cells from WT and Crlf2-/- mice. (G) Heatmap of differentially expressed genes from WT or Crlf2-/- CD8+T cells at day 8 p.i. with secondary influenza infection. Shown is the scale for fold induction or repression. Data are mean ± SEM. ns = not significant; ****p<0.0001 using a two-tailed paired students t-test. Data shown are representative of at least two independent experiments.
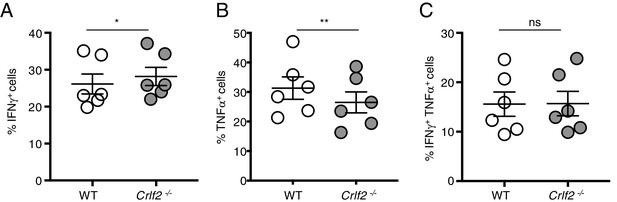
IFN-γ and TNF-α expression in WT versus Crlf2-/- mice during secondary infection.
Cells were re-stimulated with gp33 peptide in the presence of monensin and brefeldin A for 5 hr, and production of IFN-γ and TNF-α was assessed by flow cytometry. (A–C) Shown are the percentage of cells expressing IFN-γ (A), TNF-α (B), or both cytokines (C). (n = 6). Data are mean ± SEM. ns = not significant; *p<0.05; **p<0.01 using a two-tailed paired students t-test. Data shown are representative of at least two independent experiments.

Gating strategy for sorting WT and Crlf2-/-CD8+ T cells for RNA-sequencing.
Shown is forward versus side scatter (FSC versus SSC) to exclude debris (first plot), then FSC-A and FSC-H analysis was performed to exclude doublets (second plot). Then dead cells were eliminated (third plot). An empty channel was used to eliminate autofluorescent cells such as macrophages. Transferred P14 cells were identified as CD8+ Thy1.1+ cells (fourth plot) and then Crlf2-/- cells as Thy1.1+Thy1.2+ cells and WT cells as Thy1.1+Thy1.2- cells.
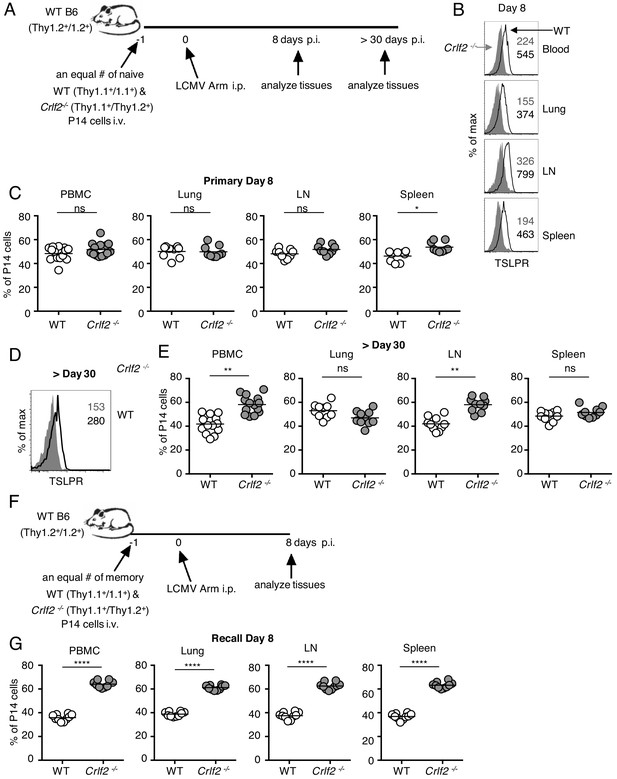
Direct actions of TSLP on virus-specific CD8+ T cells during primary and recall LCMV infection.
(A) Schematic of LCMV infection experiment. 103 of WT (Thy1.1+/1.1+) and Crlf2-/- (Thy1.1+/1.2+) P14 naïve T cells were co-transferred into naïve WT C57BL/6 mice (Thy1.2+/1.2+) and on the following day the mice were infected i.p. with 2 × 106 pfu of LCMV Armstrong. Mice were analyzed at day 8 p.i. (B and C) or at a memory time point (>day 30 p.i.) (D and E). (B) TSLPR expression was assessed on LCMV-specific P14 CD8+ T cells in the tissues on day 8 p.i. (C) Proportion of WT and Crlf2-/- T cells at day 8 p.i. in the tissues (combined data from two independent experiments shown). (D) TSLPR expression on memory P14 T cells from a mouse seeded with P14 T cells and infected with LCMV Armstrong i.p. for >30 days. (E) The proportion of WT and Crlf2-/- P14 cells of transferred cells in the tissues at >30 days, a memory time point (combined data from two independent experiments shown). (F) Schematic of LCMV infection experiment. WT (Thy1.1+/1.1+) and Crlf2-/- (Thy1.1+/1.2+) P14 T cells were isolated from mice seeded with P14 T cells and infected with LCMV Armstrong i.p. for >30 days, and equal numbers of WT and Crlf2-/- cells (103 of each population) were co-transferred into naïve WT C57BL/6 mice (Thy1.2+/1.2+) on day −1. On the following day (day 0), the mice were infected with 2 × 106 pfu LCMV Armstrong i.p. Mice were analyzed at day 8 p.i. (G) Proportion of WT and Crlf2-/- T cells at day 8 p.i. with LCMV in the tissues, (combined data from two independent experiments shown). Data are mean ± SEM. ns = not significant; *p<0.05; **p<0.01; ****p<0.0001 using a two-tailed paired students t-test. Data shown are representative of at least two independent experiments.
Additional files
-
Supplementary file 1
Differentially expressed genes from sorted naïve and memory CD8+ T cells 4 or 24 hr after incubation with medium or TSLP, without TCR stimulation.
Related to Figure 3A.
- https://cdn.elifesciences.org/articles/61912/elife-61912-supp1-v1.xlsx
-
Supplementary file 2
Differentially expressed genes from sorted naïve and memory CD8+ T cells 4 or 24 hr after incubation with medium or TSLP, with TCR stimulation.
Related to Figure 3B.
- https://cdn.elifesciences.org/articles/61912/elife-61912-supp2-v1.xlsx
-
Supplementary file 3
RNA-Seq comparison between naïve and memory CD8+ T cells 24 hr after incubation with TSLP, without TCR stimulation.
Related to Figure 3C–E.
- https://cdn.elifesciences.org/articles/61912/elife-61912-supp3-v1.xlsx
-
Supplementary file 4
Differentially expressed genes of WT versus Crlf2-/- mice, from CD8+ T cell recall responses at day eight after secondary infection.
Related to Figure 4E–G.
- https://cdn.elifesciences.org/articles/61912/elife-61912-supp4-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/61912/elife-61912-transrepform-v1.pdf