The origins and consequences of UPF1 variants in pancreatic adenosquamous carcinoma
Figures
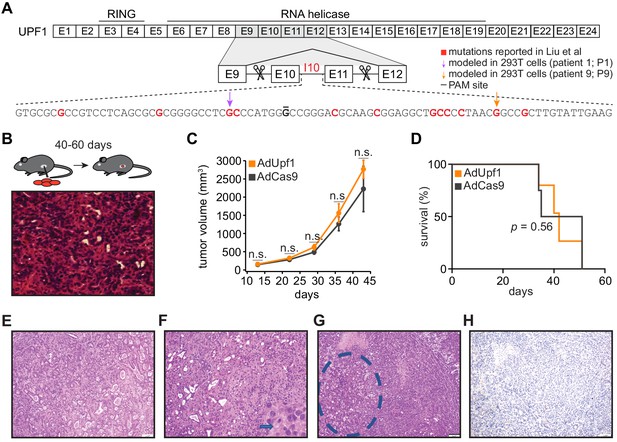
UPF1 mutations do not result in the acquisition of squamous histological features or confer a growth advantage to mutant cells in vivo.
(A) Schematic of UPF1 gene structure and corresponding encoded protein domains. Intron 10 (I10) contains the bulk of the mutations reported by Liu et al. Scissors indicate the sites targeted by the paired guide RNAs used to excise exons 10 and 11 (E10 and E11). Red nucleotides represent positions subject to point mutations reported in Liu et al. Arrows indicate specific mutations that we modeled in 293 T cells. The horizontal black line indicates the nucleotide within the protospacer adjacent motif (PAM) site that we mutated to prevent repeated cutting by Cas9 in 293 T cells. (B) Top, experimental strategy for testing whether mimicking UPF1 mis-splicing by deleting exons 10 and 11 promoted pancreatic cancer growth. Mice were orthotopically injected with mouse pancreatic cancer cells (KPC cells: KrasG12D; Trp53R172H/null; Pdx1-Cre) lacking Upf1 exons 10 and 11. Bottom, hematoxylin and eosin (H and E) stain of pancreatic tumor tissue harvested from the mice. (C) Line graph comparing tumor volume between mice injected with control (AdCas9; Cas9 only) or treatment (AdUpf1; Cas9 with Upf1-targeting guide RNAs) KPC cells. Tumor volume measured by ultrasound imaging. Error bars, standard deviation computed over surviving animals (n = 10 at first time point). n.s., not significant (p>0.05). p-values at each timepoint were calculated relative to the control group with an unpaired, two-tailed t-test. (D) Survival curves for the control (AdCas9) or treatment (AdUpf1) cohorts. Error bars, standard deviation computed over biological replicates (n = 10, each group). p-value was calculated relative to the control group by a logrank test. (E) Representative hematoxylin and eosin (H and E) staining of a pancreatic tumor resulting from orthotopic injection of control KPC cells displaying features of a moderately to poorly differentiated pancreatic ductal adenocarcinoma. Tumors were composed of medium-size duct-like structures and small tubular glands with lower mucin production. (F) Representative H and E image illustrating a moderately to poorly differentiated pancreatic ductal adenocarcinoma resulting from orthotopic injection of Upf1-targeted KPC cells. Depicted here is a section of the poorly differentiated component (arrow), which was characterized by solid sheets of tumor cells with large eosinophilic cytoplasms and marked nuclear polymorphism. (G) Representative H and E image of a pancreatic tumor resulting from orthotopic injection of Upf1-targeted KPC cells. The dashed circle marks a moderately differentiated component; the remainder is poorly differentiated. (H) Representative IHC image of a pancreatic tumor resulting from orthotopic injection of Upf1-targeted KPC cells for the squamous marker p40 (ΔNp63). No expression of the marker was observed in tumor cells.
-
Figure 1—source data 1
Source data for mouse tumor volume (Figure 1C).
- https://cdn.elifesciences.org/articles/62209/elife-62209-fig1-data1-v2.xlsx
-
Figure 1—source data 2
Source data for mouse survival (Figure 1D).
- https://cdn.elifesciences.org/articles/62209/elife-62209-fig1-data2-v2.xlsx
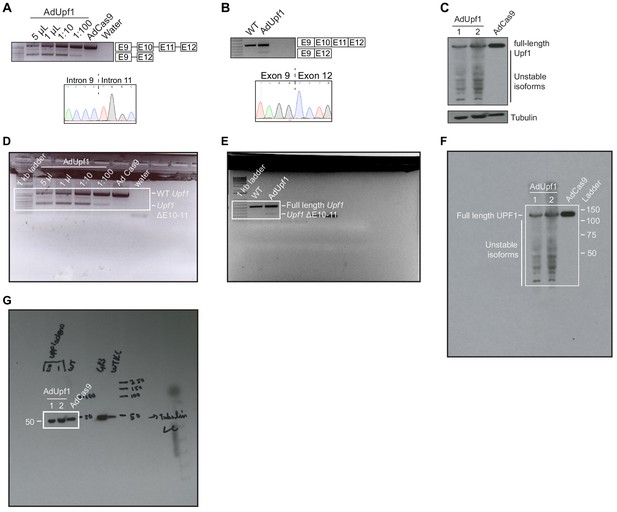
Validation experiments in mouse KPC cells.
(A) Titration of adenovirus expressing Cas9 only (AdCas9) or Cas9 and guide RNAs targeting intron 9 and intron 11 of mouse Upf1 (AdUpf1) in KPC cells. Gel corresponds to PCR using genomic DNA with primers in exons 9 and 12 (E9, E12). The lower band, corresponding to excision of exons 10 and 11, was only observed in AdUpf1 conditions. This band was excised and sequence-verified by Sanger sequencing (sequence trace displayed below). (B) As (A), but assaying a cDNA library instead of genomic DNA using RT-PCR. The lower band, corresponding to spliced mRNA lacking exons 10 and 11, was only observed in AdUpf1 conditions. (C) Immunoblot with a probe against UPF1. Levels of full-length UPF1 protein are lower in AdUpf1 conditions, as expected. 1 and 2 indicate biological replicates. Tubulin serves as a loading control. (D) Raw gel image for the genomic DNA PCR shown in (A). The cropped region is boxed in white. (E) Raw gel image for the RT-PCR shown in (B). The cropped region is boxed in white. (F) Raw image of the western blot shown in (C). The cropped region is boxed in white. (G) Raw image of the western blot for the mouse tubulin loading control shown in (C). The cropped region is boxed in white.
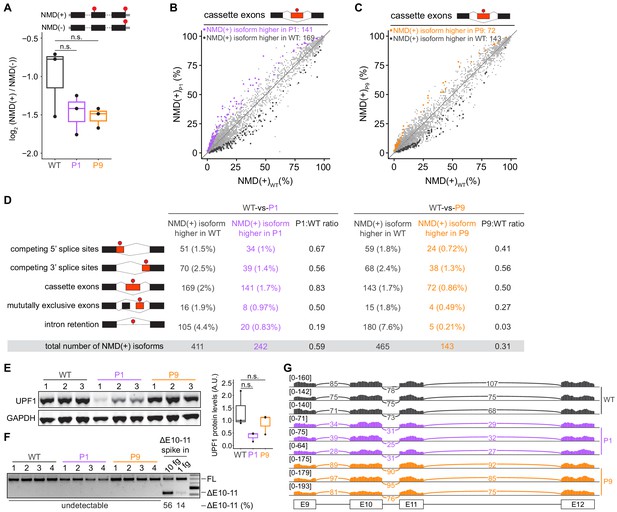
Mutations in UPF1 intron 10 do not inhibit nonsense-mediated mRNA decay (NMD) or cause exon skipping.
(A) Box plot of NMD efficiency in 293 T cells engineered to contain wild-type (WT) or mutant (P1, P9) UPF1. P1 and P9 correspond to the IVS10+31G>A and IVS10-17G>A mutations reported by Liu et al. All cells have the protospacer adjacent motif (PAM) site mutation illustrated in Figure 1A. NMD efficiency estimated via the beta-globin reporter assay11. Middle line, notches, and whiskers indicate median, first and third quartiles, and range of data. Each point corresponds to a single biological replicate. n.s., not significant (p>0.05). p-values were calculated for each variant relative to the control by a two-sided Mann–Whitney U test (p=0.40 for P1, 0.30 for P9). (B) Scatter plot showing transcriptome-wide quantification of transcripts containing NMD-promoting features in 293 T cells carrying the UPF1 mutation that was reportedly observed in patient 1 relative to control, wild-type cells. Each point corresponds to a single isoform that is a predicted NMD substrate (NMD(+)). Purple points represent NMD substrates that are significantly increased in UPF1-mutant cells relative to wild-type cells; black points represent NMD substrates that exhibit the opposite behavior. Plot is restricted to NMD substrates arising from differential inclusion of cassette exons. Significantly increased/decreased NMD substrates were defined as transcripts that displayed either an absolute increase/decrease in isoform ratio of ≥10% or an absolute log fold-change in expression of ≥2 with associated p≤0.05 (two-sided t-test). (C) As (B), but for 293 T cells carrying the UPF1 mutation that was reportedly observed in patient 9. Gold points represent NMD substrates that are significantly increased in UPF1-mutant cells relative to wild-type cells. (D) Summary of the numbers of NMD substrates arising from differential alternative splicing that exhibit significantly higher or lower levels in UPF1-mutant cells relative to wild-type cells. Analysis is identical to (B) and (C), but extended to the illustrated different types of alternative splicing. (E) Left, immunoblot of full-length UPF1 protein for the 293 T cell lines. Each lane represents a single biological replicate with the indicated genotype. GAPDH serves as a loading control. Equal amounts of protein were loaded in each lane (measured by fluorescence). Right, box plot illustrating UPF1 protein levels relative to GAPDH for each genotype. Middle line, notches, and whiskers indicate median, first and third quartiles, and range of data. Each point corresponds to a single biological replicate. Data was quantified with Fiji (v2.0.0). A.U., arbitrary units. n.s., not significant (p>0.05). p-values were calculated for each variant relative to the control by a two-sided Mann–Whitney U test (p=0.10 for P1, 1.0 for P9). (F) PCR using primers that amplify both full-length UPF1 mRNA (FL) and mRNA lacking exons 10 and 11 (ΔE10-11). UPF1 mRNA lacking exons 10 and 11 was only detected in the positive control lanes (ΔE10-11 spike in), in which DNA corresponding to UPF1 cDNA lacking exons 10 and 11 was synthesized and added to cDNA libraries created from WT cells prior to PCR. Numbers above each lane indicate biological replicates. Numbers below each lane represent the abundance of the lower band as a percentage of total intensity (see Materials and methods). Data was quantified with Fiji (v2.0.0). (G) RNA-seq read coverage across the genomic locus containing UPF1 exons 9–12 in the indicated 293 T cell lines. Each sample corresponds to a distinct biological replicate. Numbers represent read counts that supported each indicated splice junction (Katz et al., 2015).
-
Figure 2—source data 1
Source data for qPCR in HEK 293 T cell lines (Figure 2A).
- https://cdn.elifesciences.org/articles/62209/elife-62209-fig2-data1-v2.xlsx
-
Figure 2—source data 2
Source data for western blot in HEK 293 T cell lines (Figure 2E).
- https://cdn.elifesciences.org/articles/62209/elife-62209-fig2-data2-v2.xlsx
-
Figure 2—source data 3
Source data for RT-PCR in HEK 293 T cell lines (Figure 2F).
- https://cdn.elifesciences.org/articles/62209/elife-62209-fig2-data3-v2.xlsx
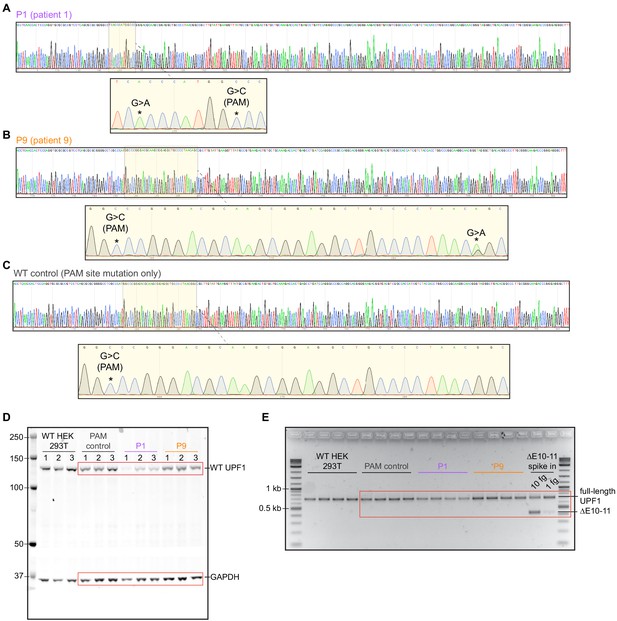
Validation experiments in human 293 T cells and raw gel images for western blot and RT-PCR.
(A) Sanger sequencing of genomic DNA from engineered 293 T cells verifying introduction of the mutation IVS10+31G>A in a homozygous state, as reported by Liu et al. for patient P1, as well as a single protospacer adjacent motif (PAM) site mutation. (B) Sanger sequencing of genomic DNA from engineered 293 T cells verifying introduction of the IVS10-17G>A mutation in a heterozygous state, as reported by Liu et al. for patient P9, as well as a single PAM site mutation. (C) Sanger sequencing of genomic DNA from engineered 293 T cells verifying introduction of a single PAM site mutation as a wild-type control. (D) Raw image of the western blot from Figure 2E. The cropped regions for UPF1 and GAPDH are boxed in red. (E) Raw image of the RT-PCR gel from Figure 2F. The cropped region for full-length UPF1 is boxed in red.
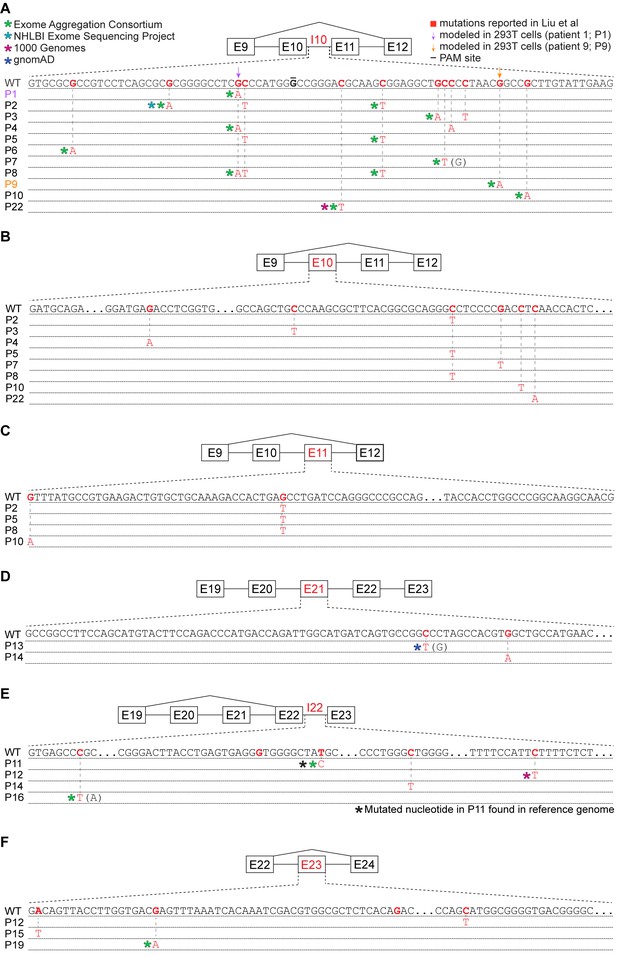
Many reported UPF1 mutations are identical to genetic variants.
(A) Illustration of the mutations in UPF1 intron 10 (I10) reported by Liu et al. Each row indicates the wild-type (WT) sequence from the reference human genome or mutations reported by Liu et al. (P1, patient 1). Purple and gold arrows indicate the mutations that we modeled with genome engineering in 293 T cells for patient 1 and patient 9, respectively. Red nucleotides represent positions subject to point mutations reported in Liu et al. The horizontal black line indicates the nucleotide within the protospacer adjacent motif (PAM) site that we mutated to prevent repeated cutting by Cas9. Parentheses indicate where we found genetic variation at a reported mutation position that differed from the specific mutated nucleotide reported by Liu et al. (B) As (A), but for UPF1 exon 10 (E10). (C) As (A), but for UPF1 exon 11 (E11). (D) As (A), but for UPF1 exon 21 (E21). (E) As (A), but for UPF1 intron 22 (I22). (F) As (A), but for UPF1 exon 23 (E23).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (H. sapiens) | UPF1 | Ensembl | ENSG00000005007 | |
| Gene (M. musculus) | Upf1 | Ensembl | ENSMUSG00000058301.8 | |
| Strain, strain background (M. musculus) | Mouse | NCI Charles River | Charles River: C57BL/6 albino | Mice for pancreatic injections |
| Strain, strain background (E. coli) | NEB Stable Competent E. coli (High efficiency) | New England Biolabs | C3040 | Chemically competent |
| Genetic reagent (M. musculus) | AdUpf1 | This paper | N/A | Adenovirus expressing CRISPR gRNAs targeting mouse Upf1 introns 9-11 |
| Cell line (H. sapiens) | HEK 293T | ATCC | CRL-11268 | Human cell line to model patient mutations |
| Cell line (M. musculus) | KPC | Generated from the PDX-1-Cre; LSL-KrasG12D/+; LSL Trp53R172H/+ | N/A | Murine cell line to model Upf1 exon skipping mutations (Hingorani et al., 2005) Provided by Robert Vonderheide |
| Antibody | Anti-human UPF1 (rabbit monoclonal) | Abcam | Cat No. ab10936 | WB: (1:1000) |
| Antibody | Anti-human GAPDH (rabbit polyclonal) | Abcam | Cat No. ab9485 | WB: (1:1000) |
| Antibody | Anti-rabbit secondary antibody (goat monoclonal) | Abcam | Cat No. ab216777 | WB: (1:10000) IRDye 680RD |
| Antibody | Anti-mouse UPF1 (rabbit monoclonal) | Cell signaling technology | Cat No.9435 | WB: (1:1000) |
| Antibody | Anti-mouse tubulin (mouse monoclonal) | Sigma-Aldrich | Cat No. T9206 | WB: (1:2000) |
| Antibody | Anti-rabbit secondary antibody (from donkey) | Amersham | NA93V | WB: (1:5000) |
| Antibody | Anti-mouse secondary antibody (from sheep) | Amersham | NA931 | WB: (1:5000) |
| Antibody | Anti-mouse p40-ΔNp63 (rabbit polyclonal) | Abcam | Cat No. ab166857 | WB: (1:100) |
| Recombinant DNA reagent | pX459/Cas9 expression plasmid | Addgene | Cat No. 48139 | Ran et al., 2013 |
| Recombinant DNA reagent | phCMV-MUP | PMID:9671053 | Plasmid | Control for transfection efficiency of pmCMV-GI-Norm and pmCMV-GI-39Ter |
| Recombinant DNA reagent | pmCMV-Gl-Norm | PMID:9671053 | Plasmid | Transient transfection construct coding for full-length β-globin |
| Recombinant DNA reagent | pmCMV-Gl-39Ter | PMID:9671053 | Plasmid | Transient transfection construct coding for truncated β-globin with PTC at amino acid 39 |
| Sequence-based reagent | mUpf1_F | This paper | PCR primer | GGTGATGAGATTGCTATTGAGC |
| Sequence-based reagent | mUpf1_R | This paper | PCR primer | TGTTCCTGATCTGGTTGTGC |
| Sequence-based reagent | mUpf1-intron_9-gDNA_F | This paper | Guide DNA Oligo | CACCGTTGTGAGGGCCATACCCTTG |
| Sequence-based reagent | mUpf1-intron_9-gDNA_R | This paper | Guide DNA Oligo | AAACCAAGGGTATGGCCCTCACAAC |
| Sequence-based reagent | mUpf1-intron_11-gDNA_F | This paper | Guide DNA Oligo | CACCGCCGTTGAGCTGATGGTGGCT |
| Sequence-based reagent | mUpf1-intron_11-gDNA_R | This paper | Guide DNA Oligo | AAACAGCCACCATCAGCTCAACGGC |
| Sequence-based reagent | hUPF1_F | This paper | Genomic DNA PCR primer | AAAACGTTTGCCGTGGATGAG |
| Sequence-based reagent | hUPF1_R | This paper | Genomic DNA PCR primer | CACATAGAGAGCGGTAGGCA |
| Sequence-based reagent | hUPF1-gDNA_F | This paper | Guide DNA oligo | GCGCGCGGGGCCTCGCCCAT |
| Sequence-based reagent | hUPF1-patient_1-HDR_R | This paper | DNA HDR ultramer | GCTCAGTGGTCTTTGCAGCACAGTCTTCACGGCATAAACCTTCAATACAAGCGGCCGTTAGGGGCAGCCTCCGCTTGCGTCCCGGGCCATGGGTGAGGCCCCGCGCGCTGAGGACGGCGCGCACCTG |
| Sequence-based reagent | hUPF1-patient_9-HDR_R | This paper | DNA HDR ultramer | GCTCAGTGGTCTTTGCAGCACAGTCTTCACGGCATAAACCTTCAATACAAGCGGCTGTTAGGGGCAGCCTCCGCTTGCGTCCCGGGCCATGGGCGAGGCCCCGCGCGCTGAGGACGGCGCGCACCTG |
| Sequence-based reagent | hUPF1-PAM-control-HDR_R (wild type) | This paper | DNA HDR ultramer | GCTCAGTGGTCTTTGCAGCACAGTCTTCACGGCATAAACCTTCAATACAAGCGGCCGTTAGGGGCAGCCTCCGCTTGCGTCCCGGGCCATGGGCGAGGCCCCGCGCGCTGAGGACGGCGCGCACCTG |
| Sequence-based reagent | hUPF1-RT-PCR_F | This paper | RT-PCR primer | GGATGAGATATGCCTGCGGT |
| Sequence-based reagent | hUPF1-RT-PCR_R | This paper | RT-PCR primer | TTCTCGTCGGCAGACGACAG |
| Sequence-based reagent | Positive control gBlock to detect UPF1 splice variant (ΔE10-11) | This paper | DNA gBlock | ACATGCGGCTCATGCAGGGGGATGAGATATGCCTGCGGTACAAAGGGGACCTTGCGCCCCTGTGGAAAGGGATCGGCCACGTCATCAAGGTCCCTGATAATTATGGCGATGAGATCGCCATTGAGCTGCGGAGCAGCGTGGGTGCACCTGTGGAGGTGACTCACAACTTCCAGGTGGATTTTGTGTGGAAGTCGACCTCCTTTGACAGGCCGGTGCTGGTGTGTGCTCCGAGCAACATCGCCGTGGACCAGCTAACGGAGAAGATCCACCAGACGGGGCTAAAGGTCGTGCGCCTCTGCGCCAAGAGCCGTGAGGCCATCGACTCCCCGGTGTCTTTTCTGGCCCTGCACAACCAGATCAGGAACATGGACAGCATGCCTGAGCTGCAGAAGCTGCAGCAGCTGAAAGACGAGACTGGGGAGCTGTCGTCTGCCGACGAGAAGCGGTACCGGGCCTTGAAGCGCACCGCAGAGAGAGAGCTGCTGATG |
| Sequence-based reagent | mMup1-qPCR_F | PMID:25564732 | qPCR primer | GACCTATCCAATGCCAATCG (exon 5/6 junction) |
| Sequence-based reagent | mMup1-qPCR_R | PMID:25564732 | qPCR primer | GATGATGGTGGAGTCCTGGT (exon 7) |
| Sequence-based reagent | hβ-globin-qPCR_F | PMID:25564732 | qPCR primer | GCTCGGTGCCTTTAGTGATG (exon 2) |
| Sequence-based reagent | mβ-globin-qPCR_R | PMID:25564732 | qPCR primer | CCCAGCACAATCACGATCATA (exon 3, mouse specific) |
| Commercial assay or kit | RNeasy Plus Mini Kit | Qiagen | Cat No. 79654 | |
| Commercial assay or kit | Zero Blunt TOPO PCR cloning system | ThermoFisher | Cat No. K280020 | |
| Chemical compound, drug | Phosphatase inhibitor | ThermoFisher | Cat No. A32959 | |
| Chemical compound, drug | Protease inhibitor | Thermofisher | Cat No. A32963 | |
| Chemical compound, drug | Penicillin/Streptomycin | GIBCO | Cat No. 15070063 | |
| Chemical compound, drug | Lipofectamine 2000 | ThermoFisher | Cat No. 11668030 | |
| Chemical compound, drug | Lipofectamine 3000 | Invitrogen | Cat No. L3000001 | |
| Chemical compound, drug | Puromycin | ThermoFisher | Cat No. A1113803 | |
| Software, algorithm | GuideScan v1.0 | PMID:28263296 | http://www.guidescan.com/ | |
| Software, algorithm | Fiji v2.0.0 | ImageJ | https://imagej.net/Fiji | |
| Software, algorithm | RSEM v1.2.4 | PMID:21816040 | deweylab.github.io/RSEM/RRID:SCR_013027 | |
| Software, algorithm | Bowtie v1.0.0 | PMID:19261174 | github.com/BenLangmead/bowtie/; RRID:SCR_005476 | |
| Software, algorithm | TopHat v2.0.8b | PMID:19289445 | ccb.jhu.edu/software/tophat/index.shtml RRID:SCR_013035 | |
| Software, algorithm | MISO v2.0 | PMID:21057496 | genes.mit.edu/burgelab/miso/ RRID:SCR_003124 | |
| Software, algorithm | IGV v2.3.90 | Thorvaldsdottir | software.broadinstitute.org/software/igv/ RRID:SCR_011793 | |
| Software, algorithm | Prism v7.0 | GraphPad Prism v7.0 | http://www.graphpad.com/ RRID:SCR_002798 | |
| Other | SuperScript IV Reverse Transcriptase | ThermoFisher | Cat No. 18090010 | |
| Other | Q5 High-Fidelity DNA Polymerase | New England Biolabs | Cat No. NEB #M0491 | |
| Other | PowerUp SYBER Green Master Mix | ThermoFisher | Cat No. A25742 | |
| Other | RNase-Free DNase Set | Qiagen | Cat No. 79254 |
Additional files
-
Supplementary file 1
(a) Histopathological classification of pancreatic tumors derived from orthotopic injection of control or Upf1-targeted KPC cells. Tumors displaying >25% squamous differentiation were classified as positive for squamous features. (b) Differentially spliced isoforms identified in 293 T cells engineered to have the ‘patient 1’ UPF1 genotype relative to WT control cells. (c) Differentially spliced isoforms identified in 293 T cells engineered to have the ‘patient 9’ UPF1 genotype relative to WT control cells. (d) Overlap between UPF1 mutations reported by Liu et al. and genetic variants present in the 1000 Genomes, NHLBI Exome Sequencing Project, ExAC, and gnomAD databases. Genomic coordinates are specified with respect to the GRCh37/hg19 genome assembly. (e) Summary statistics of somatic mutations in UPF1 identified in our reanalysis of whole-exome and whole-genome sequencing data from Fang et al. (f) All genetic differences in UPF1 and KRAS from the reference genome, including both somatic mutations and inherited genetic variants, that we identified in our reanalysis of all samples from Fang et al.’s cohorts.
- https://cdn.elifesciences.org/articles/62209/elife-62209-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/62209/elife-62209-transrepform-v2.docx





