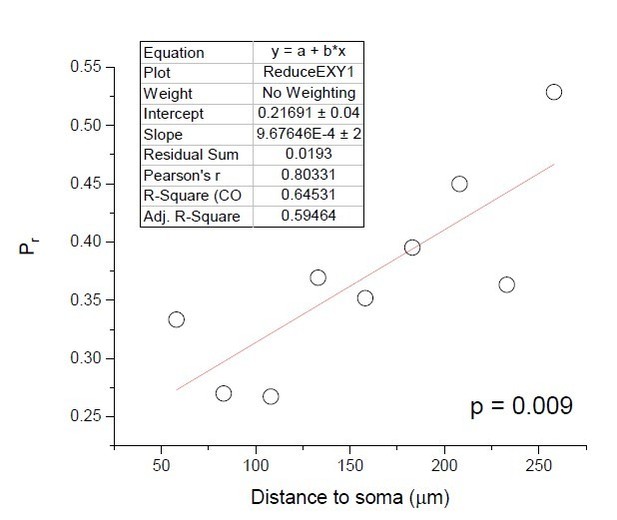
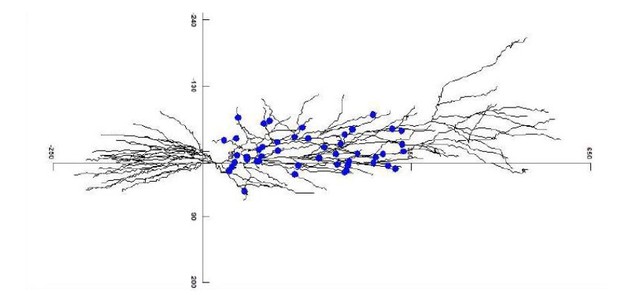
Release probability increases towards distal dendrites boosting high-frequency signal transfer in the rodent hippocampus
Figures
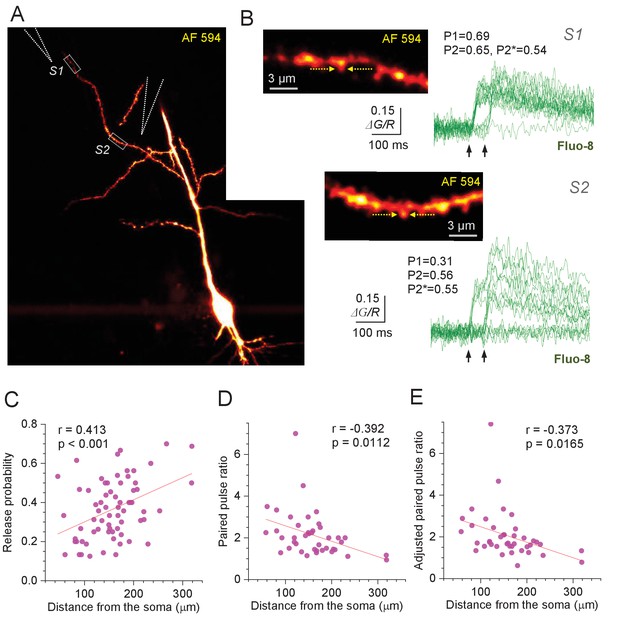
Optical quantal analysis at individual CA3-CA1 synapses reports higher release probability towards distant dendrites.
(A) CA1 pyramidal cell held in whole-cell mode (acute hippocampal slice), dialysed with 50 µM AF 594 and 300 µM Fluo-8 (75 µm z-stack average, λx2p = 800 nm; AF 594 channel). Dotted rectangles (S1 and S2), two ROIs to record from two dendritic spines; stimulating electrode positions are illustrated by dotted cones. (B) Image panels, ROIs as in (A), shown at higher magnification; arrows, linescan positioning at the dendritic spines of interest. Traces, Ca2+ signal (ΔG/R, Fluo-8 green-channel signal ΔG related to red-channel AF 594 signal R) recorded as the width-integrated linescan intensity, at two spines as indicated, in response to paired-pulse afferent stimuli (arrows). Release failures and responses to the first pulse can be clearly separated (Figure 1—figure supplement 1D). P1 and P2 show average release probability in response to the 1st and 2nd stimulus, respectively, calculated as P1 = {(1,0) + (1,1)} / N and P2 = {(0,1) + (1,1)} / N, and P2* is the adjusted 2nd-stimulus release probability calculated as P2* = (0,1) / {(0,0) + (0,1)} where brackets indicate the counts of paired-pulse successes (1) or failures (0), and N is the number of trials. (C) Average release probability (Pr; shown as P1 in B) at individual synapses plotted against distance to the soma. Solid line, linear regression (p value to reject H0 = zero slope and Pearson's r shown; n = 67). (D) Paired-pulse ratio P2/P1 plotted against distance to the soma. Other notations as in (C) (n = 41; average PPR mean ± SEM: 2.14 ± 0.17; spines with no reliable detection of 2nd responses, and no detectable (0,1) responses, were excluded). (E) Adjusted paired-pulse ratio PPR*=P2*/P1 plotted against distance to the soma. Other notations as in (C) (n = 41; average PPR*=2.05 ± 0.18).
-
Figure 1—source data 1
Original data readout for Figure 1C-E and Figure 1—figure supplement 1D.
- https://cdn.elifesciences.org/articles/62588/elife-62588-fig1-data1-v3.xlsx
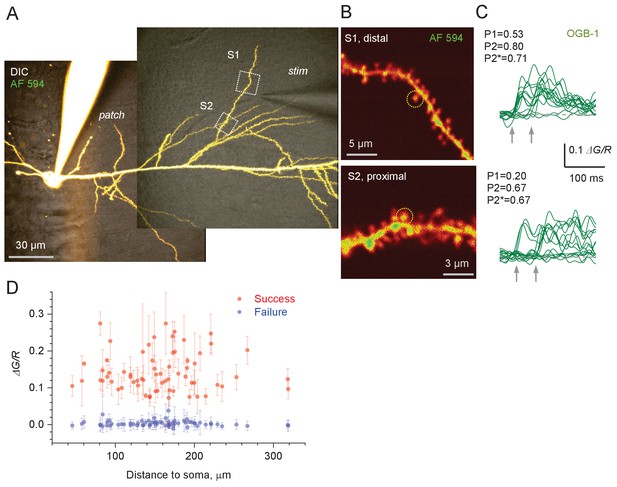
Optical quantal analysis at individual CA3-CA1 synapses in acute hippocampal slices: second-order dendrite.
(A) CA1 pyramidal cell held in whole-cell mode dialysed with 50 µM AF 594 and 400 µM OGB-1 (75 µm z-stack average, λx2p = 800 nm; DIC + AF 594 channel combined). Patch pipette (patch) and stimulating electrode (stim) can be seen; dotted rectangles (S1 and S2), ROIs to record from individual dendritic spines. (B) ROIs as shown in (A) by dotted, at higher magnification (AF 594 channel only); dotted circles, two dendritic spines of interest. (C) Examples of Ca2+ linescan signal traces (ΔG/R, green-channel OGB-1 increment signal ΔG related to red-channel AF 594 signal R) recorded in two dendritic spines as shown in (B), in response to two afferent stimuli (arrows) applied by a stimulating electrode (as in A). Release failures and successes can be clearly separated; notations P1, P2, and P2* as in Figure 1B. (D) Average amplitudes (10–45 ms interval post-pulse, mean ± SD) of the first-response successes (red) and failures (blue) for 67 dendritic spines as shown in Figure 1C.
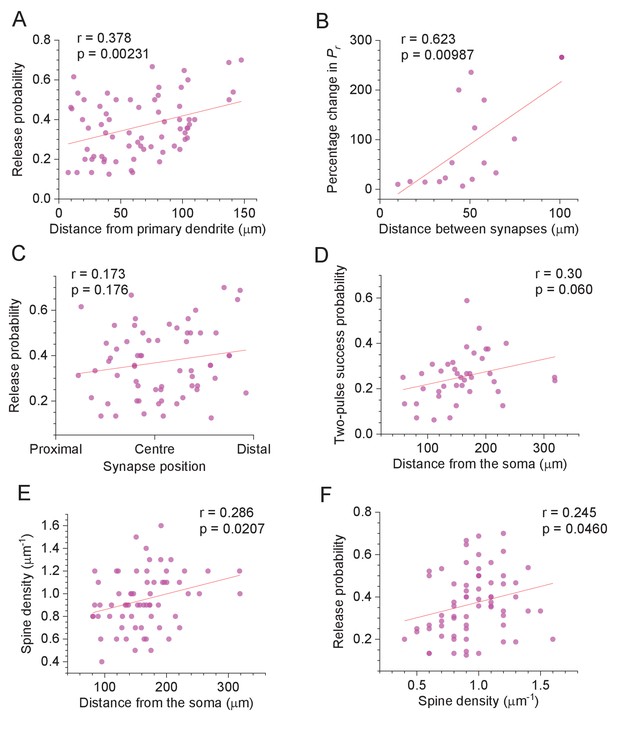
Selected features of excitatory synapses with respect to their dendritic location.
(A) Average release probability (Pr) at individual synapses plotted against distance to the first dendrite branching point. Solid line, linear regression (p value to reject H0 = zero slope and Pearson's r shown; n = 67). (B) Percentage difference in Pr between two synapses on one dendritic branch (as in Figure 1A, Figure 1—figure supplement 1A), plotted against the distance between them along the branch. Other notations as in (A) (n = 15). (C) Average release probability (Pr) plotted against relative synapse position at the dendritic branch: synapse co-ordinate was scaled to the 0–1 range representing the branch origin and the end, as indicated (n = 63; several spines with unidentifiable distal branch ends were excluded). (D) Probability of release success upon both afferent stimuli, plotted against distance from the soma. Other notations as in (A) (n = 40). (E) Apparent spine density along the dendrite (smallest/thinnest spines could be undetectable, see Discussion), plotted against distance from the soma. Other notations as in (A) (n = 65; several spines with unidentifiable local spine density were excluded). (F) Release probability (Pr), plotted against distance from the soma. Other notations as in (A) (n = 67).
-
Figure 2—source data 1
Original data readout for Figure 2A-F and Figure 2—figure supplement 1A-B.
- https://cdn.elifesciences.org/articles/62588/elife-62588-fig2-data1-v3.xlsx
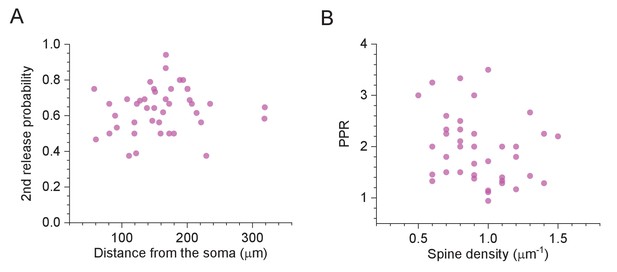
Selected features of excitatory synapses with respect to their dendritic location.
(A) Average probability of the second release (paired-pulse stimuli 50 ms apart) plotted against distance to the soma; sample mean ± SEM: 0.633 ± 0.20 (n = 41). (B) Paired-pulse ratio (as P2/P1 in C) plotted against spine density along dendrites; sample mean ± SEM: 2.14 ± 0.17 (n = 41).
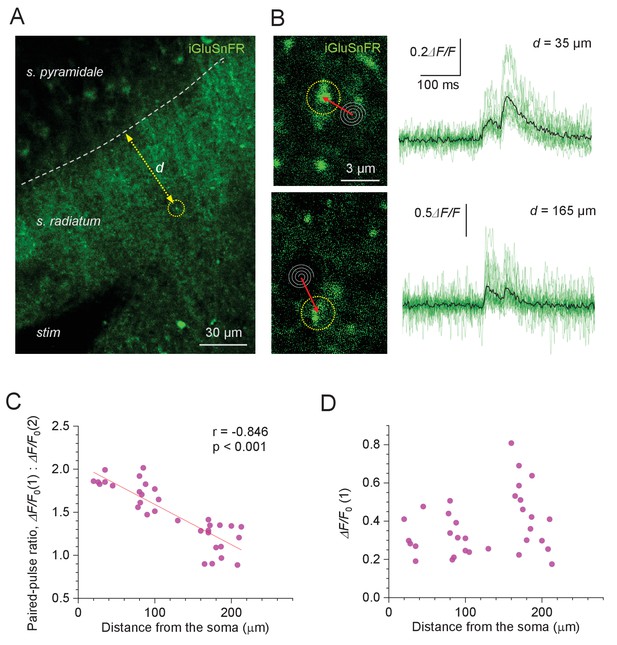
Evoked glutamate release from Schaffer collaterals shows lower paired-pulse ratios at greater distances from pyramidal cell bodies.
(A) Experimental design: area of the hippocampal slice with iGluSnFR expressed in neuronal membranes (green channel); arrow, measured distance d between CA1 pyramidal cell body layer and the axonal bouton of interest; stim, stimulating electrode. (B) Examples of recorded axonal boutons (image panels, dotted circles; position of spiral 'Tornado' linescans is illustrated) showing characteristic glutamate signals in response to two afferent stimuli 50 ms apart (green traces, individual trials; black, average), at two distances from the s. pyramidale, as indicated. The lack of release failures reflects detection of glutamate escaping from multiple neighbouring synapses. (C) Paired-pulse ratio for optical glutamate signals: ΔF/F0(1) / ΔF/F0(2) averaged over 18–36 trials at individual boutons, plotted against distance to the soma. Other notations as in Figure 1C (n = 33). (D) Amplitude of the first glutamate response, ΔF/F0(1) averaged over 18–36 trials at individual boutons, plotted against distance to the soma. The amplitude values reflect the average amount of glutamate released from the bouton of interest, and glutamate escaping from its neighbours; other notations as in (C) (n = 33).
-
Figure 3—source data 1
Original data readout for Figure 3C-D and Figure 3—figure supplement 1C.
- https://cdn.elifesciences.org/articles/62588/elife-62588-fig3-data1-v3.xlsx
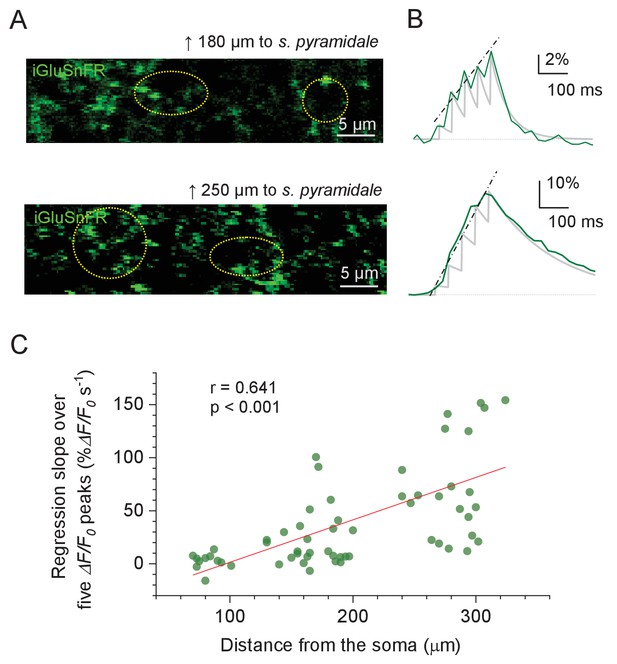
Optical (multi-synaptic) glutamate signal evoked by short bursts of Schaffer collateral stimulation at varied distances from s.pyramidale.
(A) Examples of recorded ROIs (iGluSnFR green fluorescence channel, dotted ovals) in s. radiatum, with groups of stimulation-responding tentative axonal boutons expressing iGluSnFR, at different distances to the s. pyramidale border (as illustrated in Figure 3A; distance reference shown is measured to the horizontal midline of the area shown; distance to each ROI was measured to the ROI centroid). (B) ROI-average iGluSnFR responses (green line, temporal resolution ~25 ms) to five stimuli 50 ms apart. The underlying sensor signal kinetics (light grey line) was reconstructed using the fitting algorithm (i = 1,..., 5) where Ai is the ith signal amplitude (fitted directly to the recoded amplitude), Δt = 50 ms, and the decay constant τ obtained directly from fitting the signal tail after the fifth pulse; dash-dotted lines, the linear regression line for the five ΔF/F0 response peaks (at the five pulse onsets). (C) Average slope of linear regression for the five ΔF/F0 response peaks as shown in (B), plotted against the distance from the s. pyramidale. (n = 61 boutons from N = 7 animals / slices).
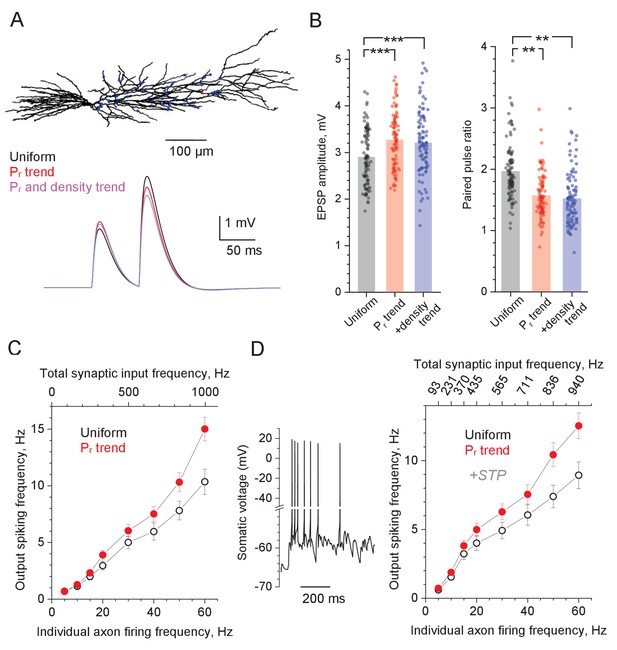
Computer simulations of CA1 pyramidal cell with stochastic excitatory synapses.
(A) Diagram, NEURON model of a reconstructed CA1 pyramidal cell (Migliore et al., 1999) (ModelDB 2796; variable time step dt, t = 34°C). 50 excitatory inputs (blue dots) generate bi-exponential conductance change (rise and decay time, 1 ms and 20 ms, respectively) stochastically, in accord with Pr. Traces, simulated somatic response to paired-pulse stimuli (50 ms apart), with Pr distributed either uniformly randomly (black), or in accord with the distance-dependent trend (as in Figure 1C; red), and both Pr and synaptic density trends (as in Figure 2E, blue; same average Pr = 0.36 ). (B) Summary of simulation tests in (A); dots, individual runs (n = 100); bars, mean EPSP amplitude (left, mean ± SEM: 2.90 ± 0.060, 3.27 ± 0.061, 3.20 ± 0.068, for the three conditions, respectively, as indicated) and paired-pulse ratios (right, mean ± SEM: 1.96 ± 0.047, 1.57 ± 0.036, 1.52 ± 0.0437, notation as above) are shown; ***p<0.005. (C) Input-output spiking rate relationship over the physiological range of input firing frequencies (per axon, bottom axis; total, top axis); hollow circles, uniform distribution of Pr; solid symbols, Pr follows the distance-dependent trend (as in Figure 1C); mean ± SEM are shown (n = 100 simulation runs). (D) Trace: A characteristic cell spiking burst (model as in A) in response to a Poisson-process afferent spiking input (~50 Hz per synapse) incorporating the experimental kinetics of short-term plasticity (STP) at CA3-CA1 synapses (Mukunda and Narayanan, 2017) (see Figure 4—figure supplement 1C–F for detail). Graph: Input-output spiking rate relationship across the physiological range of average input firing frequencies, with experimental STP incorporated; other notations as in (C); the top abscissa scale is nonlinear because STP affects average Pr in a biphasic, non-monotonous manner (see Figure 4—figure supplement 1C).
-
Figure 4—source data 1
Original data readout for Figure 4A–D and Figure 4—figure supplement 1A–F.
- https://cdn.elifesciences.org/articles/62588/elife-62588-fig4-data1-v3.xlsx
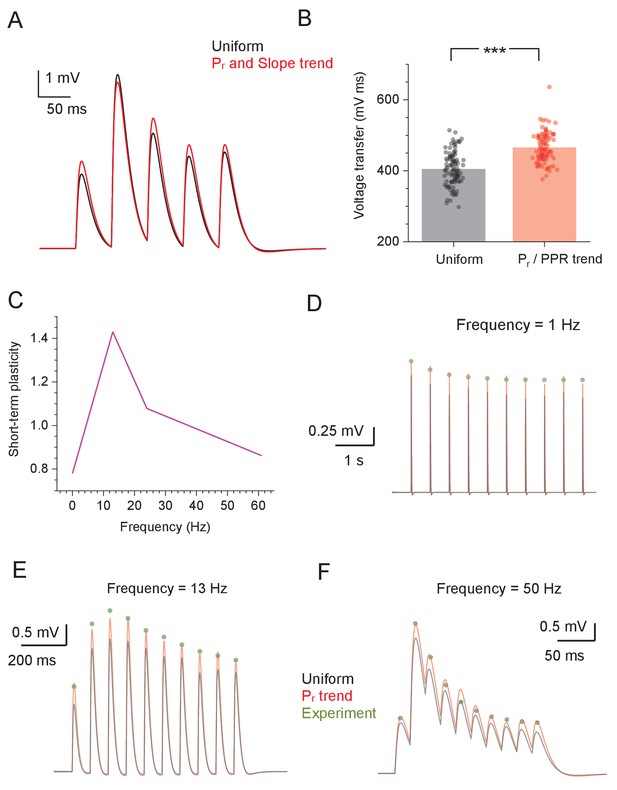
Simulating the experiment-based kinetics of short-term plasticity (STP) in the CA3-CA1 circuit.
(A) Traces show simulated average somatic response to five-burst 20 Hz stimuli (NEURON model as in Figure 4A), with synaptic Pr values distributed either uniformly randomly (black line), or in accord with the distance-dependent trend of Pr (as in Figure 1C–D) and the five-pulse ΔF/F0 slope (as in Figure 3—figure supplement 1C). (B) Summary of stochastic simulations shown in (A); ordinate, voltage transfer (area under the voltage curve over the five EPSPs); dots, individual runs (n = 100); bars, average values; ***p<0.001. (C) Linearised representation of the STP ratio profile for the 1–50 Hz range of presynaptic spiking frequencies replicating the experimental band-pass structure of the CA3–CA1 Schaffer collateral synapses (Mukunda and Narayanan, 2017). (D–F) Simulated EPSP response in a CA1 pyramidal cell (NEURON model as in Figure 4A) to a burst of afferent stimuli, with synaptic Pr values distributed either uniformly randomly (black), or in accord with the distance-dependent trend of Pr (as in Figure 1C; red); green dots, experimental data from Mukunda and Narayanan, 2017, normalised to the first EPSP amplitude, as indicated; postsynaptic response incorporates STP kinetics, as shown in (C).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Chemical compound, drug | Picrotoxin | Tocris | Cat. # 1128 | |
| Chemical compound, drug | D-Serine | Sigma Aldrich | Cat. # S4250 | |
| Chemical compound, drug | Oregon Green 488 BAPTA-1, Hexapotassium Salt | ThermoFisher | Cat. #O6806 | |
| Chemical compound, drug | Alexa Fluor-594 Hydrazide | ThermoFisher Scientific | Cat.# A10438 | |
| Chemical compound, drug | Fluo-8, potassium salt | Stratech Scientific | Cat.# 21087-AAT | Original Source: (Teflabs) No longer exists |
| Chemical compound, drug | Fluo-4, Pentapotassium Salt | ThermoFisher Scientific | Cat.# F14200 | |
| Chemical compound, drug | Agarose | Sigma-Aldrich | Cat. #9539; CAS: 9012-36-6 | Freshly prepared (4%) |
| Chemical compound, drug | D-glucose | Sigma-Aldrich | Cat. #G8270; CAS: 50-99-7 | Freshly prepared (10 mM) |
| Chemical compound, drug | KCl | Sigma-Aldrich | Cat. #P9333; CAS: 7447-40-7 | |
| Recombinant DNA reagent | AAV9.hSynap.iGluSnFr.WPRE.SV40 | Penn Vector Core | Addgene Cat. # 98929-AAV9 | |
| Strain, strain background (Sprague-Dawley rat) | Sprague-Dawley | Charles River UK | Crl: CD (SD) Strain: 0204 | |
| Strain, strain background (C57BL/6J mouse) | C57BL/6J | Charles River UK | C57BL/6NCrl Strain: 0159 | |
| Software, algorithm | ImageJ | NIH | RRID:SCR_003070 https://imagej.nih.gov/ij/ | |
| Software, algorithm | pClamp10 | Molecular Devices | RRID:SCR_011323 https://www.moleculardevices.com/products/axon-patch-clamp-system/acquisition-and-analysis-software/pclamp-software-suite | |
| Software, algorithm | OriginPro | OriginLab Inc | RRID:SCR_014212 https://www.originlab.com/origin | |
| Software, algorithm | MES 4.x-5.x | Femtonics Ltd. | RRID:SCR_018309 https://uk.mathworks.com/products/connections/product_detail/femtonics-mes.html | |
| Software, algorithm | WinWCP Versions 4.x-5.x | Strathclyde Electrophysiology Software | RRID:SCR_014270 http://spider.science.strath.ac.uk/sipbs/software_ses.htm | |
| Software, algorithm | Matlab | Mathworks | RRID:SCR_001622 https://uk.mathworks.com/products/matlab.html | |
| Software, algorithm | NEURON 7.6 × 64 | https://www.neuron.yale.edu/neuron/ | RRID:SCR_005393 | |
| Software, algorithm | Kinetics of CA1 pyramidal neuron | https://senselab.med.yale.edu/ModelDB/ShowModel?model=2796#tabs-2 | RRID:SCR_005393 model = 2796#tabs-2 | |
| Software, algorithm | reconstructed CA1 pyramidal neurons | https://senselab.med.yale.edu/ModelDB/ShowModel?model=7509#tabs-1 | RRID:SCR_005393 model = 7509#tabs-1 | |
| Other | Multiclamp 700B | Molecular Devices | RRID:SCR_018455 | |
| Other | Olympus FluoView1000 | Olympus | RRID:SCR_014215 | |
| Other | Femto3D RC | Femtonics | Femto3D RC | |
| Other | BioRad Radiance 2100 | BioRad | Radiance 2100 | |
| Other | Leica VT1200S vibratome | Leica Biosystems | RRID:SCR_018453 |