Individual differences in honey bee behavior enabled by plasticity in brain gene regulatory networks
Figures
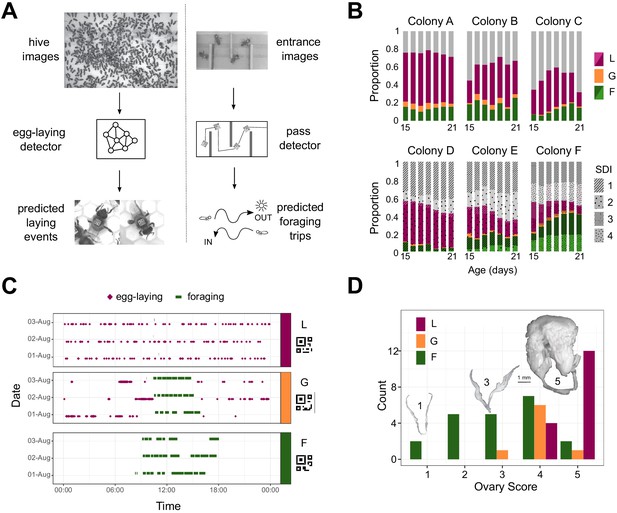
Automated monitoring of behavior in queenless colonies of laying worker honey bees.
(A) Automatic behavior monitoring was performed inside the hive and at the hive entrance to predict egg-laying and foraging events in six colonies (N = 800 bees per colony at the start of each trial). Hive images were captured 1/s for 24 h/day, and entrance images 2/s for 12 h/day beginning when adult bees were 15 days old. (B) Proportion of bees alive each day categorized as layers (purple), foragers (green), generalists (orange), or others (gray). For colonies A-C, individuals were from single source colonies headed by a naturally mated queen. For colonies D-F, individuals from two source colonies headed by queens each inseminated by semen from a single different drone (single drone inseminated, SDI) were mixed. Different source colonies are indicated by pattern and hue. (C) Ethograms for three individuals selected for sequencing (bCodes shown below group labels) across three days of tracking. (D) Distribution of ovary scores for individuals selected for sequencing. Insets are images from bees with ovary scores of 1, 3, and 5. L: layer, G: generalist, F: forager.
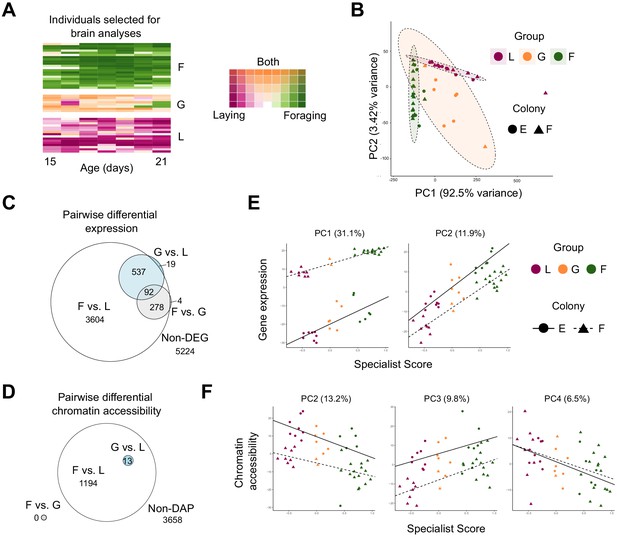
Patterns of brain gene expression and chromatin accessibility are associated with behavior.
(A) Daily rank-normalized behavior of individuals (rows) selected for brain RNAseq and ATACseq analysis converted to 2D colorspace from specialist and generalist scores. (B) Principal Component Analysis (PCA) of behavioral variation for individuals chosen for brain RNAseq and ATACseq analysis. Metrics included number of eggs laid, number of foraging events, proportion of foraging trips with evidence of nectar collection, proportion of trips with evidence of pollen collection, and proportion of trips with evidence of both nectar and pollen collection. (C) Euler diagram for overlaps of pairwise differentially expressed genes (DEGs) between behavioral groups. Note that one gene was overlapping between F vs. G and G vs. L (but not F vs. L) and is not represented in the diagram due to graphical constraints. (D) Euler diagram for overlaps of genes proximal to pairwise differentially accessible chromatin peaks (DAPs) between behavioral groups. (E) PCs from PCA of brain transcriptomic profiles regressed against specialist score (PC1: R2 = 0.947, p<0.0001; PC2: R2 = 0.838, p<0.001). (F) PCs from PCA of brain chromatin accessibility regressed against specialist score (PC2: R2 = 0.584, p<0.001; PC3: R2 = 0.543, p<0.0001; PC4: R2 = 0.187, p<0.0045; PC1: p>0.05). L: layer, G: generalist, F: forager.
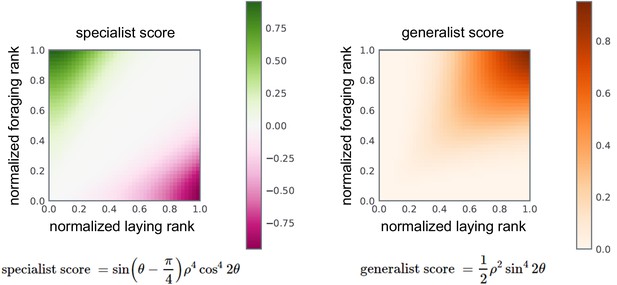
Formulae and color-space mapping for specialist (left) and generalist (right) behavioral scores.
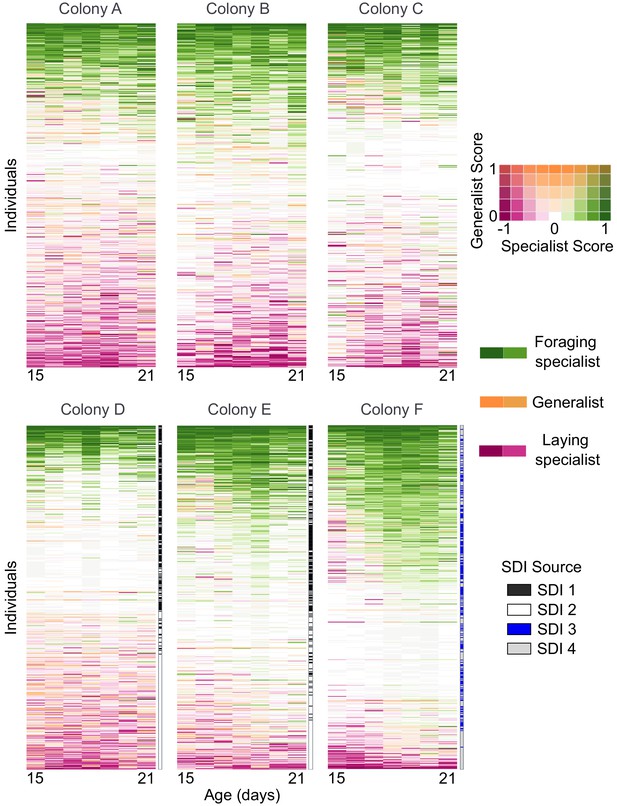
Daily behaviors of individual bees (rows) across time in each colony.
Colored rectangles indicate specialist and generalist scores represented in 2D color space as shown in legend. Individuals are sorted by median lifetime specialist score. Single-drone inseminated (SDI) queen source is shown to the right of each row for colonies D-F, where workers were known offspring of two SDI queens per colony. In colonies A-C workers were offspring of naturally mated queens.
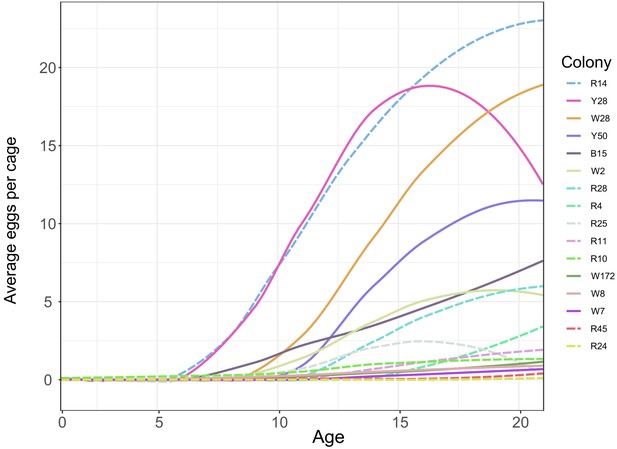
Smoothed average egg counts for laying workers in laboratory cages.
Solid lines indicate bees from source colonies headed by a naturally-mated queen, while dashed lines indicate bees from source colonies headed by queens instrumentally inseminated with semen from a single drone (SDI). Colonies used as sources for behavioral tracking included W28, W2, R25, and R45.
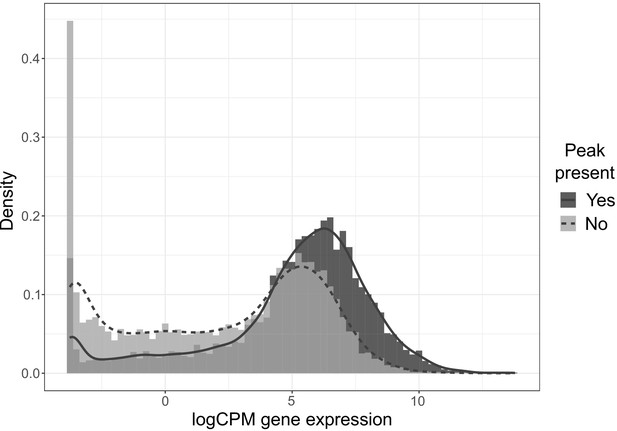
Histogram (bars) and density (lines) of normalized (logCPM) gene expression for genes with (dark gray) and without (light gray) nearby peaks of chromatin accessibility.
Distributions are significantly different (p<0.0001, Kolmogorov-Smirnov test).
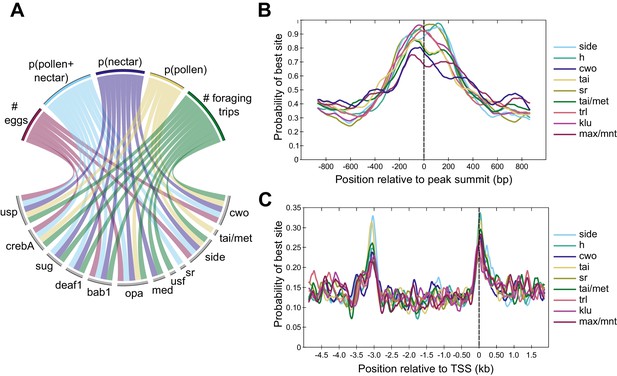
Differences in TF activity and TF motif occurrence are associated with specific behavioral phenotypes.
(A) Circos plot representing a subset of significant correlations between behaviors (top) and expression of TF modules (bottom). Lines connecting behaviors with TF modules indicate significant associations. TF modules included are those mentioned in the main text or in other figures, and five of nine traits are included for simplicity. All significant correlations between behaviors and TF modules are given in Supplementary file 8. For behaviors, p indicates proportion (e.g. p(pollen) is the proportion of returning foraging trips where the bee carried pollen). (B) Motifs enriched within DAPs show maximum binding probabilities near peak summits. (C) Motifs enriched in promoter regions of forager >layer DEGs show elevated binding probabilities ~ 3 kb upstream of and overlapping TSSs. Motif names and sequences are from FlyFactor (Zhu et al., 2011) for Drosophila melanogaster.
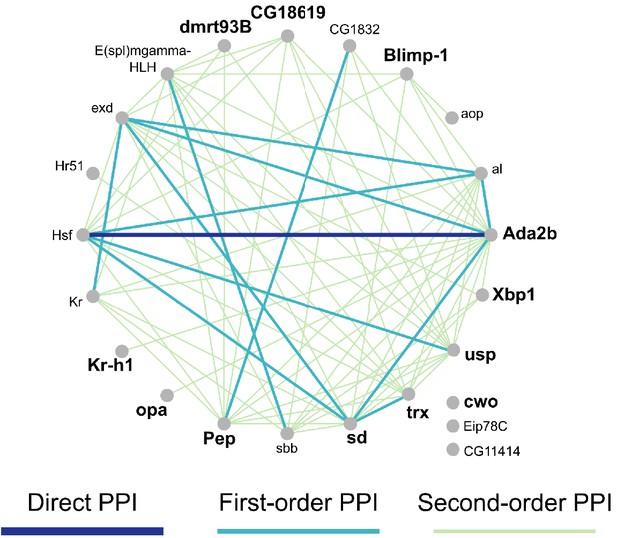
Network of 23 TFs with module expression significantly correlated with nine behavioral and physiological metrics (see Supplementary file 2) measured across individuals.
Edges indicate known interactions based on MIST database for Drosophila melanogaster. First-order PPI indicates one intermediate protein between linked nodes, while second-order PPI indicates two intermediate proteins. PPI = protein protein interaction.
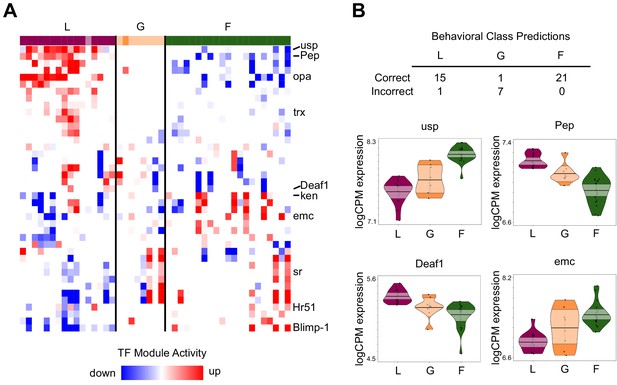
TF module activity and TF expression predict individual variation in behavior.
(A) TF modules (rows) with significant up/downregulation in at least 10 individuals, sorted by hierarchical clustering. Individuals (columns) are ordered by specialist score, with darkly colored blocks indicating correctly classified individuals based on TF expression prediction analysis and lightly colored blocks indicating incorrect classification. TF modules showed patterns of differentiation between L and F, while G were more variable in module activity. Labeled modules are those with TFs shown in panel (B) or discussed in text. (B) Class prediction analysis based on brain TF expression correctly classified all but one specialist (L: layer, F: forager) but only one generalist (G). Normalized expression (logCPM) of 4 of the top 20 informative TFs for class prediction analysis are shown (others in Figure 4—figure supplement 1). Median of points is represented by bold horizontal line within shaded 95% confidence interval, with length of shape and smoothed curve showing range and density of data, respectively.
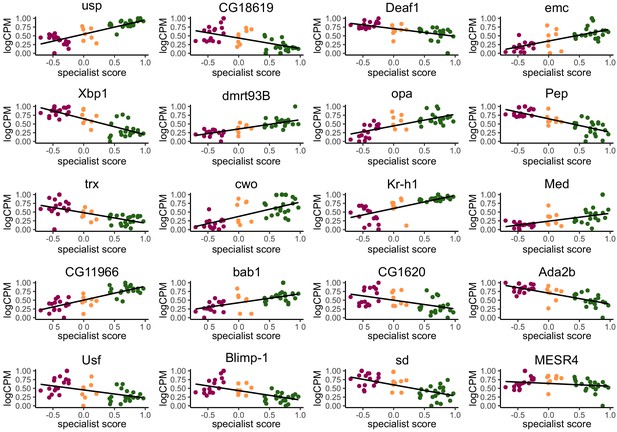
Normalized expression (logCPM, scaled to a maximum of 1 to allow for comparison across TFs) of the top 20 most informative TFs for class prediction analysis plotted against individual specialist score.
Points are colored by behavioral group (Layer: purple, Generalist: orange, Forager: green). Although some of the top predictors individually show weak correlation with specialist score, the random forest machine-learning algorithm combined multiple weak predictors together in a single model to accurately classify the three behavioral groups, suggesting the TFs act combinatorially to influence behavior.
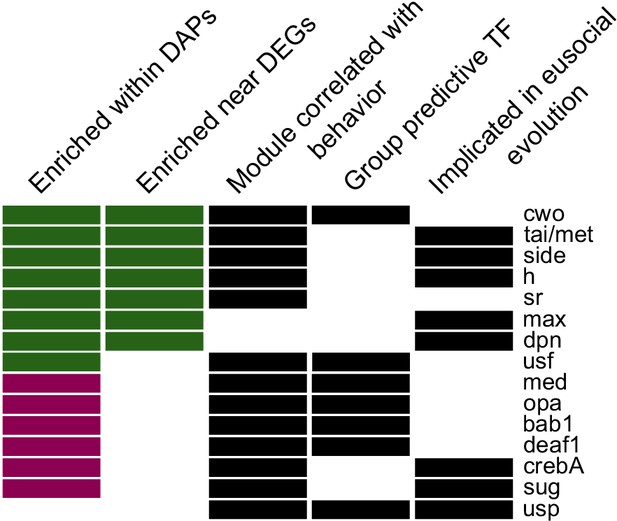
Fifteen candidate TFs predicted to regulate egg-laying and foraging behavior based on evidence across all analyses (descriptions of categories in Materials and methods).
Names given are for Drosophila melanogaster motifs (Zhu et al., 2011), with homology to honey bee genes as in Kapheim et al., 2015. Color of bar in first two columns indicates whether there was stronger enrichment among forager-biased (green) or layer-biased (purple) peaks or genes.
Tables
Description of 15 candidate TFs regulating specialist behavioral phenotypes in Figure 5.
Names given are for Drosophila melanogaster motifs (Zhu et al., 2011), with homology to honey bee genes as in Kapheim et al., 2015. Function summaries are adapted from D. melanogaster gene annotations from FlyBase (release FB2020_05; FlyBase Consortium et al., 2019). Note that terms related to ‘regulation of transcription’ apply to most TFs but were omitted for brevity.
| Motif | TF name | Function(s) |
|---|---|---|
| cwo | clockwork orange | circadian regulation of gene expression; dendrite morphogenesis |
| tai/met | taiman, Mondo | ecdysone receptor co-activator; lipid and carbohydrate metabolism |
| side | sidestep, E(spl)mgamma-HLH | pattern specification; neurogenesis; neuronal stem cell maintenance |
| h | hairy | cell morphogenesis; tracheal system development; cellular metabolism |
| sr | stripe | central nervous system development |
| max | Max | cell and organismal growth |
| dpn | deadpan | adult locomotory behavior; neuroblast development |
| usf | Usf | [unknown] |
| med | Medea | dorsal-ventral patterning; activin receptor signaling; eye morphogenesis; germ-line stem cell division and maintenance; neuron development |
| opa | odd paired | embryogenesis; midgut development; adult head morphogenesis; neural stem cell development; circadian rhythm |
| bab1 | bric a brac 1 | pattern formation; ovary morphogenesis; abdominal pigmentation; olfactory receptor neuron fate diversity |
| deaf1 | Deformed epidermal autoregulatory factor-1 | embryo development; regulation of immune response |
| crebA | Cyclic-AMP response element binding protein A | salivary gland development; cuticle development |
| sug | sugarbabe | regulates expression of insulin-like peptides and genes involved in lipid and carbohydrate metabolism |
| usp | ultraspiracle | cell migration; response to ecdysone; germ cell development; metamorphosis; mushroom body development; neuron remodeling |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Biological sample (A. mellifera) | queens | Honey Bee Insemination Service, Washington State University | SDI colonies | |
| Biological sample (A. mellifera) | workers | University of Illinois Bee Research Facility | ||
| Commercial assay or kit | RNeasy Mini Kit | Qiagen | 74104 | |
| Commercial assay or kit | TruSeq Stranded mRNA HT kit | Illumina | RS-122–2103 | |
| Commercial assay or kit | Nextera DNA Sample Preparation Kit | Illumina | FC-121–1031 | |
| Software, algorithm | Trimmomatic | RRID:SCR_011848 | v0.36 | |
| Software, algorithm | STAR | RRID:SCR_015899 | v2.5.3 | |
| Software, algorithm | bwa | RRID:SCR_010910 | v0.7.17 | |
| Software, algorithm | Picard | RRID:SCR_006525 | v2.10.1 | |
| Software, algorithm | R project for statistical computing | R Core Team | RRID:SCR_001905 |
Additional files
-
Supplementary file 1
Daily egg-laying counts, foraging counts, specialist scores, and generalist scores for individual bees.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp1-v1.xlsx
-
Supplementary file 2
Detailed behavioral and physiological information for sequenced bees.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp2-v1.xlsx
-
Supplementary file 3
Differentially expressed genes (DEGs) and Gene Ontology (GO) enrichment of DEGs for each pairwise comparison of specialists and generalists.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp3-v1.xlsx
-
Supplementary file 4
Differentially accessible peaks (DAPs) and Gene Ontology (GO) enrichment of DAPs for each pairwise comparison of specialists and generalists.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp4-v1.xlsx
-
Supplementary file 5
Lists of genes with upper and lower 5% of PC loadings for gene expression and Gene Ontology (GO) enrichment results.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp5-v1.xlsx
-
Supplementary file 6
Lists of genes with upper and lower 5% of PC loadings for chromatin accessibility and Gene Ontology (GO) enrichment results.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp6-v1.xlsx
-
Supplementary file 7
TF motif enrichment of DAPs and DEGs.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp7-v1.xlsx
-
Supplementary file 8
Predicted gene regulatory network (GRN), GRN module correlations with behavior and physiological measurements, and importance scores of TFs from class prediction analysis.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp8-v1.xlsx
-
Supplementary file 9
Overlaps and statistics for comparative gene expression datasets.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp9-v1.xlsx
-
Supplementary file 10
Details of experimental setup for recorded colonies.
- https://cdn.elifesciences.org/articles/62850/elife-62850-supp10-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/62850/elife-62850-transrepform-v1.docx





