The mitochondrial permeability transition pore activates the mitochondrial unfolded protein response and promotes aging
Figures
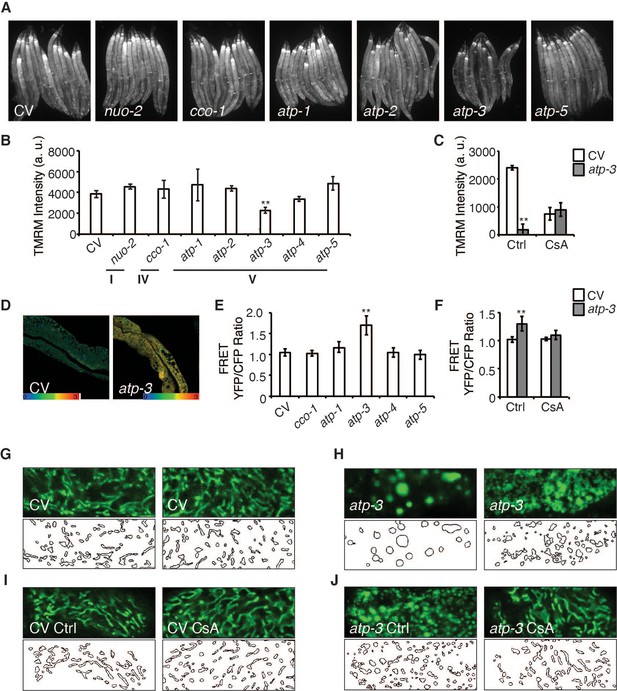
Loss of OSCP/atp-3 during adulthood selectively recapitulates mitochondrial permeability transition pore (mPTP)-like characteristics.
(A) Photomicrographs of mitochondrial membrane potential (MMP) as measured by tetramethylrhodamine methyl ester (TMRM) after RNA interference (RNAi) of OXPHOS subunits. RNAi and TMRM were administered for 48 hr beginning at young adulthood. Representative micrographs shown. (B) Quantification of TMRM intensity from (A). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.01 by Student’s t-test. I, IV, and V correspond to OXPHOS complexes. a.u.: arbitrary units. (C) Quantification of MMP after RNAi of OSCP/atp-3 and treatment with cyclosporin A (CsA: 15 μM). RNAi and CsA were administered beginning at young adulthood for 24 hr. Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. *p≤0.05, **p≤0.01 by Student’s t-test. (D) Confocal micrograph of intestinal cytosolic Ca2+ as measured by the FRET-based calcium indicator protein D3cpv/cameleon after OSCP/atp-3 RNAi. RNAi was administered for 48 hr beginning at young adulthood. Representative micrograph shown. (E) Quantification of FRET YFP/CFP ratio after RNAi of OXPHOS subunits. RNAi was administered for 48 hr beginning at young adulthood. Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.01 by Student’s t-test. (F) Quantification of FRET YFP/CFP ratio after RNAi of OSCP/atp-3 and treatment with CsA (15 μM). RNAi and CsA were administered beginning at young adulthood for 48 hr. Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.01 by Student’s t-test. (G–J) Confocal micrographs of intestinal mitochondria labeled with GFP (pges-1::GFPmt) in young adults. RNAi and CsA were administered for 48 hr beginning at young adulthood, then worms were removed from the RNAi and CsA and aged until day 7 of adulthood followed by collection for microscopy. Top panels: fluorescent channel; bottom panels: rendering of individual mitochondria. CV: control vector; Ctrl: solvent control; CsA: 15 μM. See Materials and methods for details on rendering.
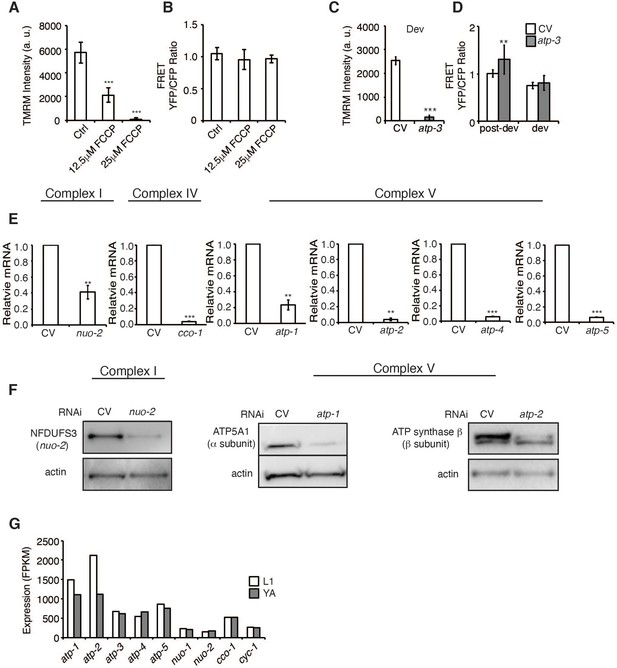
Loss of OSCP/atp-3 RNAi during adulthood uniquely causes a loss in membrane potential and a rise in cytosolic calcium.
(A) FCCP, the mitochondrial uncoupler, significantly induces a loss in mitochondrial membrane potential (MMP). FCCP was administered for 24–48 hr beginning at young adulthood (YA). Data are the mean ± SEM of ≤ 15 animals from three biological experiments. ***p≤0.0001 by Student’s t-test. (B) FCCP does not affect significantly alter cytosolic Ca2+ using the FRET-based calcium indicator protein D3cpv/cameleon. Data are the mean ± SEM of ≤ 15 animals from three biological experiments. (C) Developmental RNAi of OSCP/atp-3 induces a loss in MMP. RNAi was administered beginning from eggs for 72 hr. Tetramethylrhodamine methyl ester (TMRM) was spotted on seeded plates. ***p≤0.0001 by Student’s t-test. (D) Developmental RNAi of OSCP/atp-3 does not significantly alter cytosolic Ca2+ using the FRET-based calcium indicator protein D3cpv/cameleon compared to post-developmental RNAi. Developmental RNAi was administered beginning from eggs for 72 hr. Post-developmental RNAi was administered for 48 hr beginning from YA. **p≤0.01 by Student’s t-test. (E) qPCR from N2 worms after RNAi of subunits from complex I, IV, and V. RNAi was administered for 48 hr beginning at YA. Data are the mean ± SD ≤ 150 animals combined from two experiments. **p≤0.01, ***p≤0.0001 by Student’s t-test. (F) Immunoblots from N2 worms after RNAi of subunits from complex I and V. RNAi was administered for 48 hr beginning at YA. Representative immunoblots from two biological experiments. Actin was used as a loading control. (G) Fragments per kilobase of transcript per million (FPKM) expression values of various OXPHOS subunits collected at L1 larval stage or YA from wormbase.org.
-
Figure 1—figure supplement 1—source data 1
Source data for immunoblot (Figure 1—figure supplement 1F).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig1-figsupp1-data1-v1.jpg.zip
-
Figure 1—figure supplement 1—source data 2
Source data for immunoblot (Figure 1—figure supplement 1F).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig1-figsupp1-data2-v1.jpg.zip
-
Figure 1—figure supplement 1—source data 3
Source data for immunoblot (Figure 1—figure supplement 1F).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig1-figsupp1-data3-v1.jpg.zip
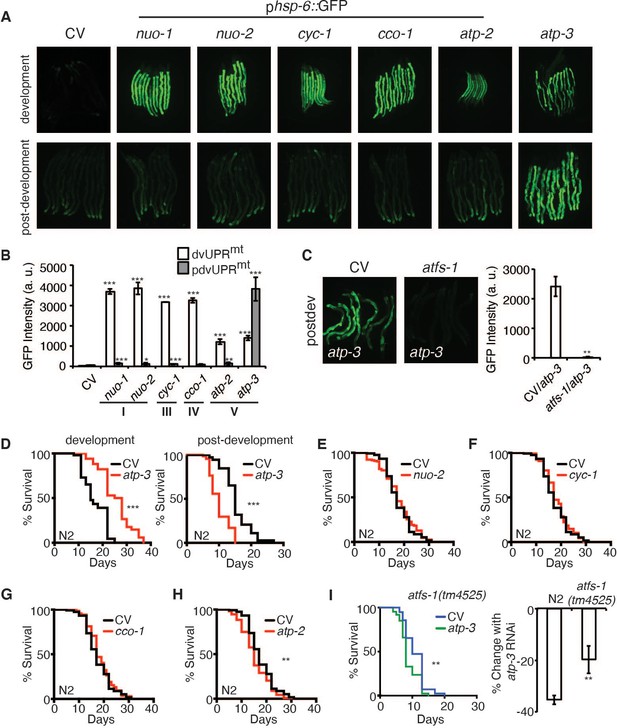
Loss of OSCP/atp-3 during adulthood initiates a unique UPRmt and shortens lifespan.
(A) Photomicrographs of phsp-6::GFP reporter after developmental or post-developmental RNA interference (RNAi) of OXPHOS subunits. For developmental treatment, worms were exposed to RNAi beginning from eggs for 72 hr. For post-developmental treatment, worms were exposed to RNAi beginning from young adulthood for 48 hr. CV: control vector; dev: development; post-dev: post-development. (B) Quantification of GFP intensity from (A). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. *p≤0.05, **p≤0.01, and ***p≤0.0001 by Student’s t-test. dv: development; pdv: post-development; I, III, IV, and V refer to OXPHOS complexes. (C) Photomicrographs of the phsp-6::GFP reporter after RNAi of OSCP/atp-3 and atfs-1. Worms were exposed to RNAi beginning from young adulthood for 48 hr. Bar graph represents quantification of GFP intensity. Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.01 by Student’s t-test. CV: control vector RNAi; pdv: post-development. (D) Survival curves of wild-type N2 animals on CV or OSCP/atp-3 RNAi. For developmental RNAi, worms were treated continuously since eggs. For post-developmental RNAi, worms were treated for 48 hr beginning at young adulthood. Representative curves selected from three biological experiments. ***p≤0.0001 by log rank (Mantel–Cox). (E–H) Survival curves of wild-type N2 animals on indicated RNAi initiated at young adulthood for 48 hr. Representative curves selected from three biological experiments. **p≤0.01 by log rank (Mantel–Cox). (I) Survival curves of atfs-1(tm4525) when OSCP/atp-3 RNAi is initiated at young adulthood for 48 hr. Representative curves selected from three biological experiments. **p≤0.01 by log rank (Mantel–Cox). Bar graph is a quantification of percent change in lifespan of N2 or atfs-1(tm4525) mutant. **p≤0.01 by Student's t-test is the mean of the percent change in lifespan ± SEM from three biological experiments.
-
Figure 2—source data 1
List of genes tested for whether they induce a developmental UPRmt (dvUPRmt) or post-developmental UPRmt (pdvUPRmt).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig2-data1-v1.xlsx
-
Figure 2—source data 2
Summary of lifespans.
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig2-data2-v1.xlsx
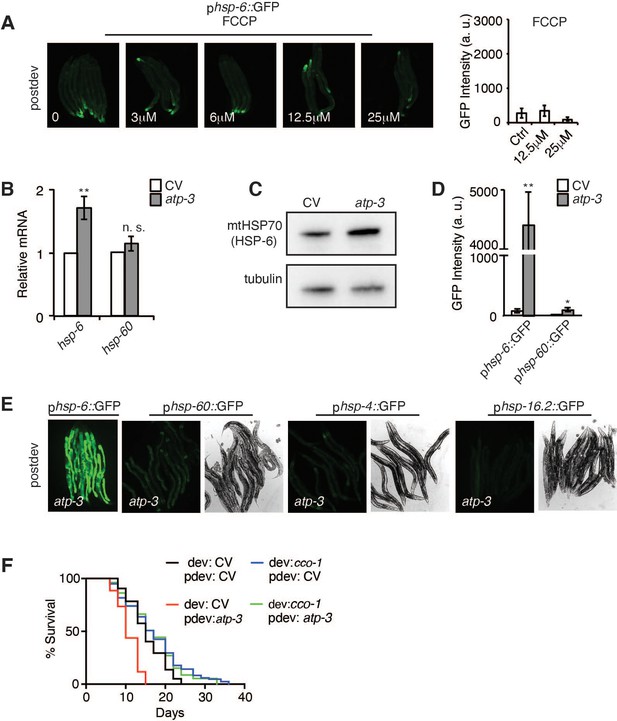
Loss of the OSCP/atp-3 during adulthood induces a specific post-developmental UPRmt.
(A) Dose–response of FCCP, the mitochondrial uncoupler, on the phsp-6::GFP reporter. Quantification of the two highest doses of FCCP from the phsp-6::GFP. (B) qPCR of hsp-6 and hsp-60 from N2 worms on post-developmental CV or OSCP/atp-3 RNAi. Data are the mean ± SEM of ≤ 150 animals combined from four biological experiments. **p≤0.01, n.s. not significant by Student’s t-test. CV: control vector RNAi. (C) Western blot of mtHSP70/HSP-6 and tubulin (loading control) from worms on post-developmental CV or atp-3 RNAi. Representative blot from three biological experiments. (D) Quantification of phsp-6::GFP and phsp-60::GFP reporters from (E). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.01, *p≤0.05 by Student’s t-test. (E) Post-developmental RNAi of OSCP/atp-3 mildly induced the mitochondrial phsp-60::GFP reporter but not the UPRER phsp-4::GFP or heat shock response (HSR) phsp-16.2::GFP reporters. Left panel: fluorescent channel; right panel: bright-field channel. (F) Survival curves of wild-type N2 treated with or without developmental COX5B/cco-1 RNAi (from eggs until young adulthood) followed by post-developmental treatment with OSCP/atp-3 RNAi (for 48 hr and then transferred to regular nematode growth medium (NGM) plates for the remainder of the lifespan). Lifespan curves from two pooled biological replicates. dev: developmental; pdev: post-developmental.
-
Figure 2—figure supplement 1—source data 1
Source data for immunoblot (Figure 2—figure supplement 1C).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig2-figsupp1-data1-v1.jpg.zip
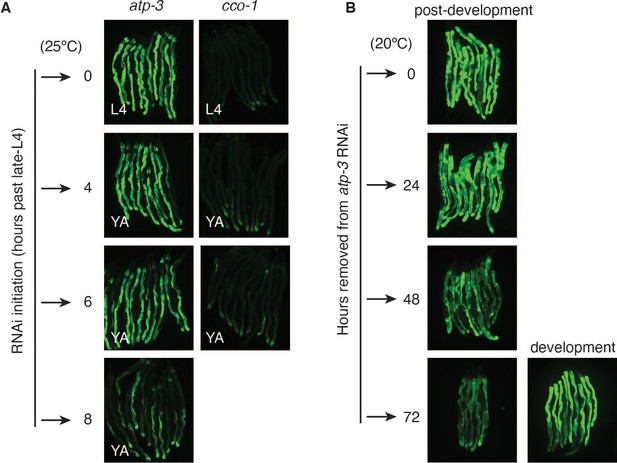
The post-developmental UPRmt is temporally confined and reversible.
(A) Post-developmental RNA interference (RNAi) of OSCP/atp-3 or COX5B/cco-1 RNAi that was initiated at L4, young adult, and adult stages using the phsp-6::GFP reporter at 25°C. GFP expression was assessed 48 hr after RNAi initiation. Representative pictures from two biological experiments. (B) The phsp-6::GFP reporter worms were treated with developmental or post-developmental OSCP/atp-3 RNAi. Worms were removed from HT115 RNAi bacteria and placed onto OP50 bacteria and GFP expression was assessed at indicated timepoints. Representative pictures from two biological experiments.
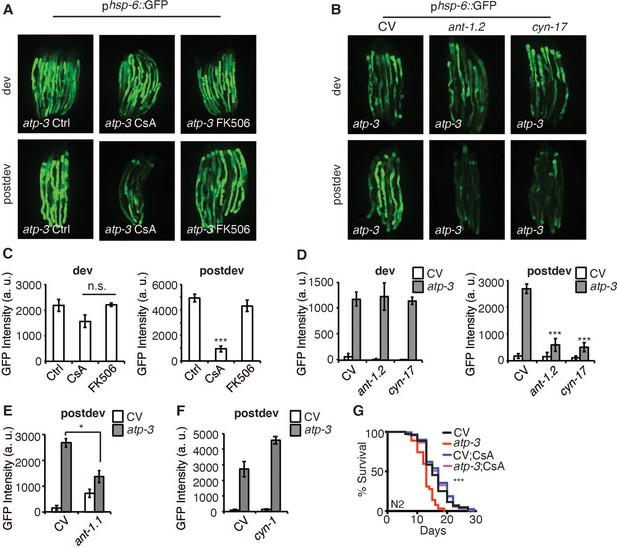
The post-developmental UPRmt is regulated by pharmacological and genetic modulators of the mitochondrial permeability transition pore (mPTP).
(A, B) Photomicrographs of phsp-6::GFP reporter after developmental or post-developmental RNA interference (RNAi) and drug treatments (cyclosporin A (CsA) and FK506, 15 μM). For developmental treatment, worms were treated beginning from eggs for 72 hr. For post-developmental treatment, worms were treated beginning from young adulthood for 48 hr. dev: development; post-dev: post-development. (C, D) Quantification of GFP intensity from (A, B). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. *p≤0.05,***p≤0.0001 by Student’s t-test; n.s., not significant; CV: control vector. (E, F) Quantification of GFP intensity from phsp-6::GFP reporter after RNAi treatment for 48 hr beginning from young adulthood. (G) Survival curves of wild-type N2 animals on CV or OSCP/atp-3 RNAi with either solvent control or CsA (15 μM). RNAi and CsA were administered beginning at young adulthood for 48 hr and then transferred to regular nematode growth medium (NGM) plates for the remainder of the lifespan. Representative curves selected from three biological experiments. ***p≤0.0001 by log rank (Mantel–Cox).
-
Figure 3—source data 1
List of C. elegans cyclophilins and their predicted mitochondrial localization using the MitoFates mitochondrial targeting sequence (MTS) prediction tool.
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig3-data1-v1.xlsx
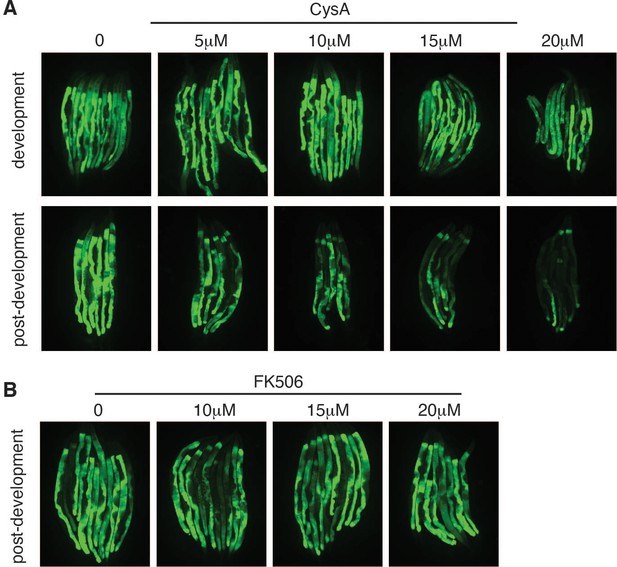
Loss of OSCP/atp-3 during adulthood recapitulates mitochondrial permeability transition pore (mPTP) characteristics.
(A) Dose–response of cyclosporin A (CsA) administered during development or post-development on OSCP/atp-3 RNA interference (RNAi)-treated phsp-6::GFP reporter worms. Developmental RNAi was administered beginning from eggs for 72 hr. Post-developmental RNAi was administered for 48 hr beginning from young adulthood. (B) Dose–response of FK506 administered during post-development on OSCP/atp-3 RNAi-treated phsp-6::GFP reporter worms. Post-developmental RNAi was administered for 48 hr beginning from young adulthood.
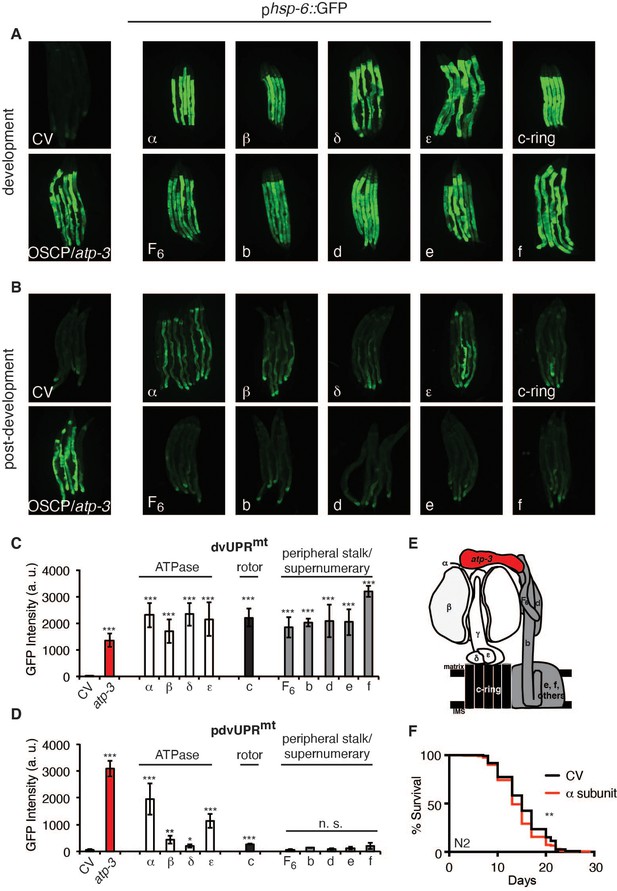
Loss of F-ATP synthase ATPases induces a post-developmental UPRmt.
(A) Developmental RNA interference (RNAi) of all ATP synthase subunits tested induced the phsp-6::GFP reporter. For developmental treatment, worms were exposed to RNAi beginning from eggs for 72 hr. (B) Post-developmental RNAi of ATPase and c-ring subunits but not peripheral stalk or supernumerary subunits of the F-ATP synthase induced the phsp-6::GFP reporter. Worms were exposed to RNAi beginning from young adulthood for 48 hr. post-dev: post-development. (C) Quantification of GFP intensity from (A). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. ***p≤0.0001 by Student’s t-test. dv: developmental. (D) Quantification of GFP intensity from (B). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. *p≤0.05, **p≤0.01, and ***p≤0.0001 by Student’s t-test. pdv: post-developmental. (E) Schematic of monomeric F-ATP synthase. White subunits: ATPase; black subunits: H+-rotor/c-ring; gray subunits: peripheral stalk and supernumerary subunits; red subunit: oligomycin sensitivity-conferring protein (OSCP/atp-3). (F) Survival curves of wild-type N2 animals on CV or α/atp-1 RNAi initiated at young adulthood for 48 hr. Pooled survival curves from four biological experiments. **p≤0.01 by log rank (Mantel–Cox). CV: control vector.
-
Figure 4—source data 1
Summary of the effects of RNA interference (RNAi) of F-ATP synthase subunits on the developmental UPRmt (dvUPRmt) or post-developmental UPRmt (pdvUPRmt).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig4-data1-v1.xlsx
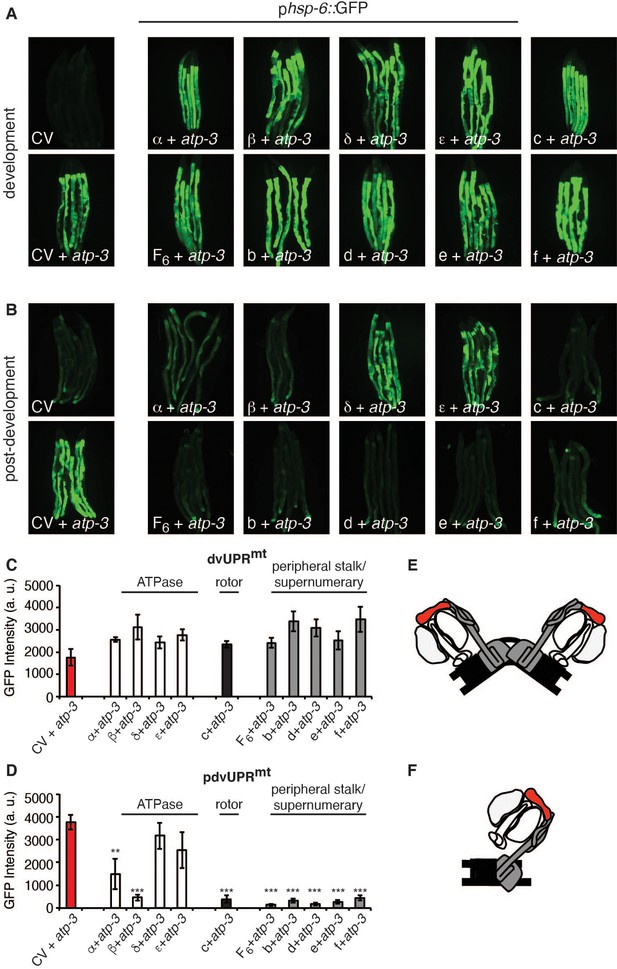
Loss of F-ATP synthase subunits important for the formation of the mitochondrial permeability transition pore (mPTP) suppresses the post-developmental UPRmt.
(A, B) Concomitant RNA interference (RNAi) of OSCP/atp-3 and individual F-ATP synthase subunits during developmental (A) or post-developmental (B) modulated the UPRmt to varying degrees in the phsp-6::GFP reporter strain. (C, D) Quantification of GFP intensity from (A, B). Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.001, ***p≤0.0001 compared to control vector (CV) + atp-3 condition by Student’s t-test. dv: developmental; pdv: post-developmental. (E, F) Models of F-ATP synthase forming a dimeric mPTP (E) or monomeric mPTP (F). White subunits: ATPase; black subunits: H+ rotor/c-ring; gray subunits: peripheral stalk and supernumerary subunits; red subunit: oligomycin sensitivity-conferring protein (OSCP/atp-3).
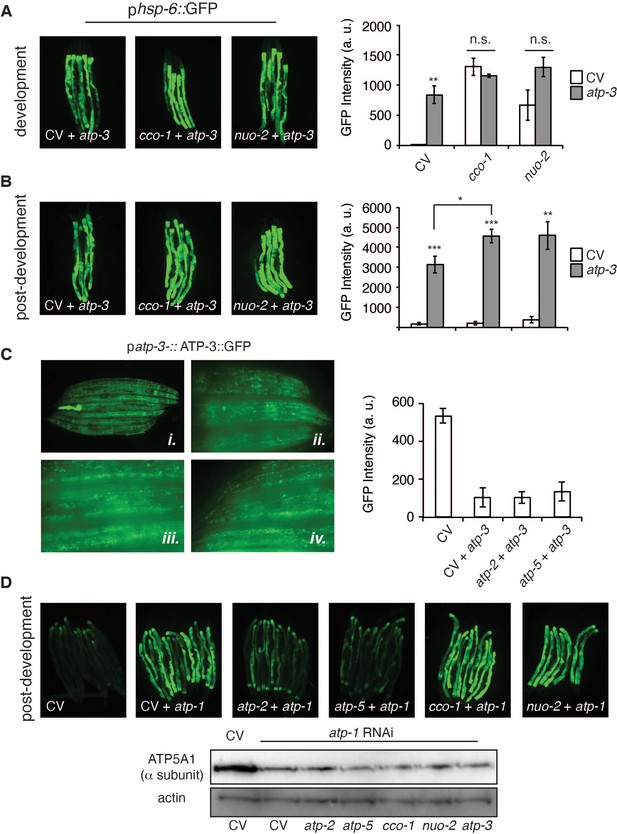
Effects of loss of OXPHOS subunits on the dvUPRmt and pdvUPRmt.
(A, B) Concomitant developmental (A) or post-developmental (B) RNA interference (RNAi) of atp-3 and either cco-1 or nuo-2 induced the phsp-6::GFP reporter. For developmental treatment, worms were exposed to RNAi beginning from eggs for 72 hr. For post-developmental treatment, worms were exposed to RNAi from young adulthood for 48 hr. Bar graphs are the mean ± SEM of ≤ 15 animals combined from three biological experiments. ** p≤0.0001, ***p≤0.0001 by Student’s t-test. n.s., no significant differences from control vector (CV) + atp-3 condition; dvUPRmt: development; pdv: post-development. (C) Dual-RNAi effectively knocks down ATP-3 protein synthesis as measured by the translational patp-3::ATP-3::GFP reporter. (i) 10× photomicrographs of homozygous and heterozygous (bright pharyngeal GFP expression) worms. (ii) 40× photomicrographs of head region of control homozygous worms. (iii) 40× photomicrographs of mid-region of control homozygous worms. (iv) 40× photomicrographs of tail region of control homozygous worms. (D) RNAi of α/atp-1 induces a post-developmental UPRmt similar to OSCP/atp-3 in the phsp-6::GFP reporter. Representative results from two biological experiments. Immunoblot of ATP-1 from phsp-6::GFP reporter worms after dual-RNAi treatment for 48 hr. Representative blots from two biological experiments.
-
Figure 5—figure supplement 1—source data 1
Source data for immunoblot (Figure 5—figure supplement 1D).
- https://cdn.elifesciences.org/articles/63453/elife-63453-fig5-figsupp1-data1-v1.jpg.zip
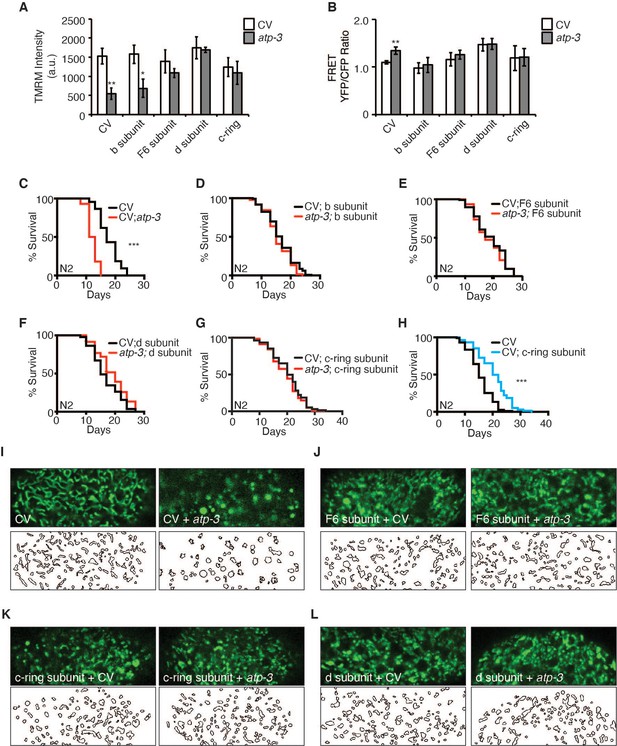
Reversal of mitochondrial permeability transition pore (mPTP) characteristics and regulation of longevity by F-ATP synthase subunits.
(A) Testing for epistatic interactions between OSCP/atp-3 and F-ATP synthase subunits on the mitochondrial membrane potential (MMP). RNA interferences (RNAis) were concomitantly administered beginning at young adulthood for 48 hr. Tetramethylrhodamine methyl ester (TMRM) was spotted on seeded plates. Data are the mean ± SEM of ≤ 15 animals combined from four biological experiments. *p≤0.05, **p≤0.01 by Student’s t-test. CV: control vector. (B) Testing for epistatic interactions between OSCP/atp-3 and F-ATP synthase subunits on cytosolic Ca2+ using the FRET-based calcium indicator protein D3cpv/cameleon. RNAis were concomitantly administered beginning at young adulthood for 48 hr. Data are the mean ± SEM of ≤ 15 animals combined from three biological experiments. **p≤0.01 by Student’s t-test. (C–G) Testing for epistatic interactions between OSCP/atp-3 and F-ATP synthase subunits on the survival. Survival curves of wild-type N2 animals treated with RNAi beginning at young adulthood for 48 hr and then transferred to regular nematode growth medium (NGM) plates for the remainder of the lifespan. Lifespan curves from two pooled biological replicates. ***p≤0.0001 by log rank (Mantel–Cox). (H) Survival curves of wild-type N2 animals on CV or CV/c-ring subunit. Lifespan curves from two pooled biological replicates. ***p≤0.0001 by log rank (Mantel–Cox). CV: control vector RNAi. (I–L) Testing for epistatic interactions between OSCP/atp-3 and F-ATP synthase subunits on mitochondrial morphology. Confocal micrographs of intestinal mitochondria labeled with GFP (pges-1::GFPmt) treated with RNAi for 48 hr beginning at young adulthood. Worms were then removed from the RNAi and aged until day 7 of adulthood followed by collection for microscopy. Top panels: fluorescent channel; bottom panels: rendering of individual mitochondria. See Materials and methods for details on rendering.
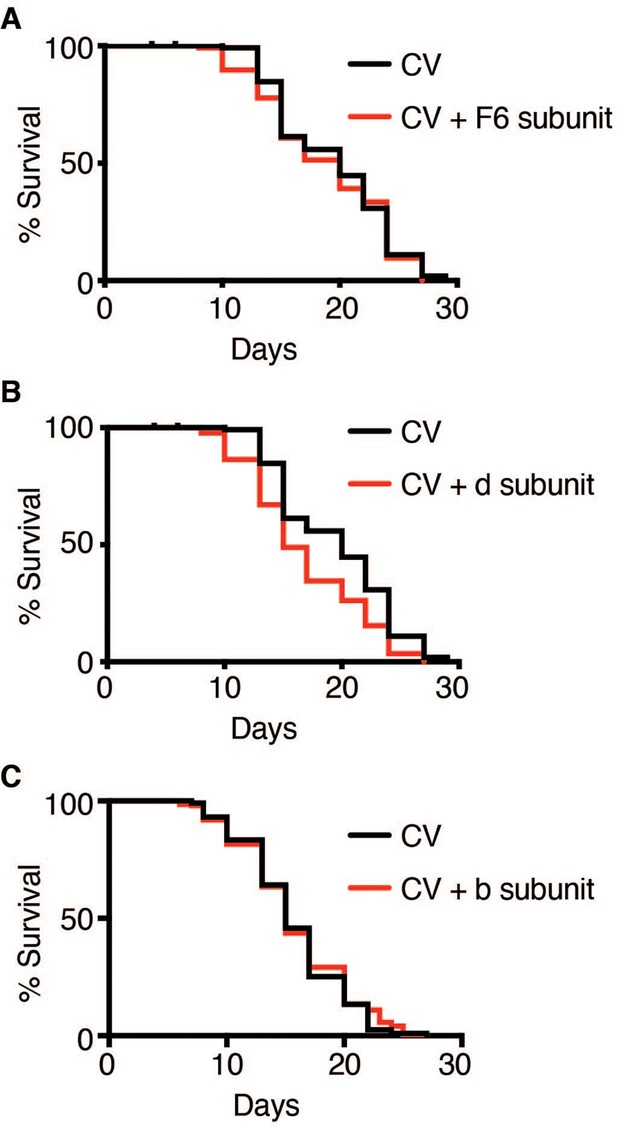
Post-developmental loss of F-ATP synthase peripheral stalk subunits.
(A–C) Survival curves of wild-type N2 worms on indicated RNA interference (RNAi). RNAi was administered starting from young adulthood for 48 hr and then transferred to regular nematode growth medium (NGM) plates for the remainder of the lifespan. Pooled survival curves from two biological experiments.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Caenorhabditis elegans) | N2 Bristol | Caenorhabditis Genetics Center | Wild-type | |
| Strain, strain background (Caenorhabditis elegans) | SJ4100 | Caenorhabditis Genetics Center | hsp-6p::GFP | |
| Strain, strain background (Caenorhabditis elegans) | SJ4058 | Caenorhabditis Genetics Center | hsp-60p::GFP | |
| Strain, strain background (Caenorhabditis elegans) | SJ4005 | Caenorhabditis Genetics Center | hsp-4p::GFP | |
| Strain, strain background (Caenorhabditis elegans) | CL2070 | Caenorhabditis Genetics Center | hsp-16.2p::GFP | |
| Strain, strain background (Caenorhabditis elegans) | KWN190 | Caenorhabditis Genetics Center | rnyEx109[nhx-2p::D3cpv + pha-1(+)], pha-1(e2123) III; (him-5(e1490)V) | |
| Strain, strain background (Caenorhabditis elegans) | ZC376.7 | National BioResource Project | atfs-1(tm4525) | |
| Strain, strain background (Caenorhabditis elegans) | PHX1826 | SunyBiotech | qIs48[atp-3(syb1826)]/hT2[bli-4(e937) let-?(q782)] | |
| Strain, strain background (Caenorhabditis elegans) | SJ4143 | Caenorhabditis Genetics Center | ges-1p::GFP(mt) | |
| Strain, strain background (Caenorhabditis elegans) | GL347 | This study | SJ4100 backcrossed 6× to N2 Bristol | |
| Antibody | Anti-GRP 75 (D9) (mouse monoclonal) | Santa Cruz Biotechnology | sc-133137 | WB(1:1000) |
| Antibody | Anti-β tubulin (D-10) (mouse monoclonal) | Santa Cruz Biotechnology | sc-5274 | WB(1:1000) |
| Antibody | Anti-β-actin (8H10D10) (mouse monoclonal) | Cell Signaling Technology | Cat# 3700 | WB(1:1000) |
| Antibody | Anti-ATP5A1 (15H4C4) (mouse monoclonal) | Thermo Fisher | Catalog # 43-9800 | WB(1:1000) |
| Antibody | Anti-ATP synthase beta (3D5AB1) (mouse monoclonal) | Thermo Fisher | Catalog # A-21351 | WB(1:1000) |
| Sequence-based reagent | Anti-NDUFS3 (17D95) (mouse monoclonal) | Thermo Fisher | Catalog # 43-9200 | WB(1:1000) |
| Chemical compound, drug | FCCP (trifluoromethoxycarbonylcyanidephenylhydrazone) | Cayman Chemical | Item # 15218 | |
| Chemical compound, drug | FK506 (tacrolimus) | Cayman Chemical | Item # 10007965 | |
| Chemical compound, drug | Tetramethylrhodamine methyl ester (perchlorate) (TMRM) | Cayman Chemical | Item # 21437 | |
| Chemical compound, drug | Cyclosporin A | Cayman Chemical | Item # 12088 | |
| Software, algorithm | GraphPad Prism | GraphPad Software | v.9 | |
| Software, algorithm | ImageJ software | ImageJ http://imagej.nih.gov/ij/ | 1.52A | |
| Other | MitoFates tool | http://mitf.cbrc.jp/MitoFates/cgi-bin/top.cgi (Fukasawa et al., 2015). | ||
| Sequence-based reagent | act-1, forward | This study | qPCR primers | ACGACGAGTCCGGCCCATCC |
| Sequence-based reagent | act-1, reverse | This study | qPCR primers | GAAAGCTGGTGGTGACGATGGTT |
| Sequence-based reagent | atp-2, forward | This study | qPCR primers | GAAGGACAAATCTCCCCACA |
| Sequence-based reagent | atp-2, reverse | This study | qPCR primers | CGCCACATTCTTCCTTTTTC |
| Sequence-based reagent | atp-4, forward | This study | qPCR primers | AATATGTTGCCTCCCGTGAT |
| Sequence-based reagent | atp-4, reverse | This study | qPCR primers | GGAACAAAAACGTTCATTCG |
| Sequence-based reagent | atp-5, forward | This study | qPCR primers | TCTTCGACGTGCCGACAA |
| Sequence-based reagent | atp-5, reverse | This study | qPCR primers | AAATGGTAGGAGAGCGATAAGG |
| Sequence-based reagent | nuo-2, forward | This study | qPCR primers | TGAAGTTGCTGAGCCAACAC |
| Sequence-based reagent | nuo-2, reverse | This study | qPCR primers | TCCACACTAACAGAAAATGAGTCT |
| Sequence-based reagent | cco-1, forward | This study | qPCR primers | TTTCGGCTATTGTTCGCATT |
| Sequence-based reagent | cco-1, reverse | This study | qPCR primers | GCCGTCTTAGCAAGTTGAGC |
| Sequence-based reagent | atp-3p::ATP-3::GFP | http://www.sunybiotech.com/ | sgRNA target site | Sg1: CCCTTGCCACCGCCATCTAAatt |
| Sequence-based reagent | atp-3p::ATP-3::GFP | http://www.sunybiotech.com/ | sgRNA target site | Sg2:CCGCCATCTAAatttttcccaaa |





