On the emergence of P-Loop NTPase and Rossmann enzymes from a Beta-Alpha-Beta ancestral fragment
Figures
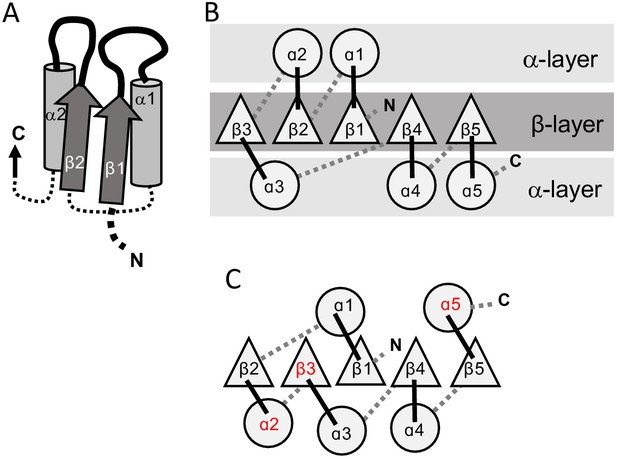
The 3-layer αβα sandwich.
(A) The αβα sandwich is a modular fold comprised of repeating β-loop-α elements. This side view shows two tandem βα elements: the functional loops are situated on the ‘top’ of the fold (thick lines) and the β-loop-α element are linked via short, bottom loops (thin, dashed lines). Shown here are the first two elements with a Rossmann topology, beginning with β1 at the N-terminus, and the first two helices (α1, α2) that, in this cartoon, comprise one external layer of the sandwich. (B) A view from the top reveals the αβα sandwich architecture with its three layers: a parallel β-sheet flanked on both sides by α-helices. The top, active-site loops face the reader whereas the N- and C-termini and the bottom, connecting loops face the back of the page. The order of the β-strands in the interior β-sheet follows the canonical Rossmann topology. (C) The most common, core P-loop NTPase (P-loops) topology. Noted in red are the differences from the Rossmann topology—migration of β3 from the edge to the center, and of α2 and α5 from one external layer to another.
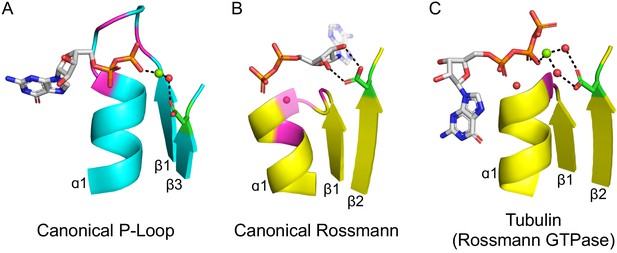
The ligand-binding modes of Rossman and P-loop proteins.
The phosphate binding loops (PBLs) of both lineages connect the C-terminus of β1 to the N-terminus of α1 (conserved glycine residues are colored magenta). The Rossmann β2-Asp, and the P-loop Walker B-Asp, are in green sticks. Water molecules are denoted by red spheres, and metal dications by green spheres. (A) The canonical P-loop NTPase binding mode. The phosphate binding loop (the P-loop Walker A motif; GxxGxGK(T/S)) begins with the first conserved Gly residue at the tip of β1 and ends with a Thr/Ser residing within α1. The Walker B-Asp, located at the tip of β3, interacts with the catalytic Mg2+, either directly or via a water molecule, as seen here. (B) The canonical Rossmann binding mode. The phosphate binding site includes a canonical water molecule (α1 has been rendered transparent so that the conserved water is visible). The Asp sidechain at the tip of β2 (β2-Asp) forms a bidentate interaction with both hydroxyls of the ribose. Note also the opposite orientations of the ribose and adenine moieties in P-loops (pointing away from the β-sheet) versus Rossmann (pointing towards the β-sheet). (C) Tubulin is a GTPase that belongs to the Rossmann lineage. It possesses the canonical Rossmann strand topology, phosphate binding loop (including the mediating water), and β2-Asp. However, the ligand, GTP, is bound in the P-loop NTPase mode (as in A). Accordingly, the β2-Asp makes a water mediated interaction with the catalytic metal cation (Ca2+or Mg2+) thus acting in effect as a Walker B-Asp (the metal cation’s coordination schemes are also identical, see Figure 2—figure supplement 3). ECOD domains used in this figure, from left to right, are e1yrbA1, e1lssA1, and e5j2tB1. All structure figures were prepared in PyMOL (pymol.org).
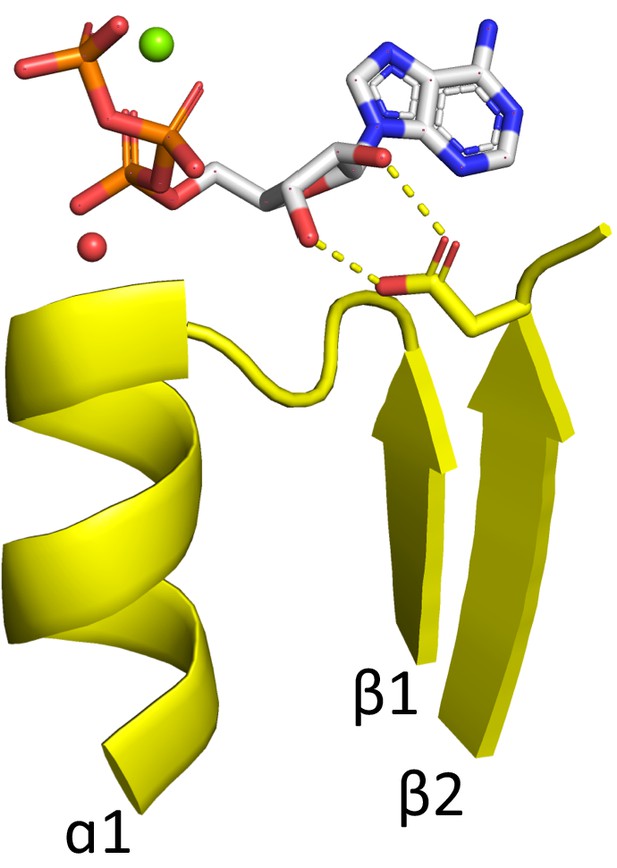
Rossmann domain binding ATP in the canonical binding mode.
The phosphate binding loop of ECOD domain e3h5nA5, a Rossmann domain from the ECOD F-group 2003.1.9.15, binds the nucleotide ATP in a largely canonical fashion, including the bidentate interaction between the β2-Asp and the ribose hydroxyls (see Main Text). The conserved water characteristic of the Rossmann fold is shown as a red sphere. An Mg2+ cation is shown as a green sphere. Other F-groups with canonical binding of an NTP include 2003.1.9.10 and 2003.1.4.3.
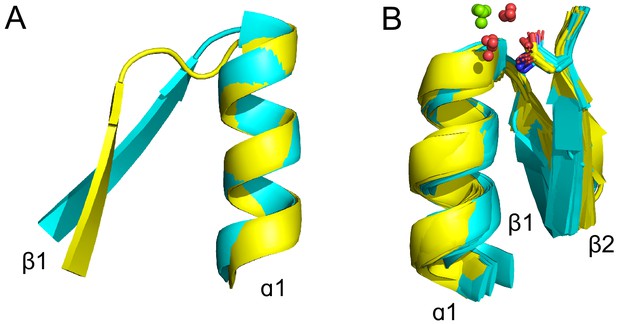
Features of the tubulin binding site (see also Supplementary file 1).
(A) The tubulin phosphate binding loop resides between β1 and α1 (cyan; ECOD domain e5j2tA1) as in all Rossmanns, yet is shorter and more compact than the canonical Rossmann binding loop (yellow; ECOD domain e1lssA1). (B) The conserved β2-Asp (yellow structures; ECOD domains e1ffxB2, e1sa1C2, e2btoA2, e2hxfA2, e3cb2A2, e3e22C2, e3r4vA1, e3zbqA2, e4ffbA4, e4ffbB1) is replaced by asparagine in some structures (cyan structures; ECOD domains e1rq2A2, e1w5fA2, e2r6r11, e2vamA1, e2vapA2, e2xkbC5, e3v3tA3, e3zidA1, e4b45A3, e4b46A1, e4dxdA1, e4e6eA1, e4m8iA2). This residue is positioned to interact with the catalytic Mg2+ (green spheres), typically via a water molecule (red spheres).
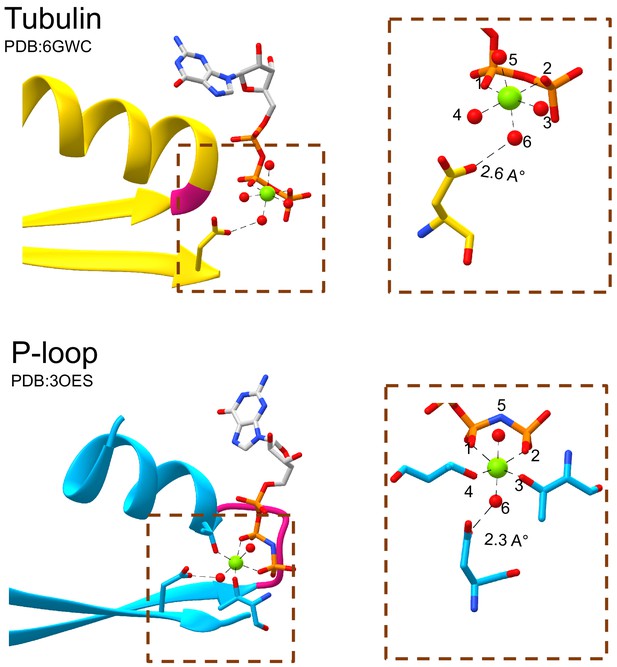
The tubulin β2-Asp and the P-loop Walker B interact with waters that occupy equivalent sites around the catalytic Mg2+ cation.
The catalytic Mg2+ cation in both tubulin (top panel) and P-loop domains (bottom panel) forms octahedral coordination complexes. Both the β2-Asp of tubulin and the Walker B of P-loop interact with the water at position 6 of the coordination sphere. Water molecules are rendered as red spheres and Mg2+ cations are rendered as green spheres.
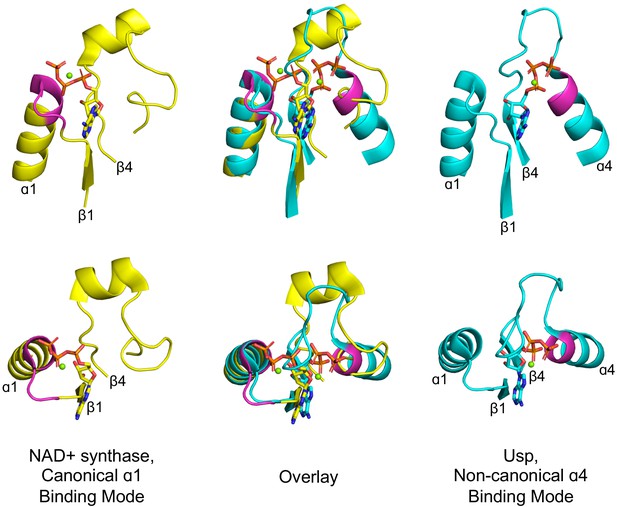
Alternative phosphate binding sites in αβα sandwich enzymes.
HUP proteins are αβα sandwich proteins with a Rossmann-like strand topology. The canonical HUP phosphate binding loop is located at the tip of α1 and colored magenta (left panels; NAD+ synthase; ECOD F-group 2005.1.1.13; shown is domain e1xngA1) as in Rossmann and P-loop NTPases (Figure 2). However, Usp (universal stress proteins) is a HUP family that exhibits kinase activity wherein phosphate binding migrated to the tip of α4 (right panels; ECOD F-group 2005.1.1.145; shown is domain e2z08A1; residues interacting with phosphate groups are colored magenta). As shown in the overlay (middle panels), despite the variation in the phosphate binding site, the ribose and adenine binding modes are identical.
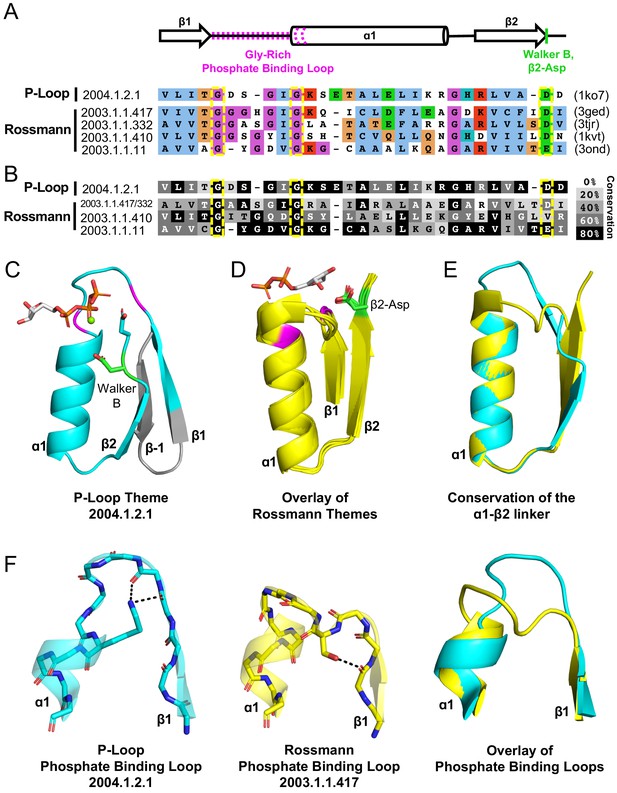
Theme sharing between Rossmann and P-loop enzymes.
(A) Sequence alignment of the shared themes. PDB codes are shown on the right, and the ECOD F-group to which they belong are on the left. The identified themes involve a segment of a single P-loop NTPase, Hpr kinase (top line, ECOD domain e1ko7A1), that aligns to a variety of Rossmanns that belong to four different F-groups (representatives shown here; see Supplementary file 3 for the complete list of bridging themes). (B) The consensus sequence of each F-group (see Methods) is shaded according to the degree of conservation. The individual sequences identified by the theme search show higher similarity by default, yet nonetheless, the family consensus sequences also align well, and the identical residues tend to be conserved. (C-D) Although detection of the shared theme was based on sequence only, structurally, the shared theme encompasses the β1-PBL-α1-β2-Asp element in both the P-loop protein (panel C; Hpr kinase, ECOD domain e1ko7A1) and the theme-related Rossmanns (panel D; ECOD domains e3gedA1, e1kvtA1, e3ondA1, and e3tjrA1; the ligand is bound by domain e3tjrA1). Note that only the pyrophosphate and ribose moieties of the ligand are shown for clarity. The conserved phosphate binding loop glycine residues are colored magenta and the β2-Asp is colored green. For panel C, the ligand binding mode was modeled using the structure of a liganded P-loop protein (see Methods). (E) An overlay of the β1-PBL-α1-β2-Asp element of the Hpr Kinase (cyan; ECOD domain e1ko7A1) and one of the theme-related Rossmann dehydrogenases (yellow; ECOD domain e3tjrA1). (F) Structural details of the phosphate binding loops: The Walker A binding loop of Hrp kinase (left panel; ECOD domain e1ko7A1); the phosphate binding loop of sorbitol dehydrogenase (middle panel; ECOD domain e1k2wA1); and an overlay of both loops (right panel).
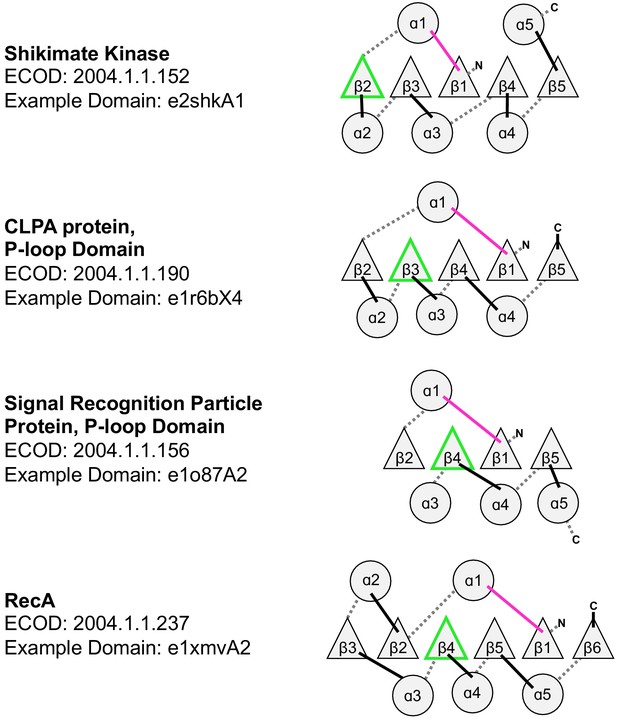
Topological diversity in the P-loop evolutionary lineage.
In addition to the examples noted here, which all have parallel β-sheets, the P-loop lineage also has instances of anti-parallel strands inserted into the β-sheet (see Figure 4C, main text). The loop bearing the Walker A and Walker B motifs are colored magenta and green, respectively.
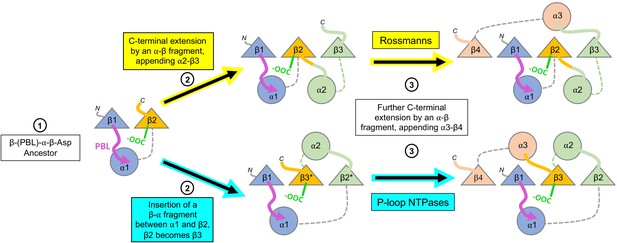
Divergence of the Rossmann and P-loop NTPase folds from a common ancestral polypeptide.
Emergence begins with a presumed β-PBL-α-β-Asp ancestor that could act as either a Rossmann or a P-loop NTPase, depending on how the phospho-ligands bind and the role taken by the β2-Asp (Figure 2A and C). In the second step, the ancestral fragment is either extended at its C-terminus by fusion of an αβ fragment to generate a Rossmann-like domain (top row); or, by insertion of a βα fragment between α1 and β2 to yield a P-loop-like domain (bottom row). Note that βα fragment sequence insertion results in the ancestral β2 that carries the Walker B-Asp becoming β3. Note also that the location of the newly added helix, α2, differs: It can pack next to α1, as in the Rossmann fold; or, it can migrate to the opposite side of the β-sheet, as in the P-loop NTPase fold. Following Figure 1, the top loops are shown as thick lines while the bottom loops are shown as thin, dashed lines. The phosphate binding loop is colored magenta, and the β2-Asp/Walker B-Asp is shown in green.
Additional files
-
Supplementary file 1
Dication binding in tubulins.
- https://cdn.elifesciences.org/articles/64415/elife-64415-supp1-v2.docx
-
Supplementary file 2
Supplementary file 2.
- https://cdn.elifesciences.org/articles/64415/elife-64415-supp2-v2.xlsx
-
Supplementary file 3
Summary of Rossmann proteins that share a bridging theme with the P-Loop domain e1ko7A1.
- https://cdn.elifesciences.org/articles/64415/elife-64415-supp3-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/64415/elife-64415-transrepform-v2.docx





