Integrated sensing of host stresses by inhibition of a cytoplasmic two-component system controls M. tuberculosis acute lung infection
Figures
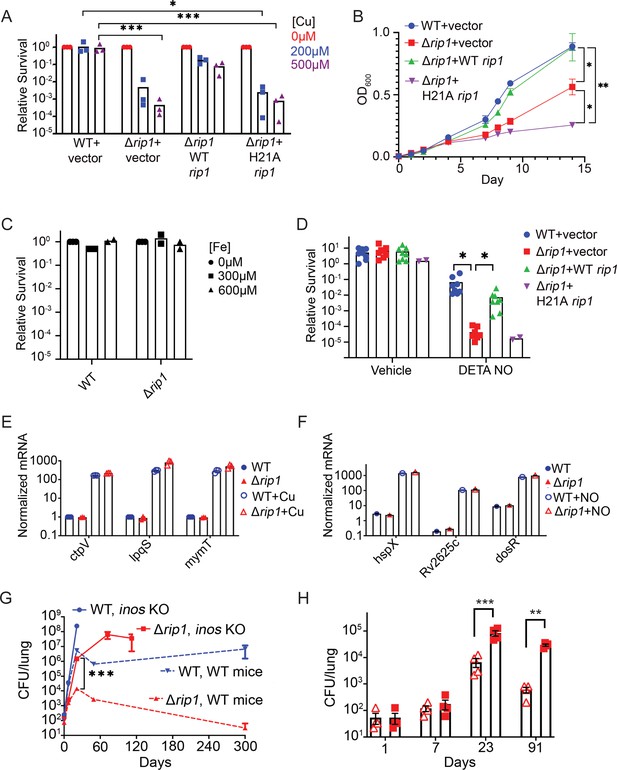
The Rip1 protease controls copper and nitric oxide resistance.
(A) The Rip1 pathway confers resistance to copper. Bacterial colony-forming units (CFUs) of the indicated strains grown on agar plates supplemented with 200 or 500 µM copper sulfate. Relative survival is normalized to untreated controls, which are set at 100 = 1. Each value is the average of technical duplicate measurements for n = 3 biological replicates. Statistical analysis by two-way ANOVA with Tukey’s multi-comparison correction *p<0.01 WT + vector 200 µM vs. Δrip1 +H21A rip1; ***p<0.001 WT + vector 500 µM vs. Δrip1 +vector; WT + vector 500 µM vs. Δrip1 +H21A rip1. (B) The Rip1 pathway confers resistance to zinc. Growth (OD600) of the indicated strains in liquid culture supplemented with 100 µM zinc sulfate. Data plotted is SEM of n = 3 biological replicates. Statistical analysis by ordinary one-way ANOVA with Tukey’s multi-comparison correction. *p<0.05, **p<0.01; day 14 WT + vector vs. Δrip1 + vector p=0.027; day 14 Δrip1 +vector vs. Δrip1 + H21A rip1 p=0.032; day 14 WT + vector vs. Δrip1 + H21A rip1 p=0.002. (C) The Rip1 pathway does not control iron sensitivity. Growth on agar plates supplemented with 300 or 600 µM iron chloride normalized to untreated controls as in (A). Each value is the average of technical duplicate measurements for n = 2 biological replicates. (D) The Rip1 pathway confers resistance to nitric oxide. CFU counts of the indicated strains post treatment with vehicle or 200 µM diethylenetriamine nitric oxide adduct (DETA-NO) for 3 days. Relative survival = CFU day 3 post treatment/CFU at day 0 for each condition n = 8 biological replicates for WT + vector, Δrip1 + vector, and Δrip1 + WT rip1, and n = 2 for Δrip1 + H21A rip1. Statistical analysis by unpaired t-test. *p<0.05; DETA-NO WT + vector vs. Δrip1 + vector p=0.041; DETA-NO Δrip1 + vector vs. Δrip1 + WT rip1 p=0.031. (E) Loss of Rip1 does not affect Cu-induced transcription. RT-qPCR quantitation of known copper-regulated transcripts ctpV, lpqS, and mymT between WT (blue) and Δrip1 (red) cells in resting (filled symbols) or 200 µM copper sulfate (open symbols) conditions for 2 hr. Values are normalized to sigA transcript levels. Values represent average of three technical replicate measurements of three biological replicates. (F) Loss of Rip1 does not affect nitric oxide-regulated gene expression through DosR. RT-qPCR comparison of nitric oxide-induced, DosR-regulated transcripts hspX, rv2625c, and dosR between WT and Δrip1 cells following 3 hr of treatment with vehicle (filled symbols) or 200 μM DETA-NO (empty symbols). Values are normalized to sigA transcript levels. Mean of biological triplicates reported. (G) NO mediates attenuation of M. tuberculosis Δrip1 in mouse lung infection. Bacterial burden (CFU) in lungs of C57bl/6 (broken lines), and age-matched Nos2-/- (solid lines) mice infected with WT M. tuberculosis (blue) or M. tuberculosis Δrip1 (red) at the indicated times (days) post infection. Data plotted is mean and SD of n = 4 biological replicates. ***p=0.0003 by unpaired t-test at 21-day time point for Δrip1 in WT vs. Nos2 -/-. (H) Repeat experiment as in (G), but with lower initial inoculum of Δrip1 M. tuberculosis in WT (open triangles) and Nos2-/- (closed squares) mice. For all strains/host day 1 n = 3 biological replicates; n = 4 biological replicates for all other time points. Statistical significance by unpaired t-test analysis is represented as *p<0.05, **p<0.01, ***p<0.001.
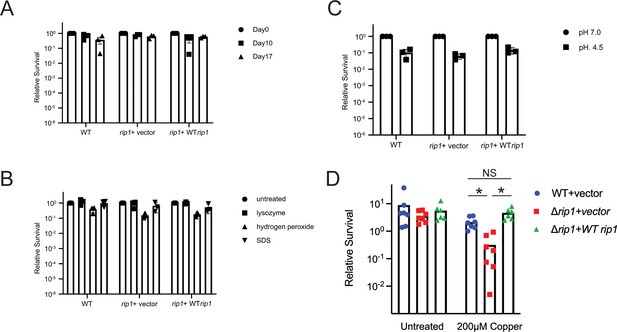
Rip1 does not defend against starvation, oxidative stress, detergent, or acid.
(A) Phosphate buffered saline (PBS) starvation sensitivity assay. Colony-forming units (CFUs) of the indicated strains were determined at the indicated time points post inoculation into standing flask containing PBS at a seeding density of 0.1 OD600. Relative survival = CFU Day X post inoculation/CFU at D0. n = 3 biological replicates. (B) Lysozyme, hydrogen peroxide, and sodium dodecyl sulfate (SDS) sensitivity assay. CFUs of the indicated strains were determined after treatment with 2.5 mg/mL lysozyme, 10 mM hydrogen peroxide, 0.05% SDS for 3 hr. Relative survival = CFU treatment/CFU matched untreated control. n = 3 biological replicates. (C) pH 4.5 survival assay. CFU of the indicated strains was determined after 7 days of incubation in 7H9 ADS media at the indicated pH. Relative survival = CFU pH 4.5/CFU pH 7.0. n = 3 biological replicates. (D) Loss of Rip1 confers sensitivity to Cu in liquid culture. The indicated strains were grown with or without 200 μM CuSO4 for 3 days and bacterial survival quantitated by culturing on agar media without copper. n = 7 biological replicates for WT + vector, Δrip1 + vector; n = 6 biological replicates for Δrip1 + WT rip1. *p<0.01 by Welch’s t-test.
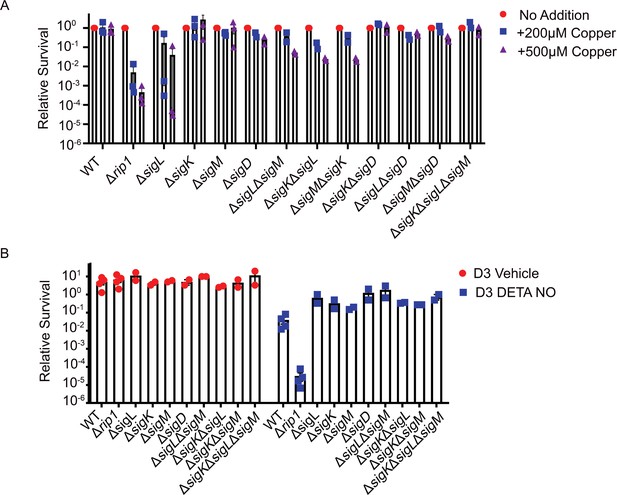
Rip1-controlled sigma factor pathways do not contribute to Cu or NO resistance.
(A) Agar plate copper sensitivity assay. Colony-forming unit (CFU) of the indicated strains grown on agar plates supplemented with 200 or 500 µM copper sulfate normalized to untreated controls. Relative survival = CFU treatment/CFU untreated (No Addition). For WT and Δrip1, each value is the average of technical duplicate measurements for n = 3 biological replicates. Values for all other strains are the average of technical duplicates measured for n = 2 biological replicates. (B) Nitric oxide sensitivity assay. CFU counts of the indicated strains post treatment with vehicle or 200 µM diethylenetriamine nitric oxide adduct (DETA-NO) for 3 days. Relative survival = CFU day 3 post treatment/CFU at day 0 for each condition. n = 4 biological replicates for WT and Δrip1. n = 2 biological replicates for all other strains.
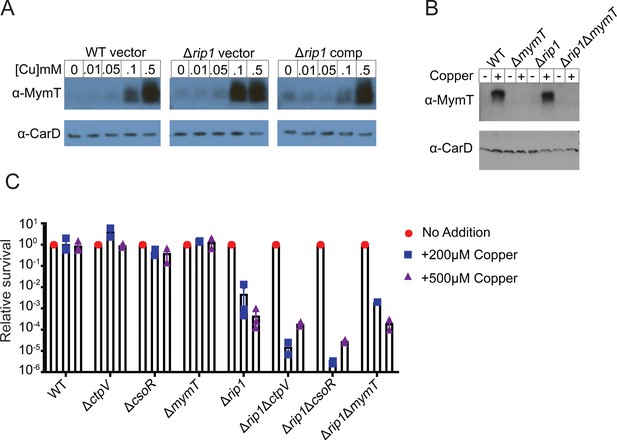
Rip1 copper sensitivity is not through known Cu resistance pathways.
(A) Rip1 does not control Cu-induced transcription through RicR. Lysates generated from the indicated strains, treated for 2 hr with the indicated concentrations of copper sulfate, were separated by SDS-PAGE and probed with anti-MymT rabbit polyclonal sera (upper panel). Blots were re-probed with rabbit polyclonal sera against CarD (lower panel) to determine relative loading. (B) Immunoblot confirmation of MymT deletion. Lysates generated from the indicated strains treated for 2 hr with 5 mM copper sulfate (+) or without addition of excess copper (-) were separated and probed as described in (A). (C) Rip1 copper sensitivity is independent of CtpV, CsoR, or MymT. Bacterial survival of the indicated strains grown on agar plates supplemented with 200 or 500 µM copper sulfate normalized to untreated controls. Relative survival = CFU treatment/CFU untreated (No Addition). For WT and Δrip1, each value is the average of technical duplicate measurements for n = 3 biological replicates. Values for all other strains are the average of technical duplicates measured for n = 2 biological replicates.
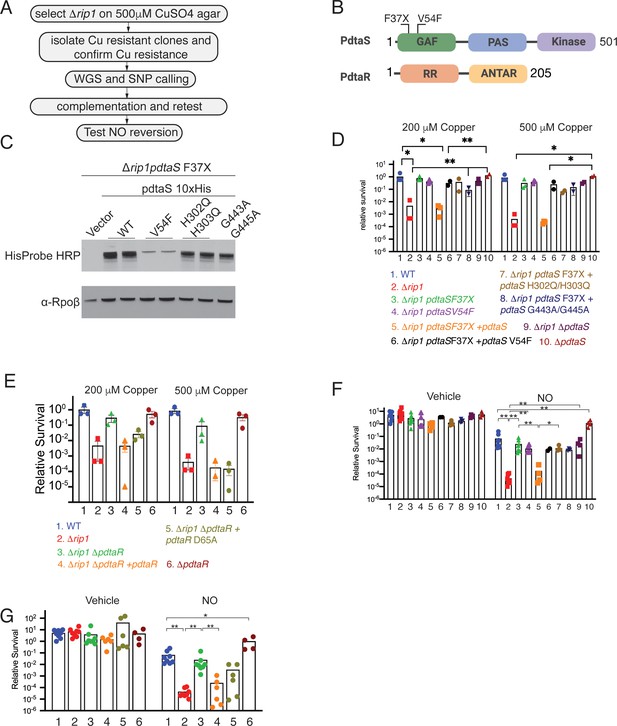
The PdtaS/PdtaR two-component system controls copper and NO resistance downstream of Rip1.
(A) Flow chart of a genetic suppressor screen to isolate reversion mutations of the Rip1 Cu sensitivity and testing their reversion of NO sensitivity. WGS: whole genome sequencing; SNP: single nucleotide polymorphism. (B) Domain structure of PdtaS with GAF, PAS, and kinase domains. Identified suppressor mutations of PdtaS are shown in the N-terminal GAF domain. The reported phosphorylation target of PdtaS is the PdtaR response regulator (RR), which contains a C-terminal ANTAR RNA binding domain. (C) Protein levels of the indicated alleles of pdtaS reintroduced into the Δrip1pdtaS F37X background as C-terminal 10X His fusion proteins with Rpoβ levels as a loading control. For WT, V54F, and H302Q/H303Q, two independent strains are shown, whereas for vector and G443A/G445A, one strain is shown. (D) Loss of pdtaS suppresses copper sensitivity of Δrip1. Relative survival of the indicated strains grown on agar plates supplemented with 200 or 500 µM copper sulfate normalized as in Figure 1. H302Q/H303Q and G443A/G445A are kinase dead alleles. Each value is the average of technical duplicate measurements for n = 3 biological replicates. Statistical analysis by two-way ANOVA with Tukey’s multi-comparison correction. (E) Loss of pdtaR suppresses copper sensitivity of Δrip1. Cu sensitivity assay as noted in (D). PdtaR D65A lacks the receiver aspartate for PdtaS phosphorylation. Each value is the average of technical duplicate measurements for n = 3 biological replicates. (F) Loss of pdtaS suppresses NO sensitivity of Δrip1. Colony-forming unit (CFU) counts of the indicated strains post treatment with vehicle or 200 µM diethylenetriamine nitric oxide (DETA-NO) for 3 days. Color coding of strain genotypes is the same as in (D). n = 8 biological replicates for WT, Δrip1, Δrip1pdtaS F37X; n = 6 biological replicates for Δrip1pdtaS V54F; n = 4 biological replicates for Δrip1pdtaS F37X + pdtaS, Δrip1pdtaS F37X + pdtaS H302Q/H303Q, Δrip1ΔpdtaS, ΔpdtaS; n = 3 biological replicates for Δrip1pdtaS F37X + pdtaS G443A/G445A; n = 2 for Δrip1pdtaS F37X + pdtaS V54F. Statistical analysis by Mann–Whitney test. (G) Loss of pdtaR suppresses NO sensitivity of Δrip1. CFU counts of the indicated strains post treatment with vehicle or 200 µM DETA-NO for 3 days. Color coding of strain genotype is the same as in (E). n = 8 biological replicates for WT, Δrip1, Δrip1 ΔpdtaR; n = 6 biological replicates for Δrip1ΔpdtaR + pdtaR and Δrip1ΔpdtaR +pdtaR D65A; n = 4 biological replicates for ΔpdtaR. Statistical analysis by Mann–Whitney test. For all panels, statistical significance is represented as *p<0.05, **p<0.01, ***p<0.001.
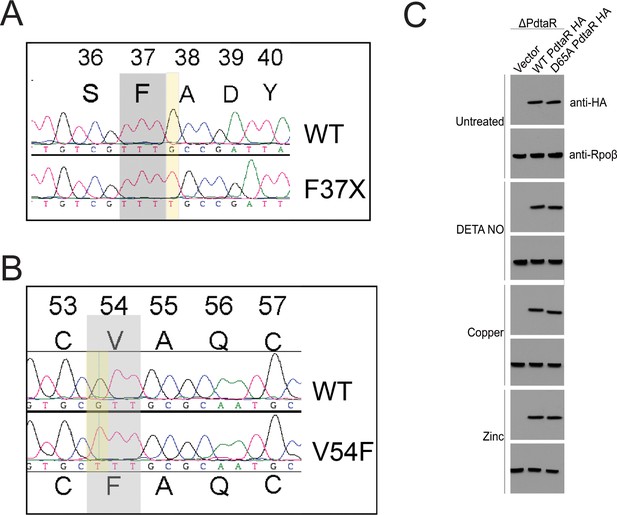
Chromosmoal PdtaS suppressor mutations and PdtaR levels under stress.
(A, B) Chromatograms from seqeuncing of amplified chromosomal segments from two Δrip1 suppressor strains with restored Cu resistance. (A) shows the T insertion at amino acid 37 of PdtaS, creating a frameshift mutation. (B) shows a G>T mutation that encodes V54F in PdtaS. (C) Western blotting with anti-HA antibodies in M. tuberculosis ΔpdtaR complemented with vector, WT PdtaR HA, or PdtaR-D65A-HA in the indicated stress conditions.
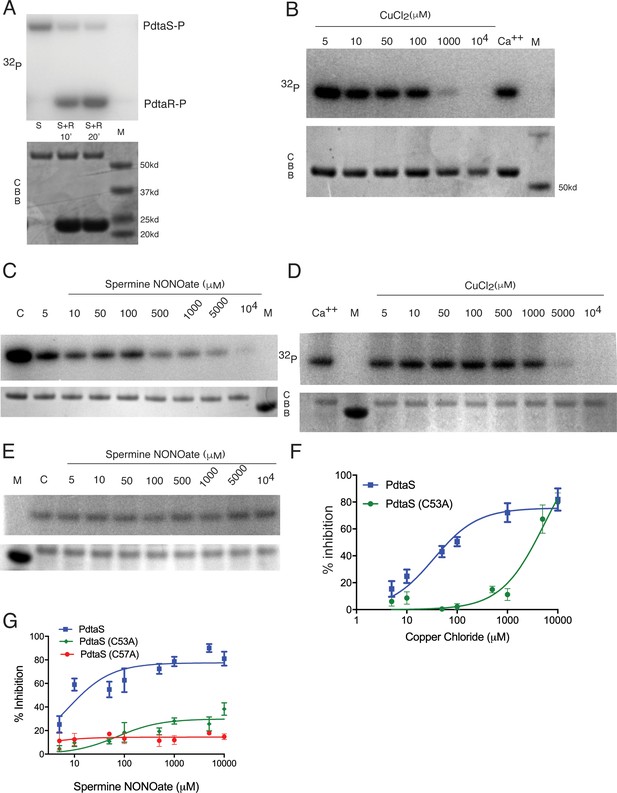
PdtaS is directly inhibited by copper and NO through a dicysteine motif in the N-terminal GAF domain.
(A) PdtaS phosphotransfer to PdtaR. Upper panel: phosphorscreen imaging of 32P incorporation; lower panel: Coomassie brilliant blue (CBB) staining to determine total protein. First reaction contains PdtaS alone, second and third lanes contain PdtaS/PdtaR incubated for 10 and 20 min, respectively. Fourth lane (M) is the molecular weight (MW) marker. (B) Cu++ inhibits PdtaS autophosphorylation. Autophosphorylation of PdtaS protein preincubated with increasing concentrations (5 µM to 104 µM) of CuCl2 or 1 mM of CaCl2 (Ca++) as a control. Upper panel: phosphorscreen imaging of 32P incorporation; lower panel: CBB staining to determine total protein. (C) NO inhibits PdtaS autophosphorylation. Autophosphorylation of PdtaS protein preincubated with increasing concentrations (5 µM to 104 µM) of spermine NONOate and 1 mM of spermine NONOate post NO release as control (designated by C). (D) PdtaS GAF domain cysteine 53 is required for copper inhibition. PdtaS-C53A protein preincubated with increasing concentrations (5 µM to 104 µM) of CuCl2 or 1 mM of CaCl2 was assayed for autophosphorylation as described in panel B . (E) PdtaS GAF domain cysteine 53 is required for NO inhibition. PdtaS-C53A protein preincubated with increasing concentrations (5 µM to 104 µM) of spermine NONOate and of 1 mM of spermine NONOate post NO release as control was followed by autophosphorylation. (F) Cu inhibition curve from replicate data represented by (B) and (D) for PdtaS (blue) and PdtaS-C53A (green) proteins yields Ki of (34 ± 8 µM) for PdtaS and (5540 ± 2381 µM) for PdtaS-C53A. Error bars are SEM for n = 3. (G) NO inhibition curve from replicate data represented by (C) and (E) and (Figure 3—figure supplement 2) for PdtaS (blue), PdtaS-C53A (green), or PdtaS-C57A (red) proteins yields Ki of (7 ± 2 µM) for PdtaS and (73 ± 34 µM) for PdtaS-C53A. Error bars are SEM for n = 3 for WT and C53A and n = 2 for C57A.
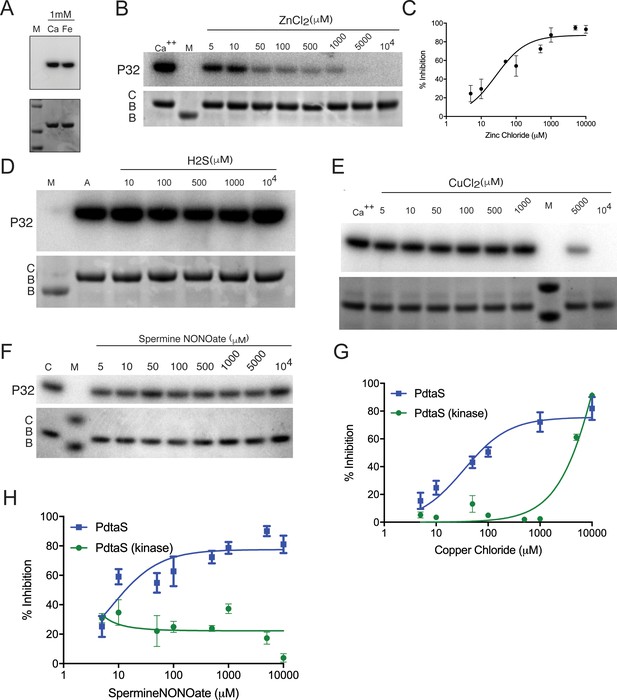
PdtaS is inhibited by Zn and Cu through its GAF domain.
(A) PdtaS is not inhibited by calcium or iron. Autophosphorylation of PdtaS protein preincubated with 1 mM CaCl2 or FeCl2. Upper panel: phosphorscreen imaging of 32P incorporation; lower panel: Coomassie brilliant blue (CBB) staining to determine total protein. M: molecular weight marker. (B) Zinc inhibits PdtaS autophosphorylation. Autophosphorylation of PdtaS protein preincubated with increasing concentrations (5 µM to 10 mM) of ZnCl2 or 1 mM of CaCl2 (Ca++) as a control. Upper panel: phosphorscreen imaging of 32P incorporation; lower panel: CBB staining to determine total protein. (C) Zn inhibition curve from replicate data represented by (B) for PdtaS protein. Error bars are SEM for n = 3. (D) PdtaS is not inhibited by H2S. Same assay as in (B), but with escalating concentration of H2S from sodium hydrosulfide hydrate. (E) The PdtaS kinase domain is not inhibited by Cu. PdtaS-kinase (amino acids 277–501) domain was assayed for autophosphorylation with the indicated concentrations of CuCl2. (F) The PdtaS kinase domain is not inhibited by NO. PdtaS-kinase (amino acids 277–501) domain was assayed for autophosphorylation with the indicated concentrations of spermine NONOate. (G) Cu inhibition curve from replicate data represented by (E) for PdtaS-kinase domain protein. The WT PdtaS curve is the same data presented in Figure 3F. Error bars are SEM for n = 3. (H) NO inhibition curve from replicate data represented by (F) for PdtaS-kinase domain protein. The WT PdtaS curve is the same data presented in Figure 3G. Error bars are SEM for n = 3.
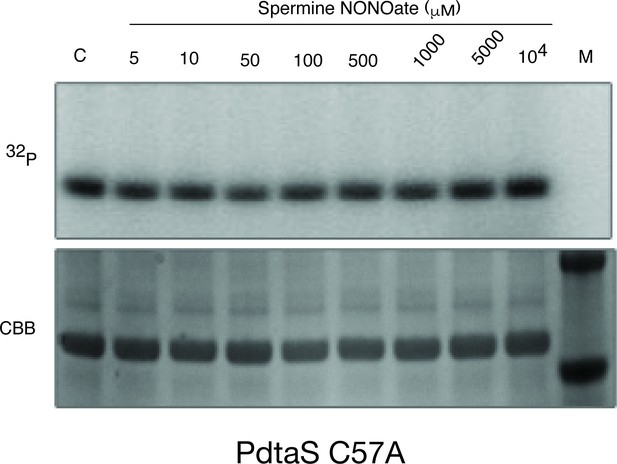
PdtaS C57A is not inhibited by NO.
Autophosphorylation of the PdtaS protein as in Figure 3 using the PdtaS-C57A protein.
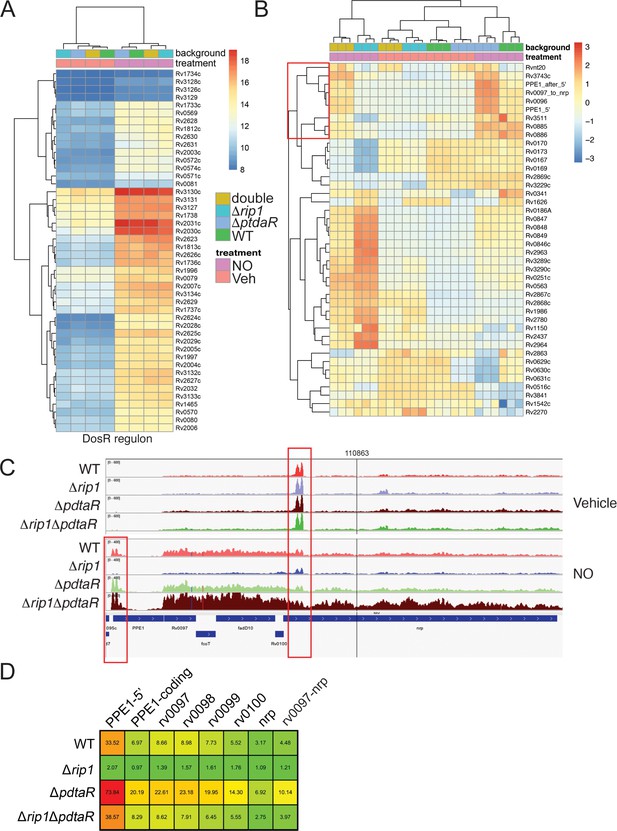
NO-induced dual regulation of the chalkophore biosynthetic operon by Rip1/PdtaS/PdtaR.
(A) Unsupervised clustering of the diethylenetriamine nitric oxide (DETA-NO) (NO)-induced, DosR-regulated transcripts across the indicated genotypes. The scale bar represents the log2 of the normalized counts for each gene. (B) Unsupervised clustering of gene expression of M. tuberculosis WT, Δrip1, ΔpdtaR, and Δrip1ΔpdaR using the same color coding as in (A) treated with vehicle (V) or DETA-NO (NO) The highlighted cluster indicated by the red box on the left includes PPE1-5′, the PPE1 coding sequences after PPE1 (PPE1-after 5′), and the Rv0097-nrp cluster as independent elements. The scale bar represents the log2 of the scaled expression level for each row. The clustering of the genes and strains/conditions is done on the raw normalized counts. (C) Read coverage tracks across the ppe1-nrp cluster. Boxed areas highlight ppe1-5′ peak at the 5′ end of PPE1 and a region of potential termination at 5′ end of nrp. (D) DETA-NO-induced fold change values (DETA-NO/vehicle) across the same coverage region as in (C). 0097-nrp designates the entire Rv0097-nrp cluster.
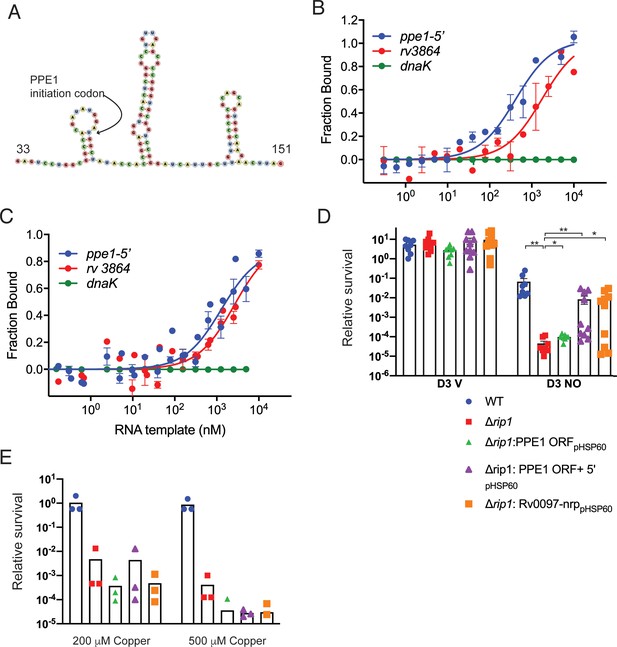
The PPE1-5′ RNA is an effector of NO resistance through sequestration of PdtaR.
(A) Predicted structure of the RNA hairpins in PPE1-5′. The nucleotide numbering refers to distance from the first transcribed nucleotide of the ppe1 RNA. The PPE1 translational initiation codon is indicated within the first hairpin. (B) Phosphorylated PdtaR directly binds to ppe1-5′. Change in the thermophoretic movement of fluorescently labeled and phosphorylated PdtaR was measured as a function of titrant RNA concentration on the X-axis for ppe1-5′, rv3864, and dnaK (ranging from 0.31 nM to 10 µM) as described in Materials and methods yielding binding affinity constants (Kd) for ppe1-5′ = 410 ± 84 nM, rv3864 = 2117 ± 798 nM while dnaK shows no binding. Error bars are SEM for n = 3. (C) RNA binding by PdtaR is partially phosphorylation dependent. Identical assay as in (B) using PdtaR without phosphorylation by PdtaS. Changes in the thermophoretic movement of fluorescently labeled PdtaR were measured as a function of titrant RNA concentration: ppe1-5′ (0.15 nM to 10 µM), rv3864 (0.61 to 10 µM), and dnaK (0.15 nM to 10 µM). Binding affinities (Kd) are ppe1-5′ = 1281 ± 322 nM, rv3864 = 2592 ± 832 nM while dnaK shows no binding. Error bars represent SEM for n = 3. (D) Rip1/PdtaS/PdtaR control of PPE1-5′ and chalkophore biosynthesis controls NO resistance. Constitutive expression of annotated protein coding region of ppe1 alone (+ppe1 ORF hsp60), ppe1 coding region plus 222-nt 5′ of start codon (+ppe1-5′), both C-terminally fused to GFP, or the chalkophore cluster (+rv0097-nrp) in Δrip1 and testing for diethylenetriamine nitric oxide (DETA-NO) sensitivity. Relative survival of the indicated strains post treatment with vehicle or 200 µM DETA-NO. n = 8 biological replicates for WT, Δrip1, Δrip1: PPE1 ORFpHSP60; n = 10 biological replicates for Δrip1:PPE1 ORF +5'pHSP60 and Δrip1: Rv0097-nrppHSP60. Statistical analysis by Mann–Whitney test. (E) Rip1/PdtaS/PdtaR control of PPE1-5′ and chalkophore biosynthesis does not control copper resistance. Relative survival of the same strains as in (D) grown on agar plates supplemented with 0, 200, or 500 µM copper sulfate normalized as in Figure 1. Each value is the average of technical duplicate measurements for n = 3 biological replicates. For all panels, statistical significance by t-test analysis is represented as *p<0.05, **p<0.01, ***p<0.001.
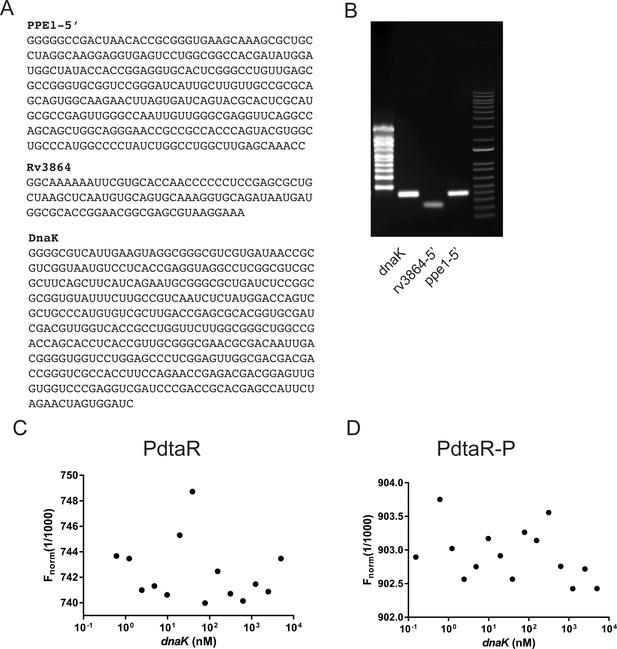
RNAs used in PdtaR binding experiments.
(A) RNA sequences of RNAs produced by in vitro transcription and used in binding studies in Figure 5B, C. (B) Gel electrophoresis of purified RNAs produced by in vitro transcription. (C, D) Microscale thermophoresis titrations for dnaK RNA showing no appreciable binding to PdtaR (C) or PdaR-P (D) for the DnaK RNA. This data is represented on the binding curves in Figure 5B, C.
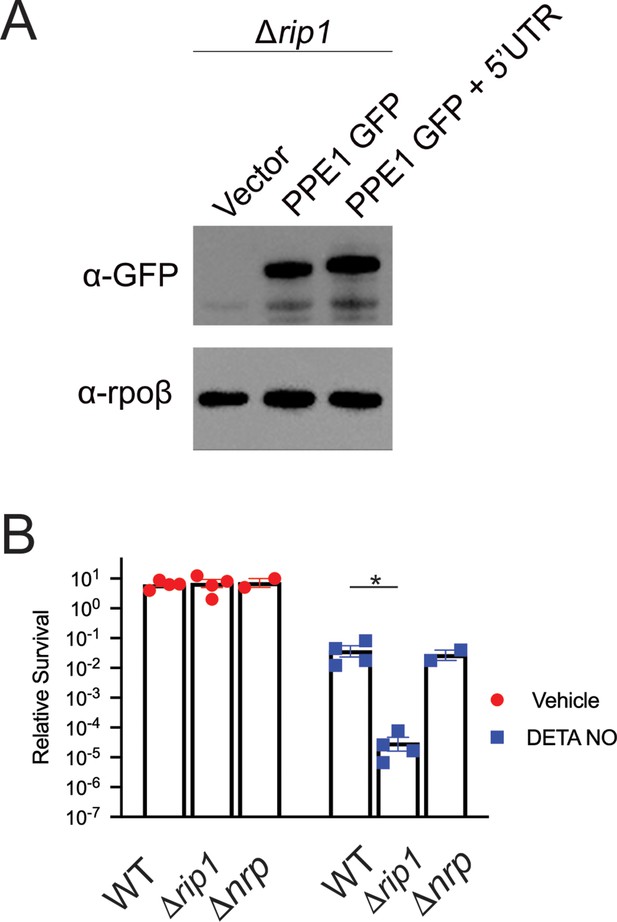
PPE1-5' UTR does not affect PPE1 protein expression and nrp loss is not sufficient for NO sensitivity.
(A) PPE1 is equivalently expressed independent of PPE-5′. Δrip1 cells transformed with vector, or plasmids encoding C-terminal GFP fused PPE1 ORF (PPE1 GFP), or C-terminal GFP fused PPE1 ORF + PPE1-5′ (PPE1 GFP + 5′UTR), both expressed from the hsp60 promoter, were lysed and blotted as described in Materials and methods. Upper panel: anti-GFP; lower panel: Rpoβ loading control. (B) Loss of chalkophore biosynthesis alone does not confer NO sensitivity. Colony-forming unit (CFU) counts of the indicated strains post treatment with vehicle or 200 µM diethylenetriamine nitric oxide (DETA-NO) for 3 days. Relative survival = CFU day 3 post treatment/CFU at day 0 for each condition. n = 4 biological replicates for WT and Δrip1; n = 2 biological replicates for Δnrp. *p<0.05 by unpaired t-test.
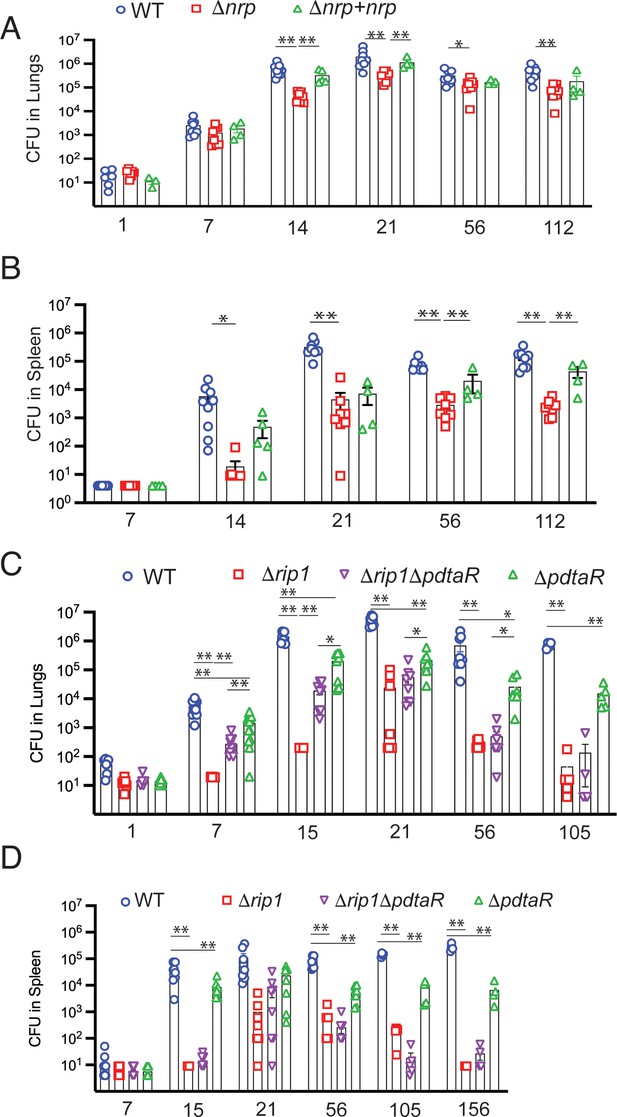
Rip1/PdtaS/PdtaR control of chalkophore biosynthesis controls early lung infection.
(A, B) Bacterial burden (colony-forming unit [CFU]) of the indicated strains in lungs (A) or spleens (B) of C57bl/6 mice infected with WT (blue circle), Δnrp (red square), or Δnrp +nrp (green triangle) at the indicated time points (days) post aerosol infection. For WT and Δnrp, n = 6 biological replicates on day 1 and n = 8 on all subsequent days. All p values calculated by unpaired t-test. (C, D) Bacterial burden (CFU) of the indicated strains in lungs (C) or spleens (D) of C57bl/6 mice infected with WT (blue circle), Δrip1 (red square), or Δrip1ΔpdtaR (purple inverted triangle) or ΔpdtaR (green triangle) at the indicated time points (days) post aerosol infection. For all strains, n = 5 biological replicates on day 1; n = 12 biological replicates on day 7; n = 8 biological replicates on days 15, 21, and 56; n = 4 biological replicates on days 105 and 156. For all panels, statistical significance by unpaired Welch’s t-test is represented as *p<0.05, **p<0.01.
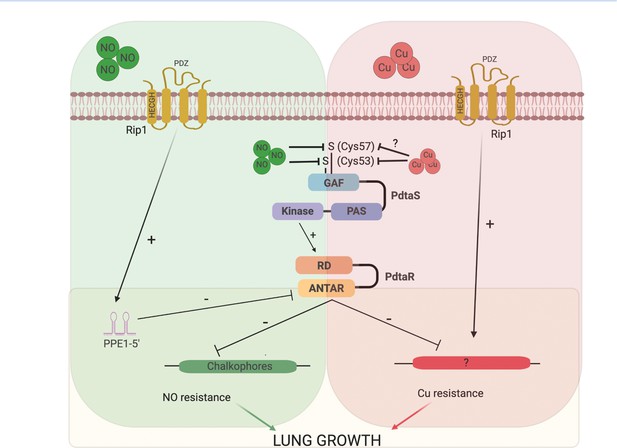
Model of the Rip1/PdtaS/PdtaR signaling circuit and its response to copper and nitric oxide.
NO and copper resistance is mediated by cell surface and cytoplasmic convergent signals that inactivate PdtaS/PdtaR. In basal conditions, PdtaS/R is constitutively active, and that activity holds virulence genes, including chalkophore biosynthesis, inactive. NO or copper directly inhibit PdtaS activity through direct sensing by PdtaS N-terminal GAF domain. NO shutoff of PdtaS kinase activity requires C53 and 57, whereas Cu shutoff requires C53 with the role of C57 unknown. Sensing of NO at the cell surface contributes a second signal to relief of PdtaS/R by inducing Rip1-dependent expression of a hairpin containing small RNA at the 5′ end of the PPE1 gene (PPE1-5′), which binds directly to the ANTAR domain containing PdtaR. These signals relieve PdtaS/R repression of virulence gene expression, including chalkophore biosynthesis, to activate NO resistance. Although our data clearly implicates the upstream Rip1/PdtaS/PdtaR cascade in Cu resistance, the downstream targets that mediate Cu resistance are not known (indicated by ?). Despite the high Cu affinity of the isonitriles produced by the nrp locus, these molecules are involved in the NO resistance arm of the pathway rather than Cu. Figure constructed with BioRender.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mycobacterium tuberculosis) | M.tb Erdman (WT, EG2) | Lab Stock | ATCC 35801 | Animal passaged |
| Strain, strain background (Escherichia coli) | DH5α | Lab Stock | ATCC SCC2197 | Plasmid maintenance strain |
| Strain, strain background (Escherichia coli) | EL350/ phAE87 | Lab Stock | Phage packaging strain | |
| Strain, strain background (Escherichia coli) | Rosetta 2 (DE3) | Millipore Sigma | Cat# 71397 | Recombinant protein expression strain |
| Strain, strain background (Mus musculus) female | C57BL/6J | Jackson Laboratory | Stock no: 000664; RRID:IMSR_JAX:000664 | |
| Strain, strain background (Mus musculus) female | B6;129P2-Nos2tm1Lau/J | Jackson Laboratory | Stock no: 002596; RRID:IMSR_JAX:002596 | |
| Gene (M. tuberculosis) | rip1 | ATCC 35801 | Erdman_3146 | |
| Gene (M. tuberculosis) | sigK | ATCC 35801 | Erdman_0488 | |
| Gene (M. tuberculosis) | sigL | ATCC 35801 | Erdman_0808 | |
| Gene (M. tuberculosis) | sigM | ATCC 35801 | Erdman_4291 | |
| Gene (M. tuberculosis) | sigD | ATCC 35801 | Erdman_3735 | |
| Gene (M. tuberculosis) | mymT | ATCC 35801 | Rv0186A | nt217703-217864 Erdman Genome |
| Gene (M. tuberculosis) | ctpV | ATCC 35801 | Erdman_1076 | |
| Gene (M. tuberculosis) | csoR | ATCC 35801 | Erdman_1074 | |
| Gene (M. tuberculosis) | nrp | ATCC 35801 | Erdman_0118 | |
| Gene (M. tuberculosis) | ppe1 | ATCC 35801 | Erdman_0113 | |
| Gene (M. tuberculosis) | rv0097 | ATCC 35801 | Erdman_0114 | |
| Gene (M. tuberculosis) | fcoT | ATCC 35801 | Erdman_0115 | |
| Gene (M. tuberculosis) | fadD10 | ATCC 35801 | Erdman_0116 | |
| Gene (M. tuberculosis) | rv0100 | ATCC 35801 | Erdman_0117 | |
| Gene (M. tuberculosis) | pdtaS | ATCC 35801 | Erdman_3533 | |
| Gene (M. tuberculosis) | pdtaR | ATCC 35801 | Erdman_1787 | |
| Genetic reagent (M. tuberculosis) | Δrip1 | Sklar et al., 2010 | MGM3206; rip1 KO | Chromosomal deletion of Erdman_3146 nt5-1212 by double crossover recombination followed by marker excision by LoxP recombination |
| Genetic reagent (M. tuberculosis) | ΔsigD | Schneider et al., 2014 | JSS0005; sigD KO | Chromosomal deletion of Erdman_3735 nt6-622 by double crossover recombination |
| Genetic reagent (M. tuberculosis) | ΔsigK | Sklar et al., 2010 | MGM3259; sigK KO | |
| Genetic reagent (M. tuberculosis) | ΔsigL | Sklar et al., 2010 | MGM3254; sigL KO | |
| Genetic reagent (M. tuberculosis) | ΔsigM | Sklar et al., 2010 | MGM3260, sigM KO | |
| Genetic reagent (M. tuberculosis) sigL sigM KO | ΔsigLΔsigM | Sklar et al., 2010 | MGM3255; sigL sigM KO | |
| Genetic reagent (M. tuberculosis) | ΔsigKΔsigLΔsigM | Schneider et al., 2014 | MGM3256; sigK sigL sigM KO | |
| Genetic Reagent (M. tuberculosis) | ΔsigKΔsigL | Sklar et al., 2010 | MGM3261; sigK sigL KO | |
| Genetic reagent (M. tuberculosis) | ΔsigKΔsigM | Sklar et al., 2010 | MGM3283; sigK sigM KO | |
| Genetic reagent (M. tuberculosis) | ΔsigKΔsigD | This study, available from corresponding author | sigK::loxP sigD::hygR; MGM3288 | Chromosomal deletion of Erdman_3735 nt6-622 by double crossover recombination MGM3259 background |
| Genetic reagent (M. tuberculosis) | ΔsigLΔsigD | This study, available from corresponding author | sigL::loxP sigD::hygR; MGM3289 | Chromosomal deletion of Erdman_3735 nt6-622 by double crossover recombination MGM3254 background |
| Genetic reagent (M. tuberculosis) | ΔsigMΔsigD | This study, available from corresponding author | sigM::loxP sigD::hygR; MGM3290 | Chromosomal deletion of Erdman_3735 nt6-622 by double crossover recombination MGM3260 background |
| Genetic reagent (M. tuberculosis) | ΔctpV | This study, available from corresponding author | ctpV::hygR | Chromosomal deletion of nt 175-2136 of Erdman_1076 by double crossover recombination |
| Genetic reagent (M. tuberculosis) | ΔcsoR | This study, available from corresponding author | csoR::hygR | Chromosomal deletion of nt 1-360 of Erdman_1074 by double crossover recombination |
| Genetic reagent (M. tuberculosis) | ΔmymT | This study, available from corresponding author | mymT::hygR | Chromosomal deletion of nt 1-162 of mymT by double crossover recombination |
| Genetic reagent (M. tuberculosis) | Δrip1ΔctpV | This study, available from corresponding author | rip1::loxP ctpV::hygR | Chromosomal deletion of nt 175-2136 of Erdman_1076 by double crossover recombination MGM3206 background |
| Genetic reagent (M. tuberculosis) | Δrip1ΔcsoR | This study, available from corresponding author | rip1::loxP csoR::hygR | Chromosomal deletion of nt 1-360 of Erdman_1074 by double crossover recombination MGM3206 background |
| Genetic reagent (M. tuberculosis) | Δrip1ΔmymT | This study, available from corresponding author | rip1::loxP mymT::hygR | Chromosomal deletion of nt 1-162 of mymT by double crossover recombination MGM 3206 background |
| Genetic reagent (M. tuberculosis) | ΔpdtaS | This study, available from corresponding author | pdtaS::hygR | Chromosomal deletion of nt1-1506 of Erdman_3533 by double crossover recombination |
| Genetic reagent (M. tuberculosis) | ΔpdtaS + Vector | This study, available from corresponding author | pdtaS::hygR:pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | ΔpdtaR | This study, available from corresponding author | pdtaR::hygR | Chromosomal deletion of nt 1-576 of Erdman_1787 by double crossover recombination |
| Genetic reagent (M. tuberculosis) | ΔpdtaR + vector | This study, available from corresponding author | pdtaR::hygR:pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δnrp | This study, available from corresponding author | nrp::hygR | Chromosomal deletion of nt 21-7515 of Edman_0118 by double crossover recombination |
| Genetic reagent (M. tuberculosis) | Δnrp + vector | This study, available from corresponding author | nrp::hygR: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δnrp + WT nrp | This study, available from corresponding author | nrp::hygR: WT nrp | WT copy of nrp integrated at AttB, Erdman_0116 promoter |
| Genetic reagent (M. tuberculosis) | Δrip1ΔpdtaS | This study, available from corresponding author | rip1::loxP pdtaS::hygR | Chromosomal deletion of nt1-1506 of Erdman_3533 by double crossover recombination MGM3206 background |
| Genetic reagent (M. tuberculosis) | Δrip1ΔpdtaS + vector | This study, available from corresponding author | rip1::loxP pdtaS::hygR: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1ΔpdtaR | This study, available from corresponding author | rip1::loxP pdtaR::hygR | Chromosomal deletion of nt 1-576 of Erdman_1787 by double crossover recombination MGM3206 background |
| Genetic reagent (M. tuberculosis) | Δrip1ΔpdtaR + vector | This study, available from corresponding author | rip1::loxP pdtaR::hygR: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1ΔpdtaR + WT pdtaR | This study, available from corresponding author | rip1::loxP pdtaR::hygR: WT pdtaR HA | WT copy of pdtaR integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1ΔpdtaR + D65A pdtaR | This study, available from corresponding author | rip1::loxP pdtaR::hygR: D65A pdtaR HA | D65A variant of pdtaR integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1Δnrp | This study, available from corresponding author | rip1::loxP nrp::hygR | Chromosomal deletion of nt 21-7515 of Edman_0118 by double crossover recombination MGM3206 background |
| Genetic reagent (M. tuberculosis) | WT + vector | Makinoshima and Glickman, 2005 | EG2: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | WT + vector | Makinoshima and Glickman, 2005 | EG2:pMV261 | Transformed with empty pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1 + vector | Sklar et al., 2010 | Δrip1: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1 + vector | Sklar et al., 2010 | rip1::loxP: pMV261 | Transformed with empty pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1 + WT Rip1 | Sklar et al., 2010 Makinoshima and Glickman, 2005 | rip1::loxP: WT rip1 | Transformed with WT rip1 pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1 + H21A Rip1 | Makinoshima and Glickman, 2005 | rip1::loxP: H21A rip1 | Transformed with H21A variant rip1 pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1: PPE1 ORF pHSP60 | This study, available from corresponding author | rip1::loxP: PPE1 GFP ORF only | Transformed with PPE1 GFP ORF only pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1: PPE1 ORF + 5' pHSP60 | This study, available from corresponding author | rip1::loxP: PPE1 GFP ORF + 5' UTR pHSP60 | Transformed with PPE1 GFP + 5' UTR pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1: Rv0097-nrp pHSP60 | This study, available from corresponding author | rip1::loxP: Rv0097-nrp | Transformed with Rv0097-nrp pMV261K episomal plasmid |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS F37X | This study, available from corresponding author | rip1::loxP pdtaS F37X | Isolated spontaneous Erdman_3533 variant in MGM3206 background |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS F37X + vector | This study, available from corresponding author | rip1::loxP pdtaS F37X: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS F37X + pdtaS | This study, available from corresponding author | rip1::loxP pdtaS F37X: WT pdtaS 10xHis | WT pdtaS 10xHis integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS F37X + pdtaS V54F | This study, available from corresponding author | rip1::loxP pdtaS F37X: V54F pdtaS 10xHis | V54F variant of Erdman_3533 10xHis integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS F37X + pdtaS H302Q/H303Q | This study, available from corresponding author | rip1::loxP pdtaS F37X: H302Q/H303Q pdtaS 10xHis | H302Q/H303Q variant of Erdman_3533 10xHis integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS F37X + pdtaS G443A/G445A | This study, available from corresponding author | rip1::loxP pdtaS F37X: G443A/G445A pdtaS 10xHis | G443A/G445A variant of Erdman_3533 10xHis integrated at AttB |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS V54F | This study, available from corresponding author | rip1::loxP pdtaS V54F | Isolated spontaneous Erdman_3533 variant in MGM3206 background |
| Genetic reagent (M. tuberculosis) | Δrip1 pdtaS V54F + vector | This study, available from corresponding author | rip1::loxP pdtaS V54F: pMV306K | Empty pMV306K integrated at AttB |
| Genetic reagent (E. coli) | Rosetta 2 DE3: pET WT PdtaS | This study, available from corresponding author | Rosetta 2 DE3: pET WT PdtaS | T7lac recombinant expression of C-terminally 10xHis tagged protein |
| Genetic reagent (E. coli) | Rosetta 2 DE3: pET WT PdtaS kinase domain | This study, available from corresponding author | Rosetta 2 DE3: pET WT PdtaS kinase domain | T7lac recombinant expression of C-terminally 10xHis tagged protein |
| Genetic reagent (E. coli) | Rosetta 2 DE3: pET C53A PdtaS | This study, available from corresponding author | Rosetta 2 DE3: pET C53A PdtaS | T7lac recombinant expression of C-terminally 10xHis tagged protein |
| Genetic reagent (E. coli) | Rosetta 2 DE3: pET C57A PdtaS | This study, available from corresponding author | Rosetta 2 DE3: pET C57A PdtaS | T7lac recombinant expression of C-terminally 10xHis tagged protein |
| Genetic reagent (E. coli) | Rosetta 2 DE3: pET WT PdtaR | This study, available from corresponding author | Rosetta 2 DE3: pET WT PdtaR | T7lac recombinant expression of C-terminally 10xHis tagged protein |
| Genetic reagent (E. coli) | Rosetta 2 DE3: pET D65A PdtaR | This study, available from corresponding author | Rosetta 2 DE3: pET D65A PdtaR | T7lac recombinant expression of C-terminally 10xHis tagged protein |
| Recombinant DNA reagent | pET SUMO (plasmid) | Reverter and Lima, 2009 | pET SUMO | T7lac promoter E. coli expression vector N-terminal 10xHIS Sumo Fusion Protein |
| Recombinant DNA reagent | pET (plasmid) | This study, available from corresponding author | pET | T7lac promoter E. coli expression vector |
| Recombinant DNA reagent | PdtaS (plasmid) | This study, available from corresponding author | pET WT PdtaS 10XHIS | nt 1-1503 of Erdman_3533 fused N-terminal to Enterkinase cleavage site followed by 10xHis tag |
| Recombinant DNA reagent | PdtaS C53A (plasmid) | This study, available from corresponding author | pET C53A PdtaS 10XHIS | nt 1-1503 of Erdman_3533 C53A variant fused N-terminal to Enterkinase cleavage site followed by 10xHis tag |
| Recombinant DNA reagent | PdtaS C57A (plasmid) | This study, available from corresponding author | pET C57A PdtaS 10XHIS | nt 1-1503 of Erdman_3533 C57A variant fused N-terminal to Enterkinase cleavage site followed by 10xHis tag |
| Recombinant DNA reagent | PdtaS kinase domain (plasmid) | This study, available from corresponding author | pET WT PdtaS kinase domain 10XHIS | nt 678-1503 of Erdman_3533 variant fused N-terminal to Enterkinase cleavage site followed by 10xHis tag |
| Recombinant DNA reagent | PdtaR D65A (plasmid) | This study, available from corresponding author | pET Sumo WT PdtaR | N-terminal 10xHIS Smt3 fused to nt 1-615 of Erdman_1787 |
| Recombinant DNA reagent | PdtaR D65A (plasmid) | This study, available from corresponding author | pET Sumo D65A PdtaR | N-terminal 10xHIS Smt3 fused to nt 1-615 of Erdman_1787 D65A variant |
| Recombinant DNA reagent | Vector (plasmid) | Sklar et al., 2010 Makinoshima and Glickman, 2005 | pMV261K | Episomal M.tb plasmid |
| Recombinant DNA reagent | Vector (plasmid) | Sklar et al., 2010 Makinoshima and Glickman, 2005 | pMV306K | AttB integrating M.tb plasmid |
| Recombinant DNA reagent | pMV261K GFP (plasmid) | This study, available from corresponding author | pMV261K GFP | Episomal M.tb plasmid for C-terminal GFP fusion constructs |
| Recombinant DNA reagent | WT rip1 (plasmid) | Makinoshima and Glickman, 2005 | pHMG121 | |
| Recombinant DNA reagent | H21A rip1 (plasmid) | Makinoshima and Glickman, 2005 | pHMG141 | |
| Recombinant DNA reagent | pdtaS (plasmid) | This study, available from corresponding author | pMV306K WT PdtaS 10xHis | nt 1-1503 of Erdman_3533 tagged at C-term with 10x His |
| Recombinant DNA reagent | pdtaS V54F (plasmid) | This study, available from corresponding author | pMV306K V54F PdtaS 10xHis | nt 1-1503 of Erdman_3533 tagged at C-term with 10x His V54F variant |
| Recombinant DNA reagent | pdtaS H302Q/H303Q (plasmid) | This study, available from corresponding author | pMV306K H302Q/H303Q PdtaS 10xHis | nt 1-1503 of Erdman_3533 tagged at C-term with 10x His H302Q/H303Q variant |
| Recombinant DNA reagent | pdtaS G443A/G445A (plasmid) | This study, available from corresponding author | pMV306K G443A/H445A PdtaS 10xHIs | nt 1-1503 of Erdman_3533 tagged at C-term with 10x His G443A/G445A variant |
| Recombinant DNA reagent | pdtaR (plasmid) | This study, available from corresponding author | pMV306K WT PdtaR HA | nt 1-615 of Edman_1787 tagged at C-term with HA epitope |
| Recombinant DNA reagent | pdtaR D65A (plasmid) | This study, available from corresponding author | pMV306K D65A PdtaR HA | nt 1-615 of Edman_1787 tagged at C-term with HA epitope D65A variant |
| Recombinant DNA reagent | PPE1 ORF pHSP60 (plasmid) | This study, available from corresponding author | pMV261K ORF PPE1 GFP | nt 1-1389 of Erdman_0113 tagged at C-term with GFP |
| Recombinant DNA reagent | PPE1 ORF + 5' pHSP60 (plasmid) | This study, available from corresponding author | pMV261K ORF PPE1 GFP + 5'PPE1 | nt -222-1389 of Erdman_0113 tagged at C-term with GFP |
| Recombinant DNA reagent | Rv0097-nrp pHRP60 (plasmid) | This study, available from corresponding author | pMV261K Rv0097-nrp | nt 1 of Erdman_0114 to nt 7539 of Erdman_0118 |
| Recombinant DNA reagent | WT nrp (plasmid) | This study, available from corresponding author | pMV306 WT nrp | nt 1-96 of Erdman_0116 (predicted TSS Wadsworth) + nt 1-7539 of Erdman_0118 |
| Recombinant DNA reagent | pmsg360Hyg (plasmid) | Sklar et al., 2010 Makinoshima and Glickman, 2005 | pmsg360Hyg | Phage packaging vector for hygR allelic exchange |
| Recombinant DNA reagent | pmsg360Zeo (plasmid) | Sklar et al., 2010 Makinoshima and Glickman, 2005 | pmsg360Zeo | Phage packaging vector for zeoR allelic exchange |
| Recombinant DNA reagent | pmsg318-2 (plasmid) | Sklar et al., 2010 Makinoshima and Glickman, 2005 | pmsg318-2 | M.tb Cre recombinase expression vector |
| Recombinant DNA reagent | pmsg360Hyg CtpV flanks (plasmid) | This study, available from corresponding author | ΔctpV targeting vector | Hyg cassette flanked by 616 bp 5' CtpV nt 175 + 600 bp 3' CtpV nt 2136 |
| Recombinant DNA reagent | pmsg360 Hyg CsoR flanks (plasmid) | This study, available from corresponding author | ΔcsoR targeting vector | Hyg cassette flanked by 600 bp 5' CsoR start codon + 600 bp 3' of CsoR stop codon |
| Recombinant DNA reagent | pmsg360 Hyg MymT flanks (plasmid) | This study, available from corresponding author | ΔmymT targeting vector | Hyg cassette flanked by 600 bp 5' MymT start codon + 600 bp 3' of MymT stop codon |
| Recombinant DNA reagent | pmsg360 Hyg pdtaS flanks (plasmid) | This study, available from corresponding author | ΔpdtaS targeting vector | Hyg cassette flanked by 600 bp 5' PdtaS start codon + 579 bp 3' PdtaS stop codon |
| Recombinant DNA reagent | pmsg360 Hyg pdtaR flanks (plasmid) | This study, available from corresponding author | ΔpdtaR targeting vector | Hyg cassette flanked by 600 bp 5' PdtaR start codon + 647 bp 3' PdtaR nt 576 |
| Recombinant DNA reagent | pmsg360 Hyg nrp Flanks (plasmid) | This study, available from corresponding author | Δnrp targeting vector | Hyg cassette flanked by 621 bp 5' nrp nt 21+ 621 bp 3' nrp nt 7515 |
| Sequence-based reagent | oSigA-1 | Integrated DNA Technologies | qPCR primer | cgtcttcatcccagacgaaat |
| Sequence-based reagent | oSigA-2 | Integrated DNA Technologies | qPCR primer | cgacgaagaccacgaagac |
| Sequence-based reagent | oCtpV-1 | Integrated DNA Technologies | qPCR primer | gtgtcccatgttcgaggtcaa |
| Sequence-based reagent | oCtpV-2 | Integrated DNA Technologies | qPCR primer | gtcaatgttcttcggtgcttac |
| Sequence-based reagent | oLpqS-1 | Integrated DNA Technologies | qPCR primer | gcatcgagttgtccaccag |
| Sequence-based reagent | oLpqS-2 | Integrated DNA Technologies | qPCR primer | tcaatgtggctcacccaaac |
| Sequence-based reagent | oMymT-1 | Integrated DNA Technologies | qPCR primer | gggtgatacgaatgacgaacta |
| Sequence-based reagent | oMymT-2 | Integrated DNA Technologies | qPCR primer | acagtggcatgggacttc |
| Sequence-based reagent | o2625c-F | Integrated DNA Technologies | qPCR primer | tcttgatcgcgttgggattg |
| Sequence-based reagent | o2625c-R | Integrated DNA Technologies | qPCR primer | cccggcgaattgatgtagag |
| Sequence-based reagent | oDesR-F | Integrated DNA Technologies | qPCR primer | tctgatcctcacgtcctacac |
| Sequence-based reagent | oDesR-R | Integrated DNA Technologies | qPCR primer | agcgcccacatctttgac |
| Sequence-based reagent | oHspX-F | Integrated DNA Technologies | qPCR primer | gaattcgcgtacggttccttc |
| Sequence-based reagent | oHspX-R | Integrated DNA Technologies | qPCR primer | gccaccgacacagtaagaatg |
| Sequence-based reagent | oPdtaS_C53A-F | Integrated DNA Technologies | PCR primer | gcgacgacggtgtcctggtggcggttgcg |
| Sequence-based reagent | oPdtaS_C53A-R | Integrated DNA Technologies | PCR primer | cgcaaccgccaccaggacaccgtcgtcgc |
| Sequence-based reagent | oPdtaS_C57A-F | Integrated DNA Technologies | PCR primer | gcgcaagcccggccgaacaccgggccgacg |
| Sequence-based reagent | oPdtaS_C57A-R | Integrated DNA Technologies | PCR primer | cgtcggcccggtgttcggccgggcttgcgc |
| Sequence-based reagent | oPdtaR_D65A-F | Integrated DNA Technologies | PCR primer | gtgatcatggccgtgaaga |
| Sequence-based reagent | oPdtaR_D65A-R | Integrated DNA Technologies | PCR primer | tcttcacggccatgatcac |
| Sequence-based reagent | oPdtaS_H302Q/H303Q-F | Integrated DNA Technologies | PCR primer | gggaaatccagcagcgggtt |
| Sequence-based reagent | oPdtaS_H302Q/H303Q-R | Integrated DNA Technologies | PCR primer | aacccgctgctggatttccc |
| Sequence-based reagent | oPdtaS_G443A/G445A-F | Integrated DNA Technologies | PCR primer | acgacgcgcttgctctgccg |
| Sequence-based reagent | oPdtaS_G443A/G445A-F | Integrated DNA Technologies | PCR primer | cggcagagcaagcgcgtcgt |
| Sequence-based reagent | Sense5'PPE1-F | Integrated DNA Technologies | PCR primer; in vitro transcription | taatacgactcactatagggggccgactaacaccgcgg |
| Sequence-based reagent | Sense5'PPE1-R | Integrated DNA Technologies | PCR primer; in vitro transcription | ggtttgctcaagccaggc |
| Sequence-based reagent | Rv3864-F | Integrated DNA Technologies | PCR primer; in vitro transcription | taatacgactcactataggcaaaaaattcgtgcaccaacc |
| Sequence-based reagent | Rv3864-R | Integrated DNA Technologies | PCR primer; in vitro transcription | tttccttacgctcgccgt |
| Sequence-based reagent | AntiDNAK-F | Integrated DNA Technologies | PCR primer; in vitro transcription | gatccacctagttctagaatggctcgtgcggtcg |
| Sequence-based reagent | AntiDNAK-R | Integrated DNA Technologies | PCR primer; in vitro transcription | taatacgactcactatagggggcgtcattgaagtaggcg |
| Antibody | Anti E. coli RNA polymerase B antibody (α-Rpoβ) | BioLegend | 663903 (RRID:AB_2564414) | 1:10,000 dilution |
| Antibody | Anti-HA.11 epitope tag antibody (α-HA) | BioLegend | 901513 (RRID:AB_2565335) | 1:1000 dilution |
| Antibody | α-CarD (Rabbit) | Pocono Rabbit Farm | Stallings et al., 2009 | 1:10,000 dilution |
| Antibody | α-MymT (Rabbit) | Gold et al., 2008 | 1:1000 dilution | |
| Antibody | Anti-GFP(Rabbit) Antibody (α-GFP) | Rockland | 600-401-215L (RRID:AB_2612813) | 1:1000 dilution |
| Chemical compound, drug | Spermine NONOate | Millipore Sigma | 567703 | |
| Chemical compound, drug | Diethylenetriamine/nitric oxide adduct (DETA-NO) | Millipore Sigma | D185 | |
| Chemical compound, drug | Hydrogen peroxide, 30% | Fisher Scientific | H325-100 | |
| Chemical compound, drug | Sodium hydrosulfide hydrate | Fisher Scientific | AC296200250 | |
| Commercial assay, kit | In-Fusion HD Cloning Kit | Takara Bio USA | 639650 | |
| Commercial assay, kit | Maxima H Minus cDNA Synthesis Master Mix, with dsDNase | Thermo Fisher Scientific | M1681 | |
| Commercial assay, kit | DyNAmo Flash SYBR Green qPCR Kit | Thermo Fisher Scientific | F415F | |
| Commercial assay, kit | HiScribe T7 High Yield RNA Synthesis Kit | New England Biolabs | E2040S | |
| Commercial assay, kit | Ribo-Zero Magnetic Bacterial Kit | Epicentre | MRZB12424 | |
| Commercial assay, kit | TruSeq Stranded Total RNA kit | Illumina | 20020599 | |
| Commercial assay, kit | KAPA Hyper Prep Kit | Roche | 7962312001 | |
| Commercial assay, kit | TruSeq SBS Kit v3 | Illumina | FC-401-3002 | |
| Commercial assay, kit | GeneJET RNA Purification Kit | Fisher Scientific | FERK0731 | |
| Commercial assay, kit | TURBO DNA-free kit | Fisher Scientific | AM1907 | |
| Peptide, recombinant protein | Phusion High Fidelity Polymerase | Fisher Scientific | F530L | |
| Commercial assay, kit | GeneJET Plasmid Miniprep Kit | Fisher Scientific | FERK0503 | |
| Commercial assay, kit | NanoTemper Technologies Inc PROTEIN LABELING KIT RED-NHS (MOL011) | Fisher Scientific | NC1491187 | |
| Commercial assay, kit | NanoTemper Technologies Inc Standard capillaries | Fisher Scientific | NC1408770 | |
| Software, algorithm | ViennaRNA Web Services | http://www.viennarna.at/forna/; RRID:SCR_008550 | ||
| Software, algorithm | fastqc | http://www.bioinformatics.babraham.ac.uk/projects/fastqc | RRID:SCR_014583 | |
| Software, algorithm | bwa mem | Li et al., 2009 | RRID:SCR_010910 | |
| Software, algorithm | samtools | Li et al., 2009 | RRID:SCR_002105 | |
| Software, algorithm | Bioconductor Rsubread package | Liao et al., 2019 | RRID:SCR_016945 | |
| Software, algorithm | DESeq2 R package | Love et al., 2014 | RRID:SCR_015687 | |
| Software, algorithm | R | The R Project for Statistical Computing; https://www.R-project.org/ | RRID:SCR_001905 | |
| Software, algorithm | GATK | Broad Institute; https://gatk.broadinstitute.org/hc/en-us | RRID:SCR_001876 | |
| Software, algorithm | pheatmap | https://www.rdocumentation.org/packages/pheatmap/versions/0.2/topics/pheatmap | RRID:SCR_016418 | |
| Other | NUPAGE 4-12% BT Gel | Fisher Scientific | NPO312BOX/NPO336BOX | |
| Other | Protran Nitrocellulose Hybridization Transfer Membrane | Perkin Elmer | NBA08C001EA | |
| Other | Immun-Blot PVDF Membrane | Bio-Rad | 1620177 | |
| Other | NanoTemper Technologies Premium Capillaries | Fisher Scientific | NC1408772 | |
| Other | HisProbe-HRP Conjugate | Thermo Fisher Scientific | 15165 |
Additional files
-
Supplementary file 1
Bacterial strains used in this work with strain ID, genotype, and relevant features and reference.
- https://cdn.elifesciences.org/articles/65351/elife-65351-supp1-v1.xlsx
-
Supplementary file 2
Plasmids used in this work.
- https://cdn.elifesciences.org/articles/65351/elife-65351-supp2-v1.xlsx
-
Supplementary file 3
Oligonucleotides used in this work.
- https://cdn.elifesciences.org/articles/65351/elife-65351-supp3-v1.xlsx
-
Supplementary file 4
RNAseq count data from RNA sequencing of WT, Δrip1, ΔpdtaR, and Δrip1/ ΔpdtaR exposed to vehicle of diethylenetriamine nitric oxide (DETA-NO) in triplicate.
Each cell contains the normalized counts for each gene in each replicate. Please see Materials and methods for details.
- https://cdn.elifesciences.org/articles/65351/elife-65351-supp4-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/65351/elife-65351-transrepform-v1.docx






