A mobile genetic element increases bacterial host fitness by manipulating development
Figures
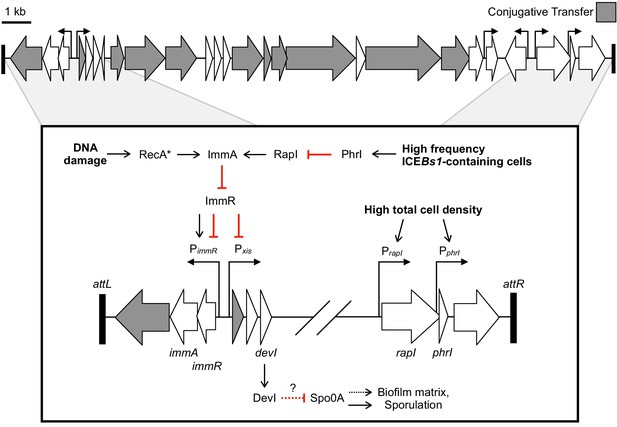
Genetic map and regulatory pathways of ICEBs1.
Genes are represented by horizontal block arrows indicating the direction of transcription. Vertical right-angle arrows mark the positions of promoters, and the arrowhead indicates the direction of transcription. Genes known to be involved in the conjugative life cycle of ICEBs1 are shaded in gray. The 60 bp direct repeats that mark the ends of ICEBs1 are shown as black rectangles. (Inset) A partial genetic map that highlights factors involved in the regulation of ICEBs1. The major promoter Pxis drives expression of most genes in ICEBs1. Pxis is repressed by the ICE-encoded repressor ImmR. Repression is relieved when ImmR is cleaved by the protease ImmA, and proteolytic cleavage is stimulated by activated RecA (RecA*) in response to DNA damage, or, independently by the cell signaling regulator RapI. RapI is made when cells are crowded by potential recipients, but repressed by the ICE-encoded secreted peptide PhrI if the neighboring cells already contain a copy of ICEBs1. devI (formerly ydcO) is the third open-reading frame downstream of Pxis. DevI inhibits sporulation and expression of biofilm matrix genes, likely by inhibiting Spo0A (directly or indirectly). In the genetic pathways, black arrows indicate activation and red T-bars indicate inhibition.
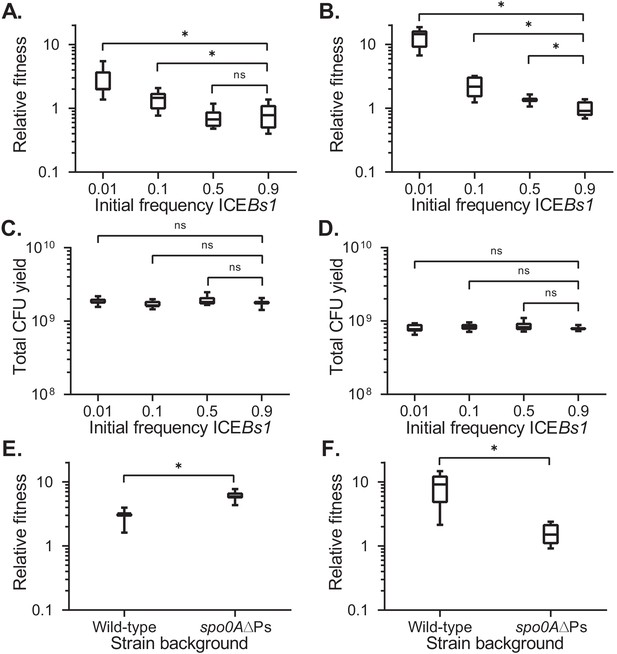
Fitness of ICEBs1-containing cells relative to ICEBs1-cured cells during development.
The fitness of ICEBs1-containing cells (JMJ593) relative to ICEBs1-cured cells (JMJ550) was measured by competition during biofilm growth (A) or growth on sporulation medium (B). Total growth yields in biofilms (C) and on sporulation medium (D) were determined by counting CFUs derived from both cells and spores. The fitness of ICEBs1-containing cells relative to ICEBs1-cured cells was compared between the wild-type strain background (JMJ593 vs. JMJ550) and in the sporulation-deficient spo0A∆Ps background (JMJ788 vs. JMJ786) by competition during biofilm growth (E) or growth on sporulation medium (F). ICEBs1-containing cells were inoculated at an initial frequency of 0.01. Data shown are pooled from three independent experiments. A total of nine populations (biological replicates) were analyzed per condition. Boxes extend from the lower to upper quartiles of the data, and the middle line indicates the median fitness. Whiskers indicate the range of the fitness measurements. Asterisks indicate a p-value<0.05 (two-tailed T-test, unequal variance). Exact p-values: (A) 3.7 × 10−5, 8.8 × 10−3, 6.0 × 10−1; (B) 4.9 × 10−11, 1.6 × 10−4, 7.1 × 10−3; (C) 3.7 × 10−1, 4.2 × 10−1, 2.2 × 10−2; (D) 8.9 × 10−1, 1.9 × 10−1, 1.7 × 10−1; (E) 5.5 × 10−6; (F) 2.4 × 10−5.
-
Figure 2—source data 1
Frequency-dependent fitness.
Counts of ICEBs1-containing and ICEBs1-cured cells at start and end of competitions on biofilm and sporulation growth medium.
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig2-data1-v2.xlsx
-
Figure 2—source data 2
Fitness dependence on sporulation.
Counts of ICEBs1-containing and ICEBs1-cured cells at start and end of competitions for strains of wild-type or sporulation-deficient backgrounds.
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig2-data2-v2.xlsx
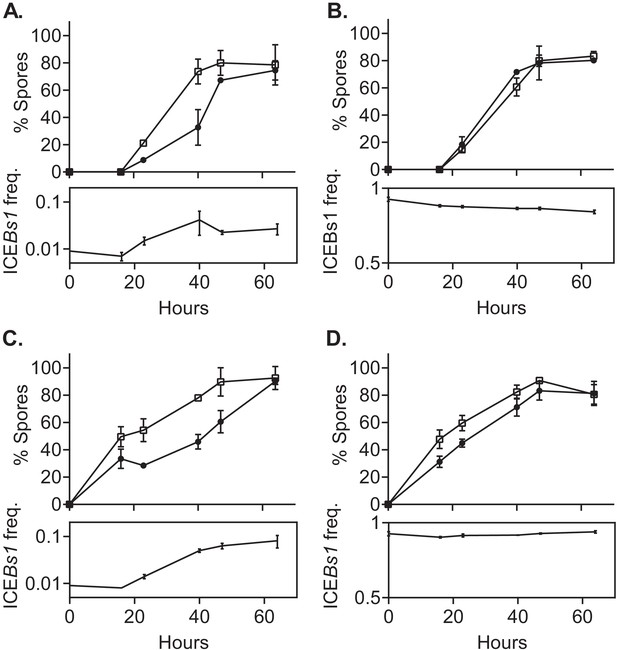
ICEBs1-containing cells delay sporulation in a frequency-dependent manner.
ICEBs1-containing cells (JMJ593, black circles) were mixed with ICEBs1-cured cells (JMJ550, open squares) at an initial frequency of 0.01 and spotted onto biofilm growth medium (A) or sporulation medium (C). Mixtures with ICEBs1-containing cells at an initial frequency of 0.9 were also spotted onto biofilm medium (B) and sporulation medium (D). Biofilms and colonies were harvested at the indicated times to determine the fraction of CFUs derived from heat-resistant spores for both the ICEBs1-containing and the ICEBs1-cured cells. Boxes below each graph indicate the frequency of ICEBs1-containing CFUs at each timepoint. Data shown are the average from two populations (biological replicates) per timepoint with error bars indicating the standard deviation. A representative experiment is shown.
-
Figure 3—source data 1
Sporulation timing during competitions.
Raw counts and calculated frequencies of spore-derived CFUs and total CFUs measured at different times during competition of ICEBs1-containing and ICEBs1-cured cells.
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig3-data1-v2.xlsx
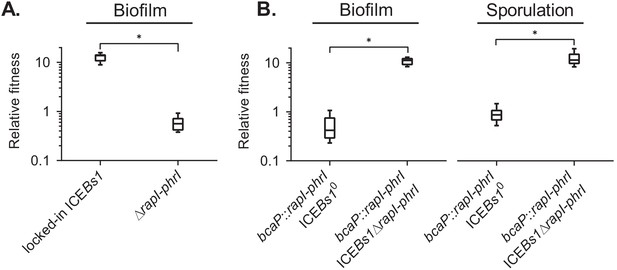
The ICEBs1 cell-cell signaling genes, rapI-phrI, are necessary but not sufficient to confer a selective advantage.
(A) The fitness of cells containing locked-in ICEBs1 (JMJ646) or an isogenic rapI-phrI mutant (JMJ686) was measured relative to ICEBs1-cured cells (JMJ550) during biofilm competitions. The ICEBs1-containing cells were started at a frequency of 0.01. (B) The fitness of cells containing rapI-phrI alone (JMJ576) or cells containing both rapI-phrI and locked-in ICEBs1∆rapI-phrI (JMJ785) was measured relative to ICEBs1-cured cells (JMJ714) during biofilm and sporulation medium competitions. JMJ576 and JMJ785 were started at a frequency of 0.01. Data shown are pooled from three independent experiments with a total of nine populations (biological replicates) analyzed per strain mixture. Boxes extend from the lower to upper quartiles, and the middle line indicates the median fitness. Whiskers indicate the range of the fitness measurements. Asterisks indicate a p-value<0.05 (two-tailed T-test, unequal variance). Exact p-values: (A) 1.3 × 10−12; (B) 2.1 × 10−8, 4.6 × 10−2.
-
Figure 4—source data 1
Fitness dependence on rapI-phrI.
Counts of ICEBs1-containing (wild-type or rapI-phrI mutant) and ICEBs1-cured cells at start and end of competitions. Counts of cells containing ectopically expressed rapI-phrI and wild-type cells at start and end of competitions.
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig4-data1-v2.xlsx
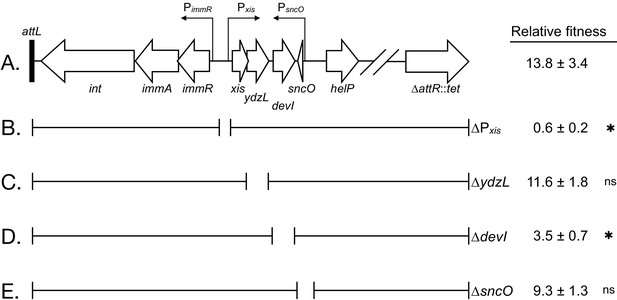
Effects of deletions in ICEBs1 on host fitness.
(A) Abbreviated genetic map of locked-in-ICEBs1 showing genes as open block arrows, promoters as thin right-angle arrows, and the left attachment site (attL) as a black bar. (B to E) Brackets under the map of ICEBs1 indicate regions contained in isogenic derivatives of locked-in-ICEBs1. Open spaces represent regions deleted. The fitness of strains containing locked-in-ICEBs1 (JMJ646) and its derivatives (∆Pxis, JMJ662; ∆ydzL, JMJ704; ∆devI, JMJ703; ∆sncO, JMJ688) is indicated at the right. Fitness was measured relative to ICEBs1-cured cells (JMJ550) during competitions in biofilms. ICEBs1-containing cells were started at a frequency of 0.01. Data shown are the median ± standard deviation from at least three independent experiments (at least nine total biological replicates per strain mixture), with the exception of JMJ688 which was measured only in one experiment (three total biological replicates). Asterisks indicate a statistically significant difference in fitness compared to JMJ646 (p-value<0.05, two-tailed T-test, unequal variance). Exact p-values: (B) 2.3 × 10−16; (C) 4.5 × 10−1; (D) 6.3 × 10−12; (E) 6.2 × 10−2.
-
Figure 5—source data 1
Effects of gene deletions on ICEBs1 fitness.
Counts of ICEBs1-containing (wild-type or deletion mutant) cells and ICEBs1-cured cells at the start and end of competitions.
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig5-data1-v2.xlsx
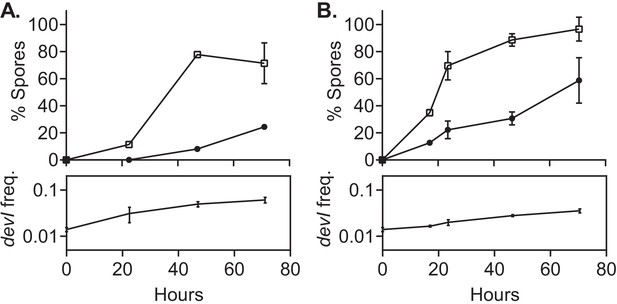
devI alone is sufficient to inhibit sporulation and provide a selective advantage.
ICEBs1-cured cells that constitutively express devI (JMJ725, black circles) were mixed at an initial frequency of 0.01 with ICEBs1-cured cells containing an empty expression construct (JMJ727, open squares). The mixture was spotted onto biofilm growth medium (A) or sporulation medium (B). Biofilms and colonies were harvested at the indicated times to determine the fraction of CFUs derived from heat-resistant spores for both the devI-containing and control cells. Boxes below each graph indicate the frequency of devI-containing CFUs at each timepoint. Data shown are the average from two populations (biological replicates) per timepoint with error bars indicating the standard deviation.
-
Figure 6—source data 1
DevI inhibits sporulation and provides benefit.
Raw counts and calculated frequencies of spore-derived CFUs and total CFUs measured at different times during competition of cells ectopically expressing devI and wild-type cells.
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig6-data1-v2.xlsx
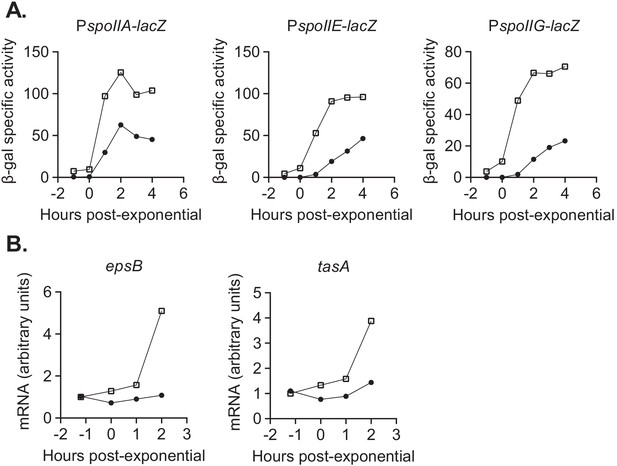
devI inhibits expression of genes associated with sporulation initiation and biofilm formation.
(A) ICEBs1-cured cells that constitutively express devI (black circles) or contain an empty expression construct (open squares) were grown in liquid sporulation medium. Cells were harvested at the indicated times and β-galactosidase specific activity was measured. Strains: PspoIIA-lacZ Pxis-devI (JMJ732), PspoIIA-lacZ WT (JMJ735), PspoIIE-lacZ Pxis-devI (JMJ731), PspoIIE-lacZ WT (JMJ734), PspoIIG-lacZ Pxis-devI (JMJ733), PspoIIG-lacZ WT (JMJ736). A representative experiment is shown. (B) ICEBs1-cured cells that constitutively express devI (JMJ725, black circles) and ICEBs1-cured cells containing an empty expression construct (JMJ727, open squares) were grown in liquid biofilm medium, and cells were harvested at the indicated times. cDNA was synthesized using reverse transcriptase and RT-qPCR was used to measure expression of biofilm-associated genes epsB and tasA. The transcript copy numbers of these genes were measured relative to a housekeeping gene gyrA. The data reported are the average of three technical replicates from one experiment. The relative expression levels are normalized to wild-type at T-1. A representative experiment is shown.
-
Figure 7—source data 1
Sporulation and biofilm gene expression.
Raw data used to calculate beta-gal activity (for early sporulation gene reporters) and mRNA transcript abundance (RT-qPCR for biofilm gene expression).
- https://cdn.elifesciences.org/articles/65924/elife-65924-fig7-data1-v2.xlsx
Tables
Frequency of transconjugants generated in biofilm matings.
| Initial frequency ICEBs1 donors* | Final frequency transconjugants† |
|---|---|
| 0.008 ± 0.002 | 0.42 ± 0.13 |
| 0.10 ± 0.03 | 0.64 ± 0.15 |
| 0.47 ± 0.05 | 0.44 ± 0.07 |
| 0.89 ± 0.03 | 0.063 ± 0.008 |
-
*ICEBs1-containing cells (JMJ592) were mixed with ICEBs1-cured cells (JMJ550). The initial frequencies reported are the average ± standard deviation from three independent experiments.
†The final frequencies of transconjugants reported are the average ± standard deviation from a total of nine biofilms from three independent experiments.
-
Table 1—source data 1
Mating in biofilms.
Counts of donors, recipients, and transconjugants for ICEBs1 mating in biofilms.
- https://cdn.elifesciences.org/articles/65924/elife-65924-table1-data1-v2.xlsx
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Bacillus subtilis NCIB3610) | DS2569 | Konkol et al., 2013. PMID:23836866 | NCIB3610 cured of pBS32 plasmid. Gift of Daniel Kearns to Avigdor Eldar. | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ550 | This paper | ICEBs10 lacA::{Ppen-mApple2 kan} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ574 | This paper | ICEBs10 lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ576 | This paper | ICEBs10 bcaP::{PrapI-rapIphrI kan} lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ592 | This paper | ICEBs1 yddJ-cat-yddK lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ593 | This paper | ICEBs1 conEK476E yddJ-cat-yddK lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ646 | This paper | ICEBs1 oriT* attR::tet lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ662 | This paper | ICEBs1 ∆Pxis oriT* attR::tet lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ686 | This paper | ICEBs1 oriT* ∆rapIphrI attR::tet lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ688 | This paper | ICEBs1 oriT* ∆sncO attR::tet lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ703 | This paper | ICEBs1 oriT* ΔdevI attR::tet lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ704 | This paper | ICEBs1 oriT* ∆ydzL attR::tet lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ714 | This paper | ICEBs10 lacA::spec bcaP::kan | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ725 | This paper | ICEBs10 lacA::{Pxis-devI mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ727 | This paper | ICEBs10 lacA::{Pxis-empty mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ731 | This paper | ICEBs10 lacA::{Pxis-devI mls} amyE::{PspoIIE-lacZ cat} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ732 | This paper | ICEBs10 lacA::{Pxis-devI mls} amyE::{PspoIIA-lacZ cat} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ733 | This paper | ICEBs10 lacA::{Pxis-devI mls} amyE::{PspoIIG-lacZ cat} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ734 | This paper | ICEBs10 lacA::{Pxis-empty mls}amyE::{PspoIIE-lacZ cat} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ735 | This paper | ICEBs10 lacA::{Pxis-empty mls} amyE::{PspoIIA-lacZ cat} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ736 | This paper | ICEBs10 lacA::{Pxis-empty mls} amyE::{PspoIIG-lacZ cat} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ785 | This paper | ICEBs1 oriT* ∆rapIphrI attR::tet bcaP::{PrapI-rapIphrI kan} lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ786 | This paper | ICEBs10 spo0A∆Ps lacA::{Ppen-mApple2 kan} | |
| Strain, strain background (Bacillus subtilis DS2569) | JMJ788 | This paper | ICEBs1 conEK476E yddJ-cat-yddK spo0A∆Ps lacA::{Pveg-mTagBFP mls} | |
| Strain, strain background (Escherichia coli MC1061) | AG1111 | Ireton et al., 1993 PMID:8436298 | E. coli strain for cloning and maintaining plasmids. MC1061 with F’ proAB+ lacIq lacZM15 Tn10. | |
| Recombinant DNA reagent | pJMJ196 (plasmid) | This paper | For generating oriT* nick; derived from pCAL1422. | |
| Recombinant DNA reagent | pJMJ430 (plasmid) | This paper | For generating unmarked rapI-phrI deletion; derived from pCAL1422. | |
| Recombinant DNA reagent | pJMJ199 (plasmid) | This paper | For generating unmarked Pxis deletion; derived from pCAL1422. | |
| Recombinant DNA reagent | pELS5 (plasmid) | Other | For generating unmarked ydzL deletion; derived from pCAL1422. From Grossman lab collection. | |
| Recombinant DNA reagent | pELS1 (plasmid) | Other | For generating unmarked devI deletion; derived from pCAL1422. From Grossman lab collection. | |
| Recombinant DNA reagent | pELC815 (plasmid) | Other | For generating unmarked sncO deletion; derived from pCAL1422. From Grossman lab collection. | |
| Recombinant DNA reagent | pJT245 (plasmid) | Other | Source of oriT* nicK allele. From Grossman lab collection. | |
| Recombinant DNA reagent | pCAL1422 (plasmid) | Thomas et al., 2013. PMID:23326247 | For generating markerless deletions/mutations. | |
| Recombinant DNA reagent | pMMH253 (plasmid) | Other | Vector for integration of constructs at bcaP. From Grossman lab collection. | |
| Recombinant DNA reagent | pJMJ354 (plasmid) | This paper | Native rapI-phrI expression construct for integration at bcaP; derived from pMMH253 | |
| Sequence-based reagent | oJJ363 | Sigma-Aldrich | qPCR primer | CGGAACAATATCGCACCATTC |
| Sequence-based reagent | oJJ364 | Sigma-Aldrich | qPCR primer | CGCTGCACTGAACGATTTAC |
| Sequence-based reagent | oJJ367 | Sigma-Aldrich | qPCR primer | GGATCACTTGCGATCAAAGAAG |
| Sequence-based reagent | oJJ368 | Sigma-Aldrich | qPCR primer | CTTCAAACTGGCTGAGGAAATC |
| Sequence-based reagent | oMEA128 | Sigma-Aldrich | qPCR primer | TGGAGCATTACCTTGACCATC |
| Sequence-based reagent | oMEA129 | Sigma-Aldrich | qPCR primer | AGCTCTCGCTTCTGCTTTAC |
| Commercial assay or kit | RNeasy PLUS | Qiagen | Cat No. 74136 | |
| Commercial assay or kit | iScript Reverse Transcription Supermix | Bio-Rad | Cat No. 1708840 | |
| Commercial assay or kit | SsoAdvanced SYBR master mix | Bio-Rad | Cat No. 1725270 |
B. subtilis strains used*.
| Strain | Relevant genotype |
|---|---|
| JMJ550 | ICEBs10 lacA::{Ppen-mApple2 kan} |
| JMJ574 | ICEBs10 lacA::{Pveg-mTagBFP mls} |
| JMJ576 | ICEBs10 bcaP::{PrapI-rapIphrI kan} lacA::{Pveg-mTagBFP mls} |
| JMJ592 | ICEBs1 yddJ-cat-yddK lacA::{Pveg-mTagBFP mls} |
| JMJ593 | ICEBs1 conEK476E yddJ-cat-yddK lacA::{Pveg-mTagBFP mls} |
| JMJ646 | ICEBs1 oriT* attR::tet lacA::{Pveg-mTagBFP mls} |
| JMJ662 | ICEBs1 ∆Pxis oriT* attR::tet lacA::{Pveg-mTagBFP mls} |
| JMJ686 | ICEBs1 oriT* ∆rapIphrI attR::tet lacA::{Pveg-mTagBFP mls} |
| JMJ688 | ICEBs1 oriT* ∆sncO attR::tet lacA::{Pveg-mTagBFP mls} |
| JMJ703 | ICEBs1 oriT* ΔdevI attR::tet lacA::{Pveg-mTagBFP mls} |
| JMJ704 | ICEBs1 oriT* ∆ydzL attR::tet lacA::{Pveg-mTagBFP mls} |
| JMJ714 | ICEBs10 lacA::spec bcaP::kan |
| JMJ725 | ICEBs10 lacA::{Pxis-devI mls} |
| JMJ727 | ICEBs10 lacA::{Pxis-empty mls} |
| JMJ731 | ICEBs10 lacA::{Pxis-devI mls} amyE::{PspoIIE-lacZ cat} |
| JMJ732 | ICEBs10 lacA::{Pxis-devI mls} amyE::{PspoIIA-lacZ cat} |
| JMJ733 | ICEBs10 lacA::{Pxis-devI mls} amyE::{PspoIIG-lacZ cat} |
| JMJ734 | ICEBs10 lacA::{Pxis-empty mls} amyE::{PspoIIE-lacZ cat} |
| JMJ735 | ICEBs10 lacA::{Pxis-empty mls} amyE::{PspoIIA-lacZ cat} |
| JMJ736 | ICEBs10 lacA::{Pxis-empty mls} amyE::{PspoIIG-lacZ cat} |
| JMJ785 | ICEBs1 oriT* ∆rapIphrI attR::tet bcaP::{PrapI-rapIphrI kan} lacA::{Pveg-mTagBFP mls} |
| JMJ786 | ICEBs10 spo0A∆Ps lacA::{Ppen-mApple2 kan} |
| JMJ788 | ICEBs1 conEK476E yddJ-cat-yddK spo0A∆Ps lacA::{Pveg-mTagBFP mls} |
-
1All strains derived from NCIB3610 plasmid-free.
Additional files
-
Source data 1
Neutral fitness due to marker effects.
Counts of two ICEBs1-cured strains each bearing an antibiotic resistance marker (used to distinguish strains) at the start and end of competitions.
- https://cdn.elifesciences.org/articles/65924/elife-65924-data1-v2.xlsx





