A hypothesis on the capacity of plant odorant-binding proteins to bind volatile isoprenoids based on in silico evidences
Figures
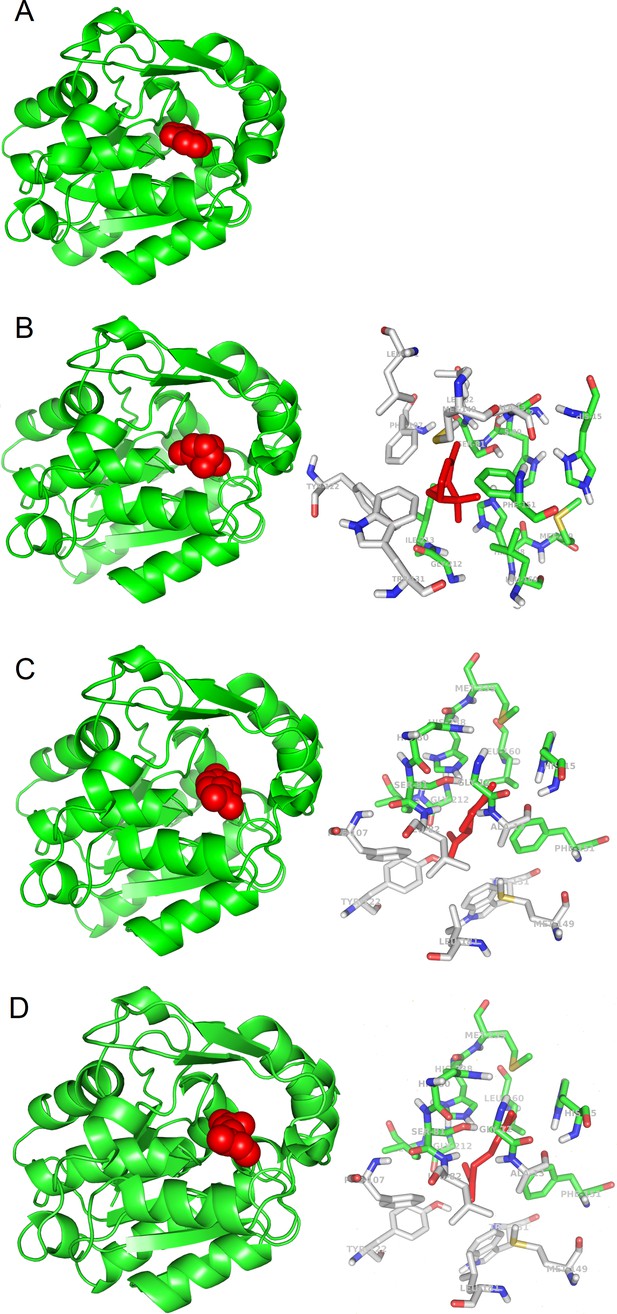
3D models of protein-ligand interactions.
(A) Experimental structure of SABP2 protein structure with salicylic acid in the binding site (PDB structure 1Y7I). The architecture of SABP2 is schematized by the backbone structure (green), with ribbons and arrows to evidence helices and beta strands, respectively. The salicylic acid molecule (colored in red) is in space-fill representation. (B) Left: Molecular docking simulation of SABP2 protein structure with α-pinene (red molecule) in the binding site. SABP2 is shown with the same spatial orientation as in panel (A) to emphasize that α-pinene occupies the same binding site of salicylic acid. Right: Focus on the binding site of SABP2. α-Pinene (red molecule) interacts directly with amino acid residues labeled with carbon atoms in green. Amino acid residues with carbon atom in grey are also part of the binding site, although not directly in contact with α-pinene. Standard colors are used for the other amino acid atoms (red = oxygen, blue = nitrogen, yellow = sulphur, white = hydrogen). (C) Left: SABP2 protein with limonene (red molecule) in the binding site. SABP2 is shown with the same spatial orientation of (A), to better show that limonene occupies the same binding site of the salicylic acid. Right: Focus of the binding site of SABP2. Details as in panel (B). (D) Left: SABP2 protein with β-myrcene (red molecule) in the binding site. SABP2 is shown with the same spatial orientation of (A), to better show that β-myrcene occupies the same binding site of the salicylic acid. Right: Focus of the binding site of SABP2. Details as in (B).
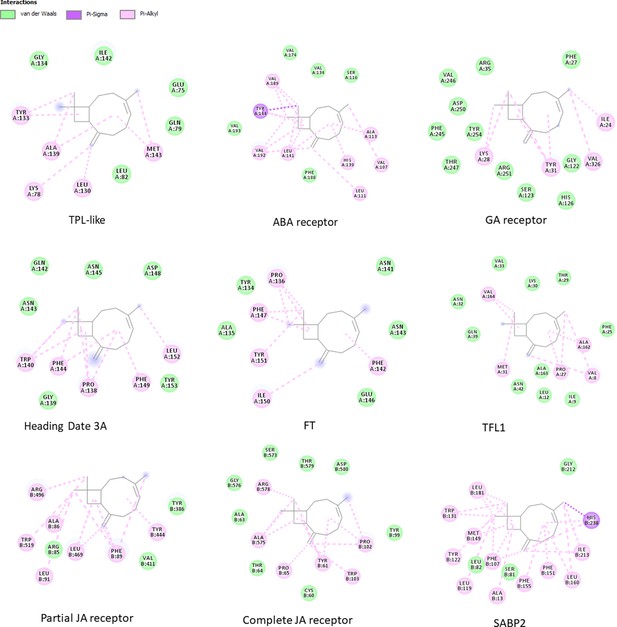
The environment of the binding pocket of β-caryophyllene in the complexes with best binding energy values.
For each candidate OBP, the 2D image shows β-caryophyllene in the center, surrounded by the residues with which it interacts. The type of interaction is defined according to the color legend in figure. Images are obtained with DiscoveryStudio software.
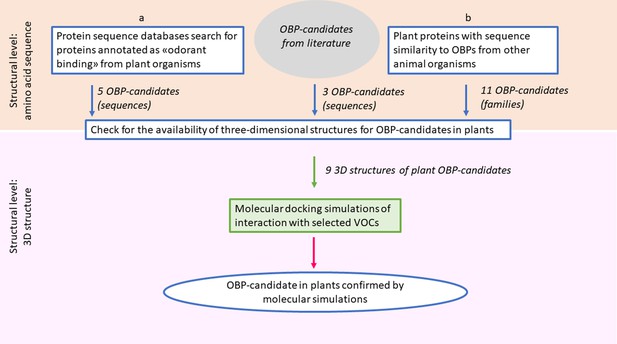
Schematic workflow for the search of candidate odorant-binding proteins in plants.
The search was performed at two levels of investigation. The first level was the identification of candidate plant OBPs, based on features and similarities of protein sequences from public data bases (boxes labeled a and b) and on literature evidences. This level identified proteins or protein families, for which amino acid sequences are available. The second level was the validation by in silico molecular simulations, to verify the potential capability of the candidate proteins to bind selected VOCs. This level can be applied only to the candidate proteins, selected from the first level, for which the three-dimensional structure is made available by experimental studies.
Tables
Similarities of animal and plant protein families with odorant receptor/transporter/channel functions.
| Animal protein family | Similar plant proteins | Identity percentage range | Sequence coverage range | BLAST E-value |
|---|---|---|---|---|
| Bovine cyclic nucleotide gated olfactory channel of Bos Taurus | Potassium voltage-gated channel | 25–30% | 60–75% | ≤3e-27 |
| Chloride channel Anoctamin-2 of Mus musculus | anoctamin-like protein | 22–27% | 50–70% | ≤3e-20 |
| Human/mouse receptor expression enhancing molecules | HVA22 e HVA22-like plants proteins | 27–44% | 40–96% | ≤7e-21 |
| BPI fold containing family B member three from Mus musculus and Rattus norvegicus | putative BPI, lipid-binding protein, hypothetical protein, and unnamed protein | 20–30% | 25–60% | ≤6e-07 |
| putative OBP 5a from Drosophila melanogaster | FT, D3-like, protein ‘Mother of FT and TFL1-like (Terminal flower 1-like)’ (MFT), ZCN9 (MFT-like), Heading Date 3A-like | 28–41% | 51–86% | ≤5e-18 |
Binding energy values (Kcal mol−1), obtained by docking simulations, between putative plant OBPs and isoprenoid VOCs.
| OBPs | α-Pinene | Limonene | β-Myrcene | β-Caryophyllene | Isoprene | Linalool |
|---|---|---|---|---|---|---|
| TPL-like | −5.06 | −4.76 | −3.95 | −6.16 | −3.39 | −3.94 |
| ABA receptor | −4.69 | −4.73 | −3.84 | −6.32 | −3.35 | −4.12 |
| GA receptor | −5.50 | −5.44 | −4.92 | −6.94 | −3.78 | −4.69 |
| Heading Date 3A | −5.46 | −4.99 | −4.29 | −6.83 | −3.44 | −4.79 |
| FT | −5.23 | −5.07 | −4.26 | −6.65 | −3.15 | −4.98 |
| TFL1 | −5.30 | −5.53 | −4.38 | −7.15 | −3.70 | −4.98 |
| Partial JA receptor | −5.23 | −4.79 | −4.16 | −6.05 | −2.78 | −4.21 |
| Complete JA receptor | −5.92 | −5.11 | −4.41 | −6.61 | −2.82 | −4.49 |
| SABP2 | −6.03 | −6.12 | −5.14 | −6.73 | −3.25 | −4.70 |
| Reference protein* | −5.42 | −6.29 | −4.15 | Not available | Not available | Not available |
-
*Reference protein is the protein for which it has been found an experimental complex with the ligand. For α-pinene, limonene, and β-myrcene, the reference proteins are cytochrome P450 2B6 complexed with α-pinene (PDB code: 4I91), lipid binding protein complexed with limonene 1,2 epoxide (PDB code: 2A2G, and linalool dehydratase/isomerase complexed with β-myrcene [PDB code: 5HSS]), respectively.
Additional files
-
Source data 1
Database accession numbers.
Related to Table 1, Figure 1 and Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/66741/elife-66741-data1-v1.xlsx
-
Source data 2
Docking results.
- https://cdn.elifesciences.org/articles/66741/elife-66741-data2-v1.xlsx
-
Source data 3
PDB structures of SABP2 complexes.
Related to Figure 1.
- https://cdn.elifesciences.org/articles/66741/elife-66741-data3-v1.zip
-
Source data 4
PDB structures of protein - beta caryophyllene complexes.
Related to Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/66741/elife-66741-data4-v1.zip
-
Supplementary file 1
Plant proteins from sequence databases annotated as ‘odorant-binding protein’.
- https://cdn.elifesciences.org/articles/66741/elife-66741-supp1-v1.docx
-
Supplementary file 2
BLAST results for non-plant proteins with similarity to ‘general odorant-binding protein 56d’ from Anthurium amnicola.
Only the best ten results are shown.
- https://cdn.elifesciences.org/articles/66741/elife-66741-supp2-v1.docx
-
Supplementary file 3
Complete results of docking simulation experiments.
Docking simulations by AutoDock suite generated results summarized in the table. For each receptor-ligand simulation, the best 100 conformations are clusterized by AutoDock on the basis of ligand position, and three values are reported for each cluster: the mean binding energy for the clusterized conformations, the best binding energy, the number of conformations in the cluster. An additional parameter computed by AutoDock is the predicted Ki value. We reported the results for the best cluster in terms of energy and population, or in some cases, alternative clusters, with reference to different pockets on the receptor surface.
- https://cdn.elifesciences.org/articles/66741/elife-66741-supp3-v1.docx
-
Supplementary file 4
List of 432 protein sequences selected as OBP from animal sources, used for searching public database for similar plant proteins.
- https://cdn.elifesciences.org/articles/66741/elife-66741-supp4-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/66741/elife-66741-transrepform-v1.docx





