ADF and cofilin-1 collaborate to promote cortical actin flow and the leader bleb-based migration of confined cells
Figures
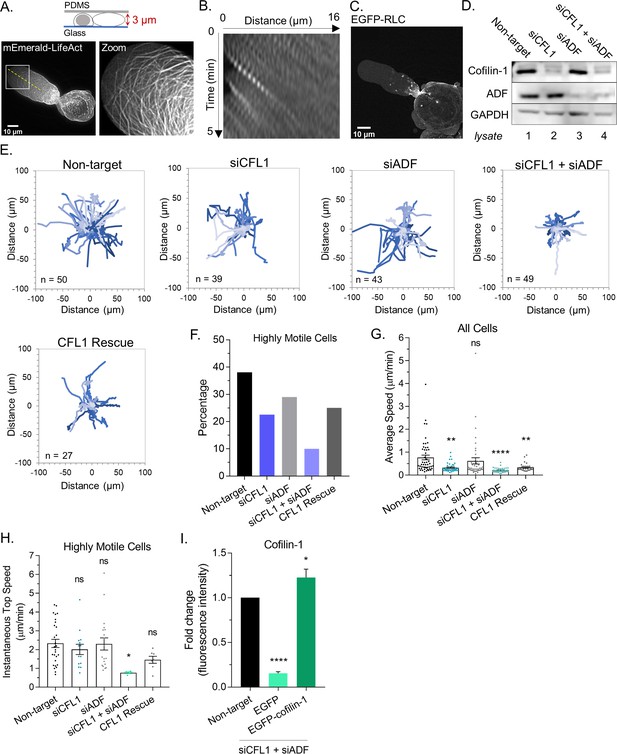
ADF and cofilin-1 are required for leader bleb-based migration.
(A) Ventral Z-section of a melanoma A375-M2 cell, which has been confined down to 3 µm, with mEmerald-LifeAct. (B) Kymograph from (A; dashed line), showing cortical actin flow. (C) Ventral Z-section of a melanoma A375-M2 cells, which has been confined down to 3 µm, with EGFP tagged regulatory light chain (EGFP-RLC). (D) Western blot confirming CFL1, actin depolymerizing factor (ADF), and ADF + CFL1 RNAi in melanoma A375-M2 cells. (E) Individual cell migration tracks (plot of origin) for non-targeting, CFL1, ADF, and CFL1 + ADF RNAi cells, as well as CFL1 + ADF RNAi cells rescued by transfection with EGFP-cofilin-1 plasmid. In each, cells were tracked over a period of 5 hr. Relative y (µm) and relative × (µm) are shown in each. (F) Percentage of highly motile cells from (E). Cells that traveled a distance equivalent to at least one cell length over the course of the 5 hr time-lapse were classified as highly motile. (G) Average speed (µm/min) from cells in (E; mean ± SEM). Statistical significance was determined by one-way ANOVA and a Dunnett’s post hoc test. (H) Instantaneous top speed (µm/min) for highly motile cells in (E; mean ± SEM). (I) Cofilin-1 levels (fold change; fluorescence intensity) of adhered RNAi cells by immunofluorescence confirming rescue by transfection with EGFP-cofilin-1 or not rescued with EGFP. Statistical significance was determined by an unpaired one-sample t-test. All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
-
Figure 1—source data 1
Raw data from manual tracking.
Spreadsheet of x-y coordinates of cells for each frame within a video. Time interval between frames for all cells is 8 min. Data for CFL1, actin depolymerizing factor (ADF), CFL1 + ADF RNAi, as well as CFL1 + ADF RNAi cells rescued by transfection with EGFP-cofilin-1 plasmid are provided. These data are displayed as plots of origin in Figure 1E.
- https://cdn.elifesciences.org/articles/67856/elife-67856-fig1-data1-v2.xlsx
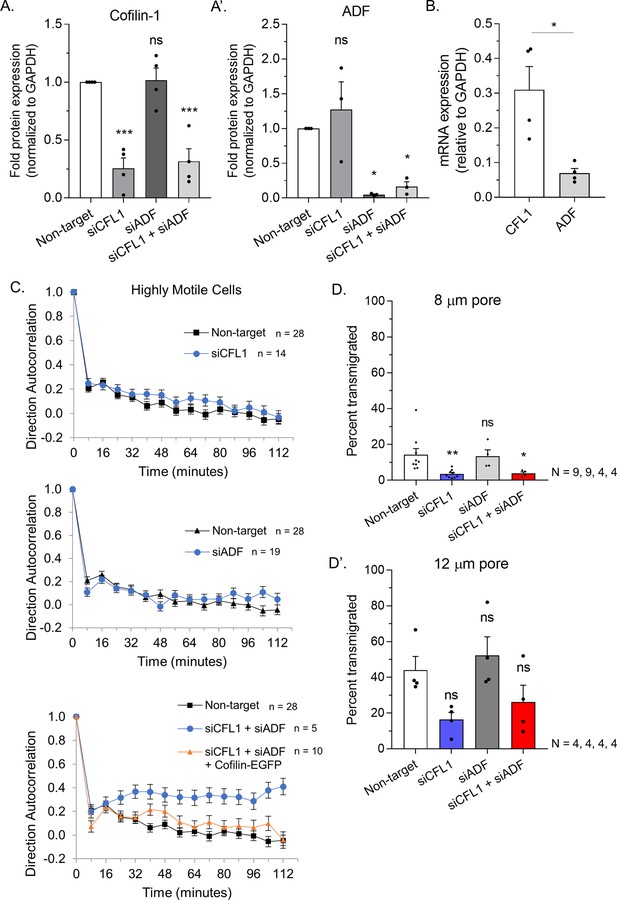
Protein levels and migration parameters.
(A) Quantitative evaluation of cofilin-1 (left) and actin depolymerizing factor (ADF) (right) RNAi as measured by Western blotting of total cell lysates. (B) Quantitative evaluation of cofilin-1 and ADF mRNA levels by qRT-PCR in A375-M2 cells (mean ± SEM). (C) Direction autocorrelation plots of confined highly motile cells after RNAi. (D) Transmigration of melanoma A375-M2 cells through 8 (above) or 12 µm (below) pores after non-targeting, CFL1, ADF, and CFL1 + ADF RNAi. Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
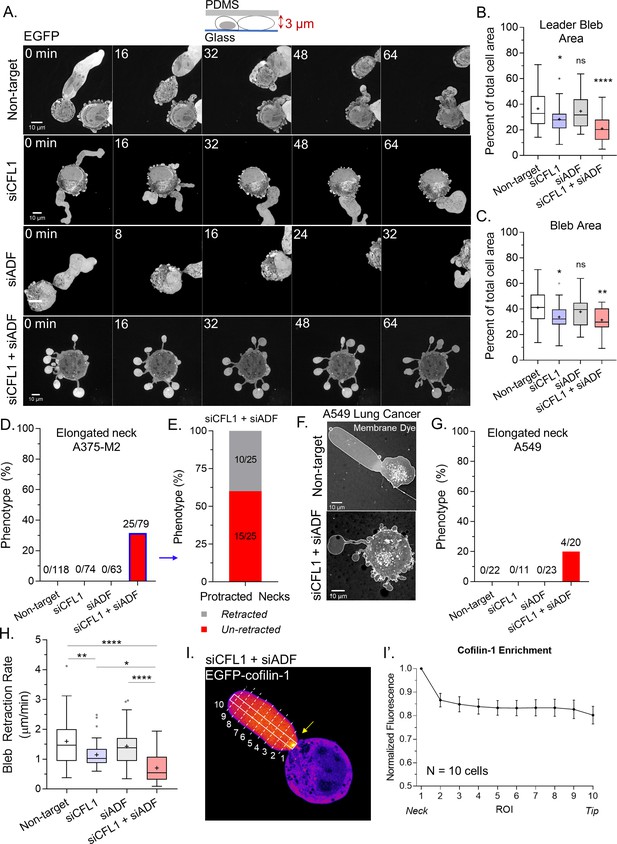
Together, actin depolymerizing factor (ADF) and cofilin-1 are required to retract blebs.
(A) Montage of non-targeting, CFL1, ADF, and CFL1 + ADF RNAi with EGFP alone (volume marker) in melanoma A375-M2 cells. (B–C) Quantitation of area for leader (A) and all blebs (B) after non-targeting, CFL1, ADF, and ADF + CFL1 RNAi. Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (D) Percent of non-targeting, CFL1, ADF, and CFL1 + ADF RNAi cells with elongated bleb necks. (E) Percent of ADF + CFL1 RNAi cells from (D) with elongated bleb necks that retract vs. un-retracted. (F–G) Lung adenocarcinoma A549 cells after non-targeting and CFL1 + ADF RNAi stained with a far-red fluorescent membrane dye (F). Percent of non-targeting, CFL1, ADF, and CFL1 + ADF RNAi cells with elongated bleb necks (G). (H) Bleb retraction rates for non-targeting (45 blebs; 26 cells), CFL1 (40 blebs; 20 cells), ADF (48 blebs; 30 cells), and CFL1 + ADF RNAi (38 blebs; 23 cells). Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (I) EGFP-cofilin-1 localization in an A375-M2 cell confined down to 3 µm. Arrow points to an enrichment of cofilin-1 at the leader bleb neck. (I’) Regional analysis of EGFP-cofilin-1 average fluorescence intensity in ROIs sampled from bleb neck to tip (mean ± SEM). Representative regions taken within white box and dashed lines in (I). All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
-
Figure 2—source data 1
Raw leader bleb and bleb area measurements.
Spreadsheet of leader bleb and bleb (i.e., all blebs) area measurements. Each measurement is a percent of the total cell area. Data for CFL1, actin depolymerizing factor (ADF), and CFL1 + ADF RNAi are provided. These data are graphed in Figure 2B–C.
- https://cdn.elifesciences.org/articles/67856/elife-67856-fig2-data1-v2.xlsx
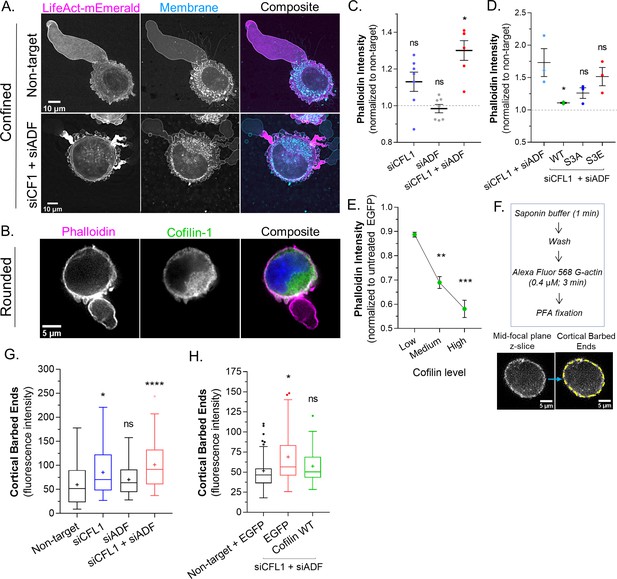
Actin depolymerizing factor ( ADF) and cofilin-1 rapidly disassemble cortical actin.
(A) mEmerald-LifeAct and far-red fluorescent membrane dye in cells after non-targeting and CFL1 + ADF RNAi. (B) Cells freshly plated on poly-L-lysine coated cover glass stained for endogenous cofilin-1 and filamentous-actin (F-actin) (phalloidin). (C) F-actin levels (normalized to non-target; mean ± SEM) after CFL1, ADF, and CFL1 + ADF RNAi in trypsinized (spherical) cells, as determined by flow cytometry. Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (D) F-actin levels (normalized to non-target; mean ± SEM) after CFL1 + ADF RNAi, as well as after CFL1 + ADF RNAi with EGFP-cofilin-1 WT, S3A, or S3E, as determined by flow cytometry. Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (E) F-actin level (normalized to EGFP alone; mean ± SEM) as a function of increasing EGFP-cofilin-1 in cells depleted of endogenous cofilin-1 and ADF by RNAi, as determined by flow cytometry. Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (F) Top, barbed end assay workflow. Bottom, representative image of a freshly plated (spherical) cell subjected to the barbed end assay. (G) As shown in (F; bottom), the level of cortical barbed ends was measured in cells after non-targeting (71 cells), CFL1 (53 cells), ADF (47 cells), and CFL1 + ADF RNAi (83 cells). Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (H) As shown in (F; bottom), the level of cortical barbed ends was measured in cells with non-targeting and EGFP (42 cells), as well as after CFL1 + ADF RNAi with EGFP (32 cells) or EGFP-cofilin-1 (27 cells). Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
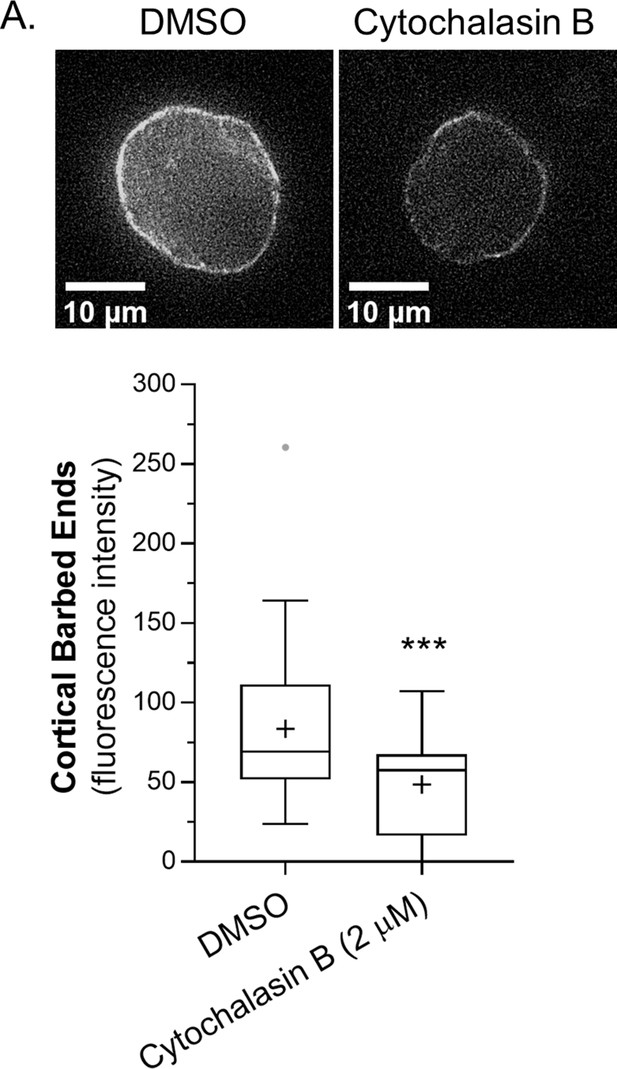
Cytochalasin B blocks access to cortical barbed ends.
(A) Cortical barbed end levels in melanoma A375-M2 cells after vehicle (DMSO; 53 cells) or cytochalasin B (2 µM; 24 cells) treatment. Statistical significance was determined using an unpaired Student’s t-test. All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
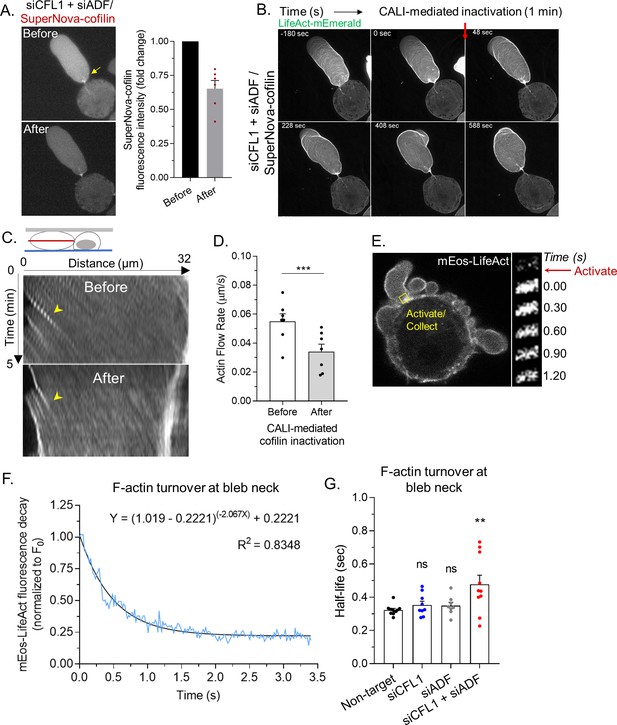
Rapid cortical actin flow requires actin depolymerizing factor (ADF) and cofilin-1 severing at leader bleb necks.
(A) Left, SuperNova-cofilin-1 localization in cells depleted of endogenous cofilin-1 and ADF by RNAi before and after 1 min of red light irradiation. Right, quantitative analysis of CALI, as determined by the fold change in SuperNova emission. (B) Montage of mEmerald-LifeAct before and after cofilin-1 inactivation in a cell depleted of endogenous cofilin-1 and ADF by RNAi. (C) Kymographs of cortical actin (mEmerald-LifeAct) flow from the leader bleb tip before and after cofilin-1 inactivation. (D) Quantitative evaluation of cortical actin flow rates before and after cofilin-1 inactivation. Statistical significance was determined by a paired Student’s t-test. (E) Left, representative image of a freshly plated (spherical) cells with mEos3.2-LifeAct. Right, montage of mEos3.2-LifeAct within the shown ROI before and after photoactivation. (F) Average decay curve for mEos3.2-LifeAct at bleb necks (normalized to the initial fluorescence level; F/F0). The curve was fit using a non-linear single phase decay function. (G) t1/2 for mEos3.2-LifeAct after photoactivation at bleb necks for non-targeting, CFL1, ADF, and CFL1 + ADF RNAi. Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
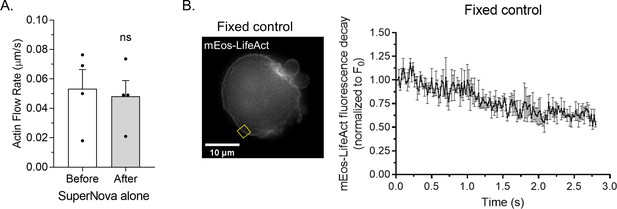
Actin turnover measurement controls.
(A) Cortical actin (mEmerald-LifeAct) flow rates before and after the inactivation (i.e., red light irradiation) of SuperNova alone. (B) Left, sample ROI from a paraformaldehyde (PFA) treated melanoma A375-M2 cell with mEos3.2-LifeAct. Right, average mEos3.2-LifeAct decay (three cells; mean ± SEM) after photoactivation. All data are representative of at least three independent experiments.
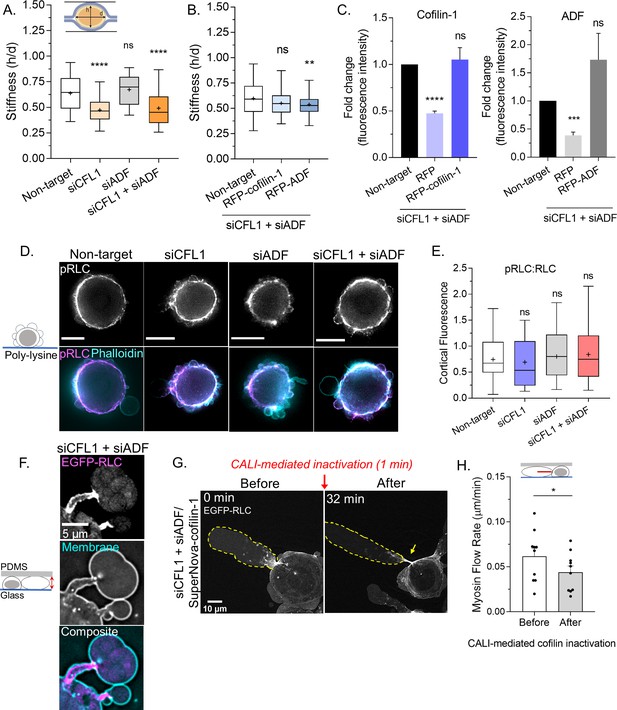
Cofilin-1 supports both actin turnover and myosin contractility at leader bleb necks.
(A) A previously described gel sandwich assay was used to measure the stiffness (h/d) of spherical cells after non-targeting (91 cells), CFL1 (30 cells), actin depolymerizing factor (ADF) (25 cells), and CFL1 + ADF RNAi (42 cells). (B) Cell stiffness (h/d) of spherical cells with non-targeting (181 cells) or siCFL1 + siADF RNAi rescued with RFP-cofilin-1 (41 cells) or RFP-ADF (74 cells). (A–B) Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (C) Cofilin-1 (left) and ADF (right) levels (fluorescence intensity; fold change) of adhered RNAi cells by immunofluorescence confirming rescue by transfection with RFP-cofilin-1 (left) or RFP-ADF (right). Statistical significance was determined by an unpaired one-sample t-test. (D) Immunofluorescence imaging of endogenous phosphorylated regulatory light chain (pRLC) (S18), total RLC, and filamentous-actin (F-actin) (phalloidin) in freshly plated (spherical) cells after non-targeting, CFL1, and CFL1 + ADF RNAi. (E) Ratio of cortical pRLC (S18) to total RLC fluorescence intensity after non-targeting (114 cells), CFL1 (107 cells), ADF (124 cells), and CFL1 + ADF RNAi (91 cells). Statistical significance was determined by one-way ANOVA and a Dunnet’s post hoc test. (F) Localization of EGFP tagged regulatory light chain (EGFP-RLC) in a cell confined down to 3 µm after CFL1 + ADF RNAi. (G) EGFP-RLC dynamics in a cell depleted of cofilin-1 and ADF before and after chromophore assisted light inactivation (CALI) of SuperNova-cofilin-1. Arrow points to myosin accumulating at an elongating leader bleb neck after cofilin-1 inactivation. (H) Myosin minifilament flow rate before and after cofilin-1 inactivation. Statistical significance was determined using a paired Student’s t-test. All data are representative of at least three independent experiments. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
-
Figure 5—source data 1
Raw stiffness measurements.
Spreadsheet of stiffness (h/d) measurements for CFL1, actin depolymerizing factor (ADF), CFL1 + ADF RNAi, as well as CFL1 + ADF RNAi cells rescued by transfection with RFP-cofilin-1 or RFP-ADF plasmid are provided. These data are graphed in Figure 5A–B.
- https://cdn.elifesciences.org/articles/67856/elife-67856-fig5-data1-v2.xlsx
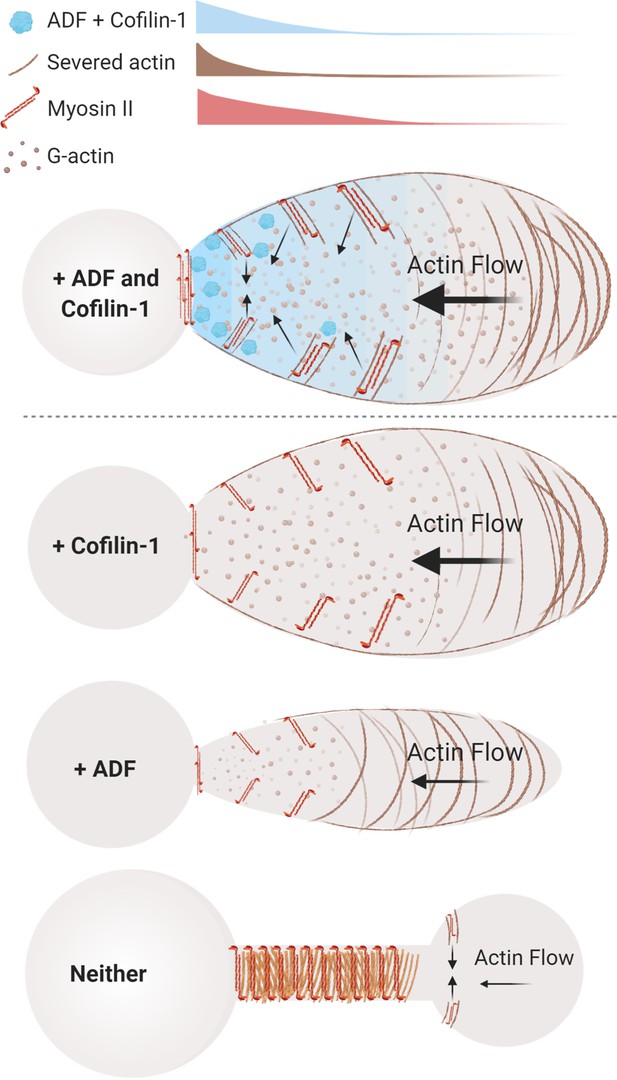
Model of actin depolymerizing factor (ADF) and cofilin-1 function within leader blebs.
Top, in the presence of both ADF and cofilin-1, cells display large blebs with rapid cortical actin flow. Below, in the absence of cofilin-1 or ADF, cells form smaller blebs with slower cortical actin flow. Bottom, without both ADF and cofilin-1, blebs display several defects, including a failure to retract and an accumulation of actomyosin at elongated necks.
Videos
Time-lapse imaging of melanoma A375-M2 cells confined down to 3 µm with far-red plasma membrane dye.
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with mScarlet-LifeAct and EGFP tagged regulatory light chain (EGFP-RLC).
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with the volume marker, mScarlet, after control (non-targeting) RNAi.
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with the volume marker, EGFP, after RNAi of actin depolymerizing factor (ADF) alone.
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with the volume marker, mScarlet, after RNAi of CFL1 alone.
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with the volume marker, EGFP, after RNAi of CFL1 + actin depolymerizing factor (ADF).
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with mEmerald-LifeAct after chromophore assisted light inactivation (CALI) of SuperNova-cofilin-1.
The cell was depleted of endogenous actin depolymerizing factor (ADF) and cofilin-1 by RNAi.
Time-lapse imaging of a melanoma A375-M2 cell confined down to 3 µm with EGFP tagged regulatory light chain (EGFP-RLC) after chromophore assisted light inactivation (CALI) of SuperNova-cofilin-1.
The cell was depleted of endogenous actin depolymerizing factor (ADF) and cofilin-1 by RNAi.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | A375-M2 | ATCC | CRL-3223 | Metastatic melanoma |
| Cell line (Homo sapiens) | A549 | ATCC | CCL-185 | Lung adenocarcinoma |
| Chemical compound, drug | SYLGARD 184 | Dow Corning | Cat no. 24236–10 | PDMS |
| Transfected construct (Homo sapiens) | EGFP-cofilin-1 WT, S3A, and S3E | Addgene (a gift from Dr James Bamburg) | Plasmid no. 50859, 50854, and 50855 | Plasmid constructs to transfect |
| Transfected construct (Homo sapiens) | RFP-cofilin-1 | Dr James Bamburg (Colorado State University) | n/a | Plasmid construct to transfect |
| Transfected construct (Homo sapiens) | RFP-ADF | Dr James Bamburg (Colorado State University) | n/a | Plasmid construct to transfect |
| Transfected construct (Homo sapiens) | SuperNova-cofilin-1 | Dr Kazuyo Sakai (Osaka University, Osaka, Japan) | n/a | Plasmid construct to transfect and destroy cofilin-1 by CALI |
| Transfected construct (Saccharomyces cerevisiae) | mEos3.2-LifeAct | Addgene (a gift from Michael Davidson) | Plasmid no. 54696 | Plasmid construct to transfect and monitor F-actin dynamics |
| Sequence-based reagent (Homo sapiens) | Non-targeting siRNA | Thermo Fisher | Cat no. 4390844 | Control siRNA to transfect |
| Sequence-based reagent (Homo sapiens) | Cofilin-1 siRNA | Thermo Fisher | Cat no. 4392420; s2936 | Cofilin-1 siRNA to transfect |
| Sequence-based reagent (Homo sapiens) | ADF siRNA | Thermo Fisher | Cat no. 4392422; s21737 | ADF siRNA to transfect |
| Antibody | Anti-cofilin-1 (mouse monoclonal) | Thermo Fisher | Cat no. MA5-17275 | WB (1:1000), IF (1:250) |
| Antibody | Anti-ADF (mouse monoclonal) | Thermo Fisher | Cat no. MA5-25485 | WB (1:1000), IF (1:250) |
| Sequence-based reagent (Homo sapiens) | Cofilin-1 forward qPCR primer | Thermo Fisher | n/a | GCAACCTATGAGACCAAGGAGAG |
| Sequence-based reagent (Homo sapiens) | ADF forward qPCR primer | Thermo Fisher | n/a | GCACCAGAACTAGCACCTCTGA |
| Sequence-based reagent (Homo sapiens) | GAPDH forward qPCR primer | Thermo Fisher | n/a | GTCTCCTCTGACTTCAACAGCG |
| Recombinant DNA protein | Alexa Fluor 568-conjugated G-actin from rabbit muscle | Thermo Fisher | Cat no. A12374 | Fluorescent G-actin to label actin barbed ends |
| Software | Fiji | n/a | https://imagej.net/Fiji | Microscopy |
| Software | Prism | GraphPad | n/a | Statistical analyses |
| Software | BioRender | Toronto, ON | n/a | Illustration |
| Other | DeltaVision Elite | GE | n/a | Commercial deconvolution microscopy system |
| Other | LSM880 with fast Airy Scan | Zeiss | n/a | Commercial point scanning confocal microscopy system |





