Ait1 regulates TORC1 signaling and localization in budding yeast
Figures
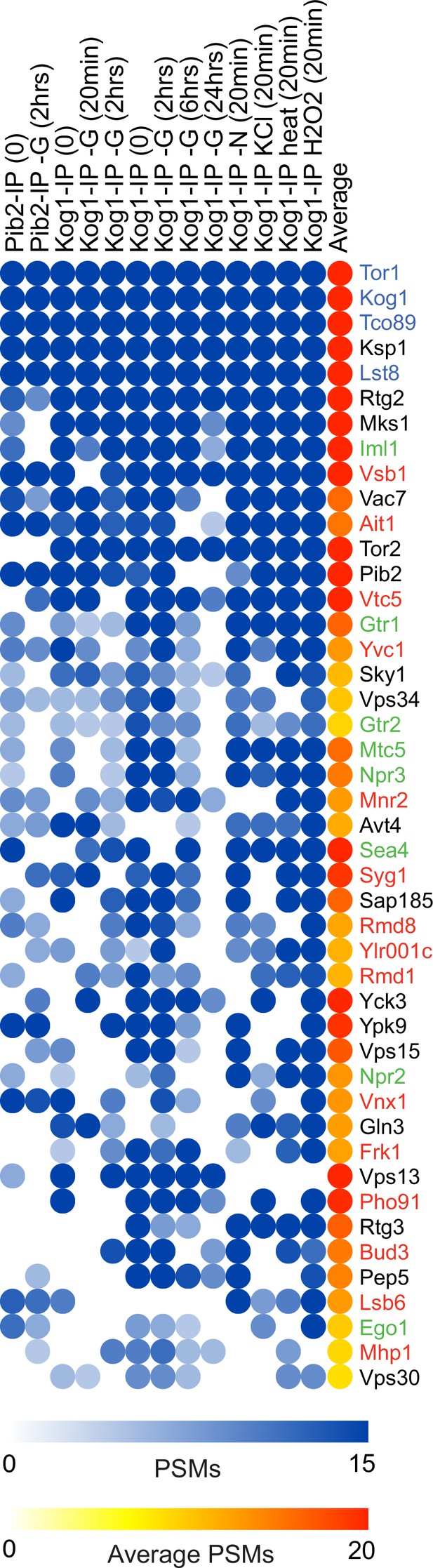
The target of rapamycin complex I (TORC1) interactome in budding yeast.
Blue circles show the number of background corrected peptide spectral maps (PSMs) from each protein identified in a specific Kog1 or Pib2 immunopurification, while the red to yellow scale shows the average number of PSMs across all experiments. The figure shows data for the top 45 TORC1 interactors (those identified in seven or more immunopurifications). The complete dataset is in Supplementary file 1.
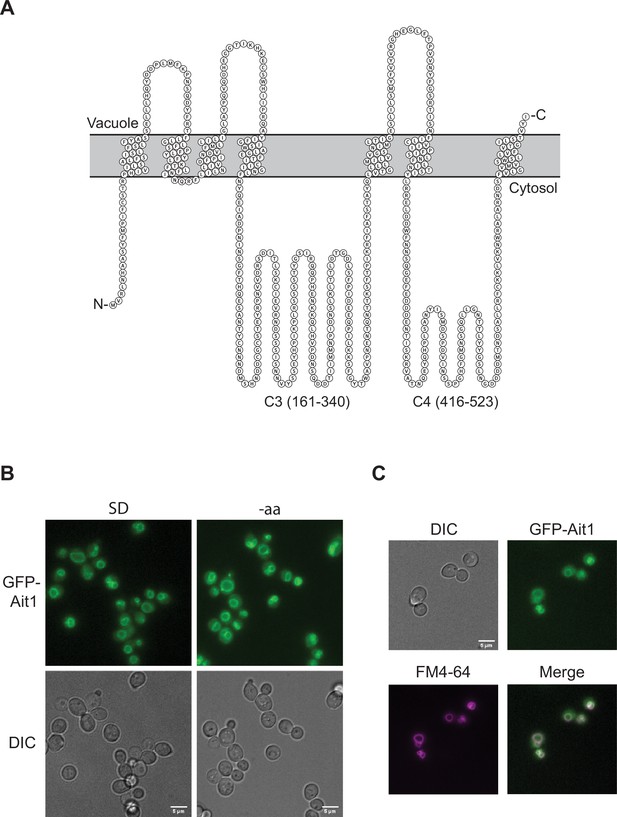
Ait1 is a putative seven-helical transmembrane (GPCR-like) protein that localizes to the vacuolar membrane.
(A) The predicted topology of Ait1/Ydl180w from Protter 1.0 (Omasits et al., 2014). The two large cytosolic loops in Ait1, both of which are predicted to be intrinsically disordered (C3 and C4), are labeled. (B, C) GFP-Ait1 localizes to the vacuolar membrane, as shown by the overlap between the GFP-Ait1 signal and the vacuolar membrane stain FM4-64 signal and does not relocalize in amino acid starvation (shown), or other starvation conditions (not shown).
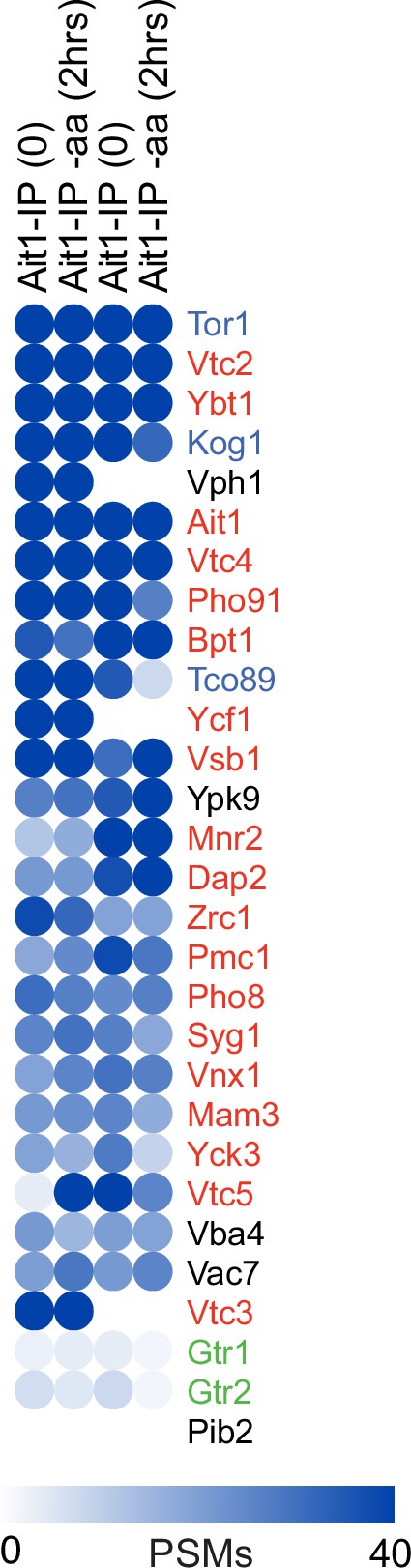
The Ait1 interactome.
Blue circles show the number of background corrected peptide spectral maps (PSMs) for the top 25 proteins identified in the GFP-Ait1 immunopurification (based on the average number of PSMs in the four experiments), along with the data for Gtr1, Gtr2, and Pib2 for comparison.
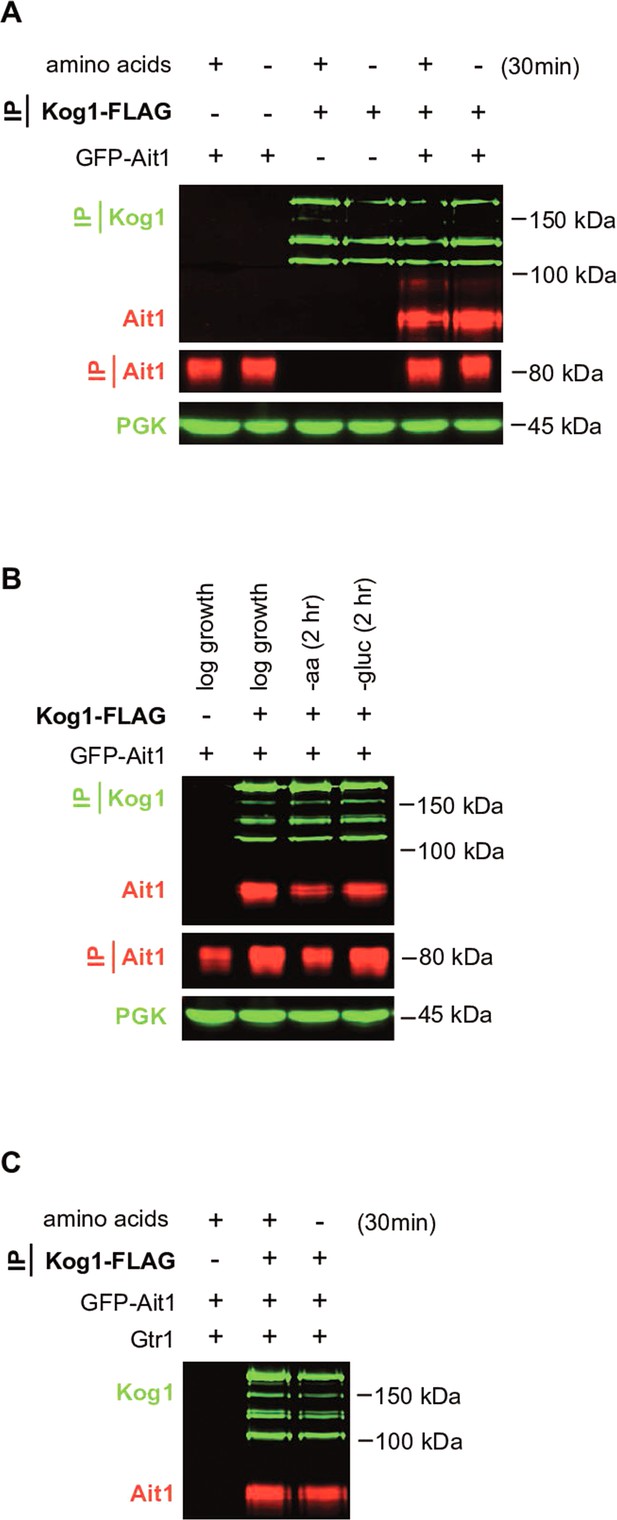
Target of rapamycin complex I (TORC1) interacts with Ait1.
Immunopurification experiments, carried out after treatment with the dithiobis(succinimidyl propionate) (DSP) crosslinker, confirm that Ait1 interacts with TORC1 in vivo. (A) GFP-Ait1 copurifies with Kog1-FLAG both in the presence, and absence, of amino acids. (B) GFP-Ait1 copurifies with Kog1-FLAG even after long-term amino acid and glucose starvation, but at 50% of the level found in log-phase growth. Note (1) that Kog1 partially degrades during the IP and thus shows up as multiple bands and (2) that we were unable to detect Kog1-FLAG or Ait1 after 2 hr of nitrogen starvation (three total attempts), likely due to changes in the solubility of localization of the TORC1–Ait1 complex. (C) Coimmunopurification carried out as in (A) but with 4× protease inhibitors. The interaction between Kog1-FLAG and GFP-Ait1 was not detected in the absence of the crosslinker (data not shown).
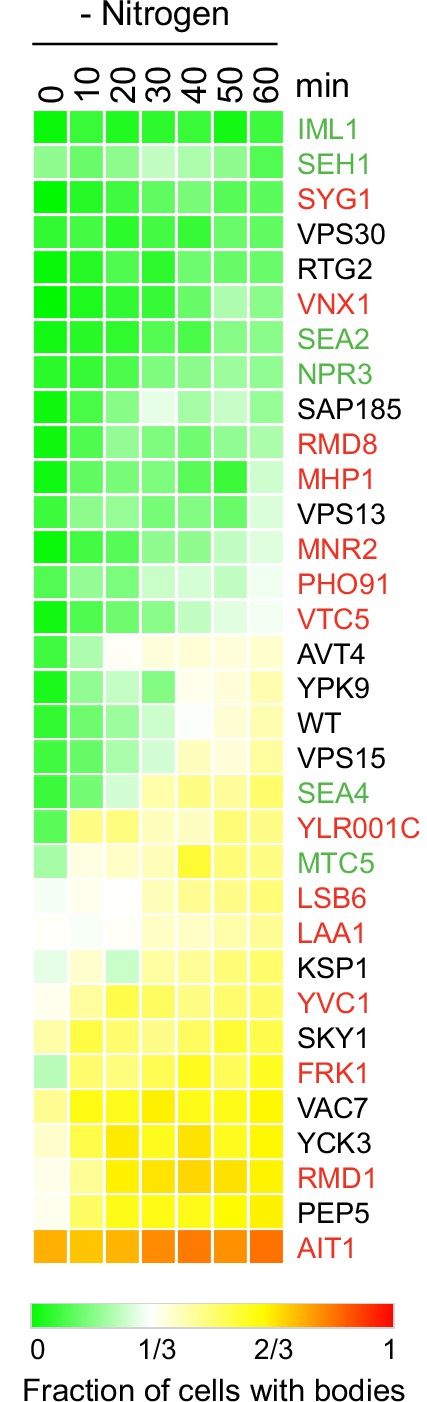
Target of rapamycin complex I (TORC1)-body formation during nitrogen starvation in strains missing key TORC1 interactors.
Each square on the heat map shows the fraction of cells with a Kog1-YFP focus/body at a specific timepoint, calculated by examining the images of >200 cells, per strain, per timepoint. Replicate experiments confirmed the severe defects in the syg1Δ, vps30Δ, rtg2Δ, and vnx1Δ strains (<15% bodies after 1 hr of nitrogen starvation). These follow-up experiments also revealed dramatic variation in the results for vsb1Δcells (even comparing between colonies) leading us to drop the strain from our analysis.
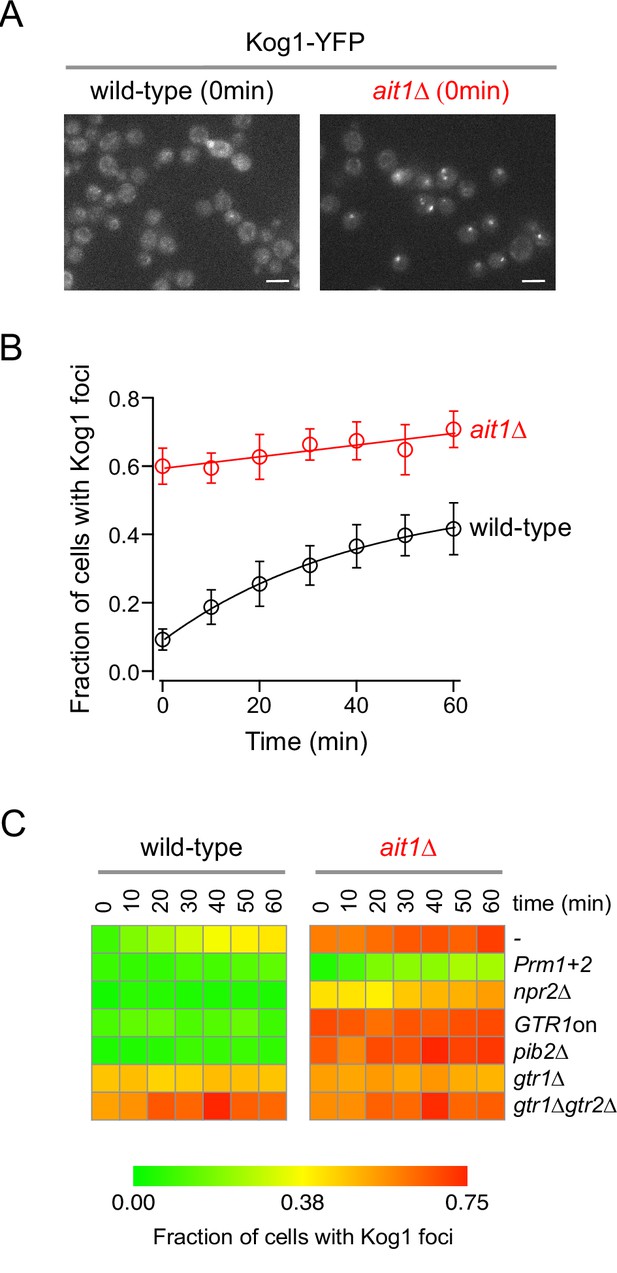
Target of rapamycin complex I (TORC1)-body formation in the ait1Δ strain.
(A) Kog1-YFP localization in the wild-type and ait1Δ strains, during log-phase growth in nutrient-rich (SD) medium. The white bar shows 5 μm. (B) Fraction of wild-type and ait1Δ cells that form Kog1-YFP foci during nitrogen starvation. The points and error bars show the average and standard deviation from five replicate experiments, with at least 100 cells examined, per timepoint, per replicate. (C) Impact of the Ait1 deletion on TORC1-body formation in the wild-type and various mutant strains (compare left and right columns). Experiments were carried out in at least duplicate with over 200 cells examined per timepoint, per mutant. Individual timepoints have errors ranging from 0.05 to 0.10.
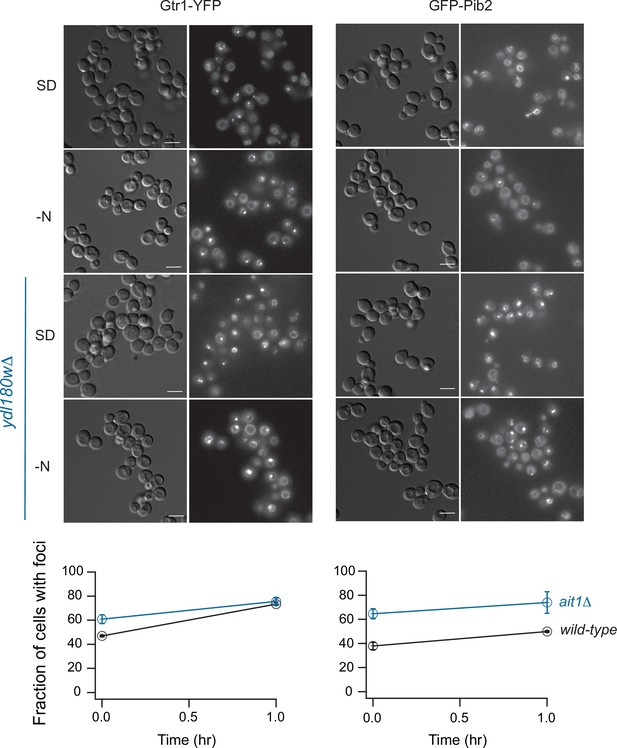
Gtr1 and Pib2 localization in the ait1Δ strain.
(Upper panel) Gtr1-YFP and GFP-Pib2 localization in the wild-type and ait1Δ strains, during log-phase growth in nutrient-rich (SD) medium and 1 hr of complete nitrogen starvation (-N). The white bar in the differential interference contrast image (left column) shows 5 μm; fluorescence images are on the right. (Lower panel) Fraction of wild-type and ait1Δ cells that form Gtr1-YFP and GFP-Pib2 foci during nitrogen starvation. The points and error bars show the average and standard deviation from three replicate experiments, with at least 200 cells examined, per timepoint, per replicate.
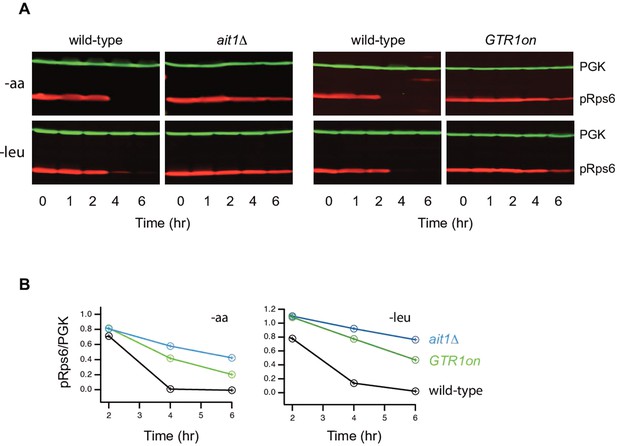
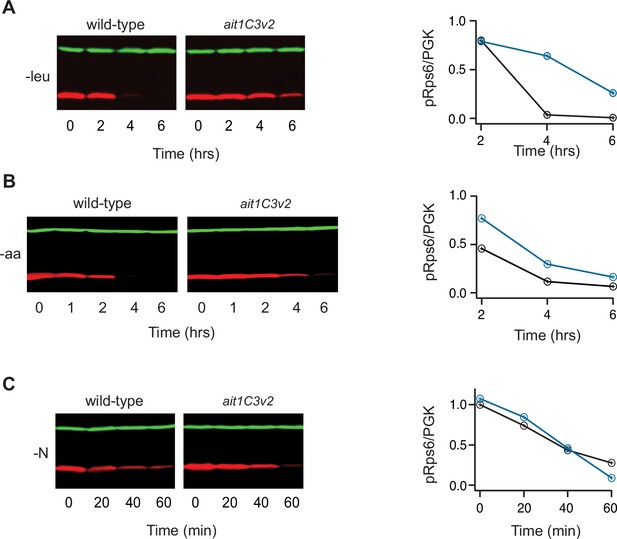
Impact of Ait1 on target of rapamycin complex I (TORC1) activity during amino acid starvation.
(B) TORC1 activity during complete amino acid starvation (top), and leucine starvation (bottom), in wild-type, ait1Δ, and GTR1Q65L (Gtr1on) strains, as measured by western blot using an anti phospho-Rps6 antibody. (b) Values show the ratio of the p-Rps6 signal divided by the PGK (loading control) signal in each lane, relative to the value for the wild-type strain at time = 0. Wild-type and mutant strains were grown and processed together and run on the same gel.
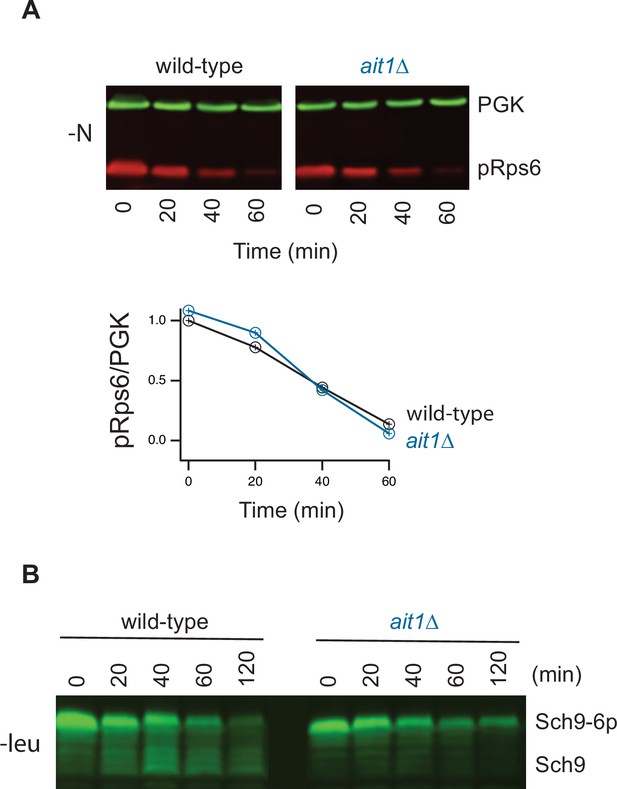
Impact of Ait1 on target of rapamycin complex I (TORC1) activity during nitrogen and amino acid starvation.
(A) TORC1 activity during complete nitrogen starvation in wild-type and ait1Δstrains, as measured by western blot using an anti phospho-Rps6 antibody. Graphs show the ratio of the p-Rps6 signal divided by the PGK (loading control) signal in each lane, relative to the value for the wild-type strain at time = 0. Wild-type and mutant strains were grown and processed together and run on the same gel. (B) Mobility shift assay following the phosphorylation of Sch9-HA during leucine starvation on a western blot.
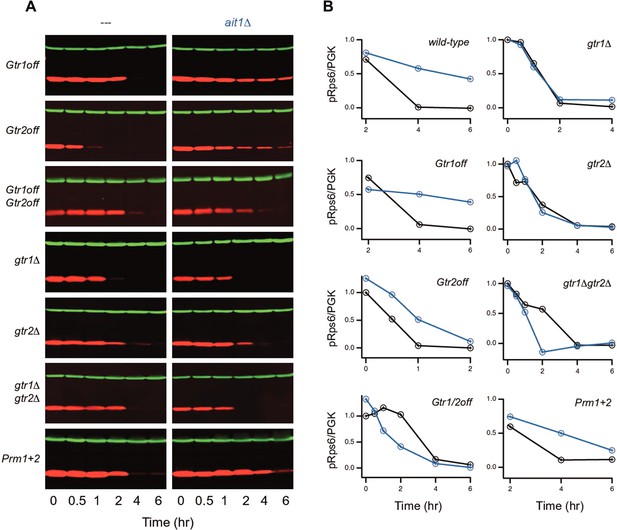
Impact of Ait1 on target of rapamycin complex I (TORC1) activity in strains with mutations in Gtr1/2, or the prion domains in Kog1, during amino acid starvation.
(A) TORC1 activity during complete amino acid starvation in mutant strains with (left column) and without Ait1 (right column), measured using a western blot, as described in Figure 6. (B) Values show the ratio of the p-Rps6 signal divided by the PGK (loading control) signal in each lane, relative to the value for the wild-type strain at time = 0. Mutant and double mutant strains were grown and collected together and run on the same gel.
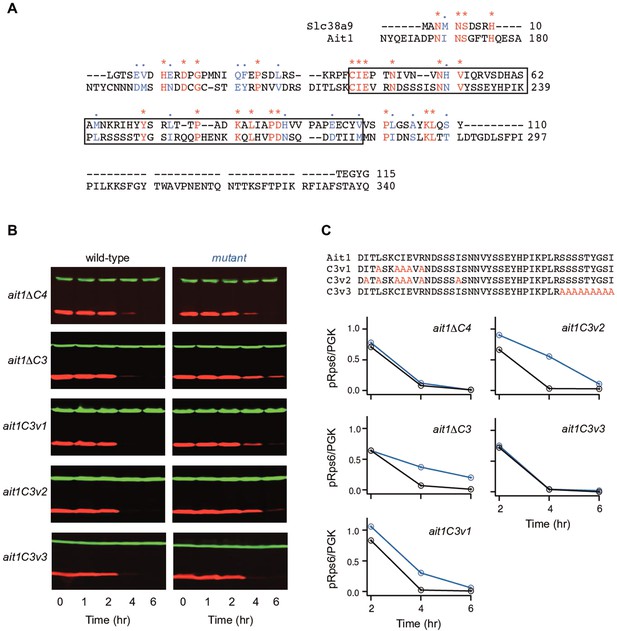
Role of the Ait1 C3 loop in target of rapamycin complex I (TORC1) regulation.
(A) BLAST alignment of the SLC38A9 (top) and Ait1 sequences, showing the entire C3 loop (no other sequences in these proteins align). (B, C) TORC1 activity in Ait1 C3 and C4 loop mutants during leucine starvation, measured as described in Figure 6. Values show the ratio of the p-Rps6 signal divided by the PGK (loading control) signal in each lane, relative to the value for the wild-type strain at time = 0. Mutant and wild-type strains were grown and collected together and run on the same gel.
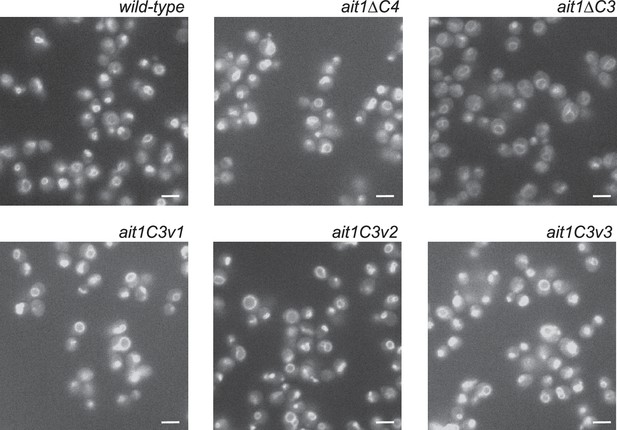
Localization of Ait1 and the Ait1 C3 and C4 loop mutants.
Images taken of strains expressing GFP-Ait1, GFP-AitΔC3, GFP-AitΔC4, GFP-Ait1C3v1, GFP-Ait1C3v2, and GFP-Ait1C3v3 show that the mutants fold and are transported to the vacuolar membrane correctly. Note, however, that the abundance of GFP-AitΔC3 is decreased to approximately 65% of that found in the wild-type (values from line-plot analysis of 10 cells in each image). The white bar shows 5 μm.
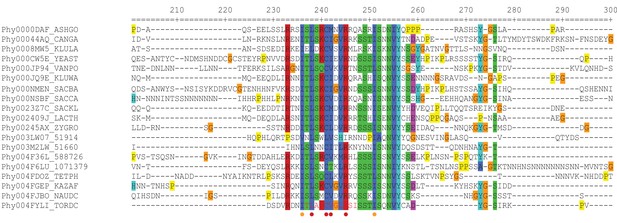
Sequence alignments of Ait1 from a variety of yeast species, showing the high degree of conservation at the center of the C3 loop.
Red dots show the sites mutated in Ait1C3v1 and red and yellow dots show the sites mutated in Ait1C3v2.
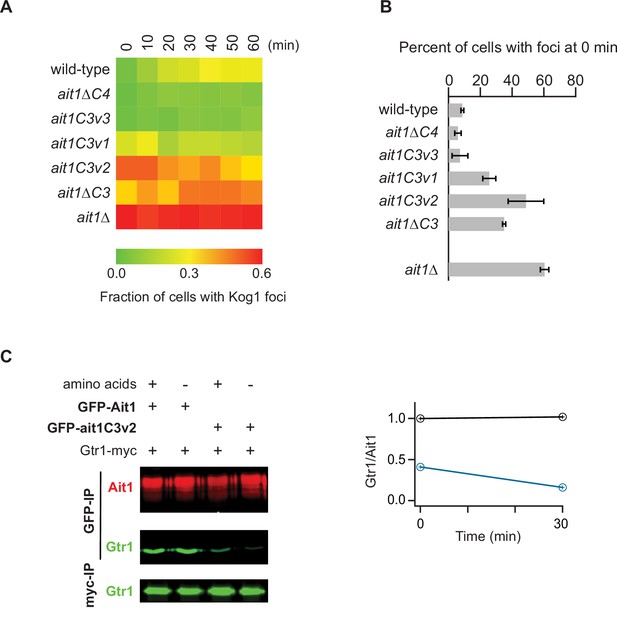
Impact of AIt1 C3 and C4 loop mutations on target of rapamycin complex I (TORC1) localization and Gtr1/2 binding.
(A) Each square on the heat map shows the fraction of cells with a Kog1-YFP focus/body at a specific timepoint (as labeled), calculated by averaging the data from three replicate experiments (>100 cells analyzed at each timepoint and replicate). Individual timepoints have errors ranging from 0.02 to 0.12 (average 0.06). (B) Bar graph showing the fraction of cells with a TORC1 body during log-phase growth in SD medium. (C) Coimmunoprecipitation showing a strong interaction between GFP-Ait1 and Gtr1-myc, but not GFP-Ait1C3v2 and Gtr1-myc. The graph shows the ratio of the Gtr1 and Ait1 signals in the wild-type (black line) and Ait1C3v2 (blue line) strains, before, and 30 min after, amino acid starvation. The Immunoprecipitation data with a full set of controls are shown in Figure 9—figure supplement 1.
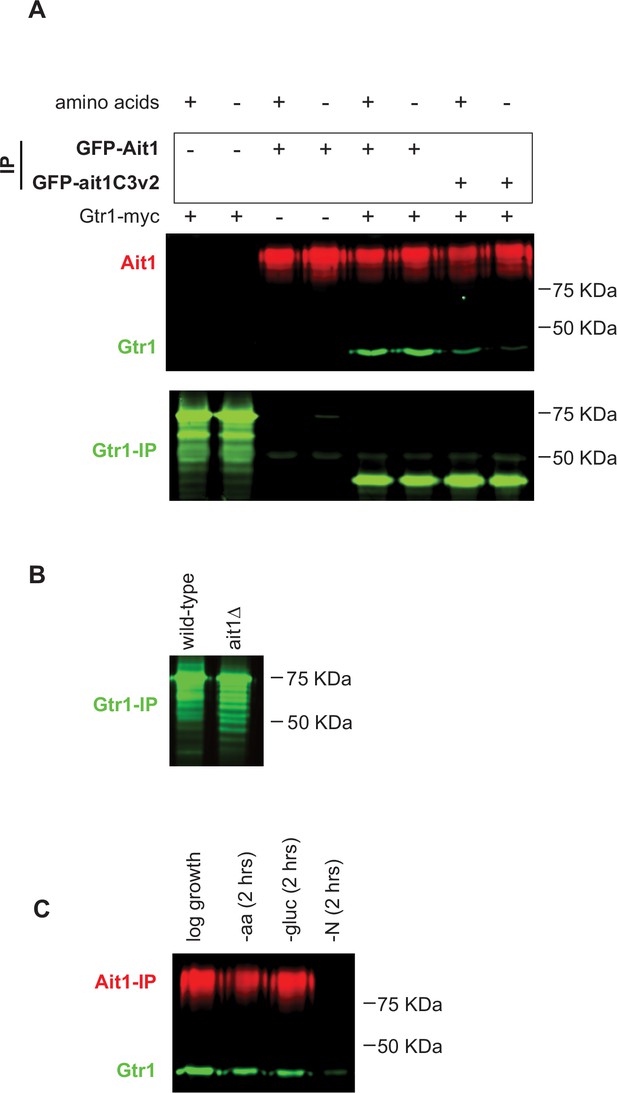
Coimmunoprecipitation of GFP-Ait1 and Gtr1-myc.
(A) GFP-Ait1 and GFP-Ait1C3v2 were immunopurified in strains with or without a myc-tag on Gtr1 (as labeled). As a control we also immunopurified Ait1 from a strain carrying Gtr1-myc, but without a GFP tag on Ait1. These experiments showed a clear interaction between Ait1 and Gtr1 (i.e., dependent on the C3 loop). However, Gtr1-myc was difficult to detect in the extract. Therefore, to test if Gtr1-myc levels are the same in the GFP-Ait1 and GFP-Ait1C3v2 strains we also carried out a separate immunopurification of Gtr1-myc (bottom panel). This revealed that there is a dramatic decrease in the apparent weight of Gtr1 in strains carrying GFP-Ait1 (unmodified Gtr1-myc is 38 kDa). Thus, Gtr1 is subject to post-translation modification (likely ubiquitination) and this modification is blocked by the GFP on the N-terminus of Ait1. (B) Deletion of Ait1 does not significantly impact the weight of Gtr1, and thus Ait1 does not regulate Gtr1/target of rapamycin complex I (TORC1) via the observed posttranslational modification. (C) Gtr1-myc copurifies with GFP-Ait1 even after long-term amino acid and glucose starvation, but at 60% of the level found in log-phase growth. We were unable to detect GFP-Ait1 after 2 hr of nitrogen starvation, likely due to a change in the solubility or localization of Ait1-Gtr1/2-TORC1.
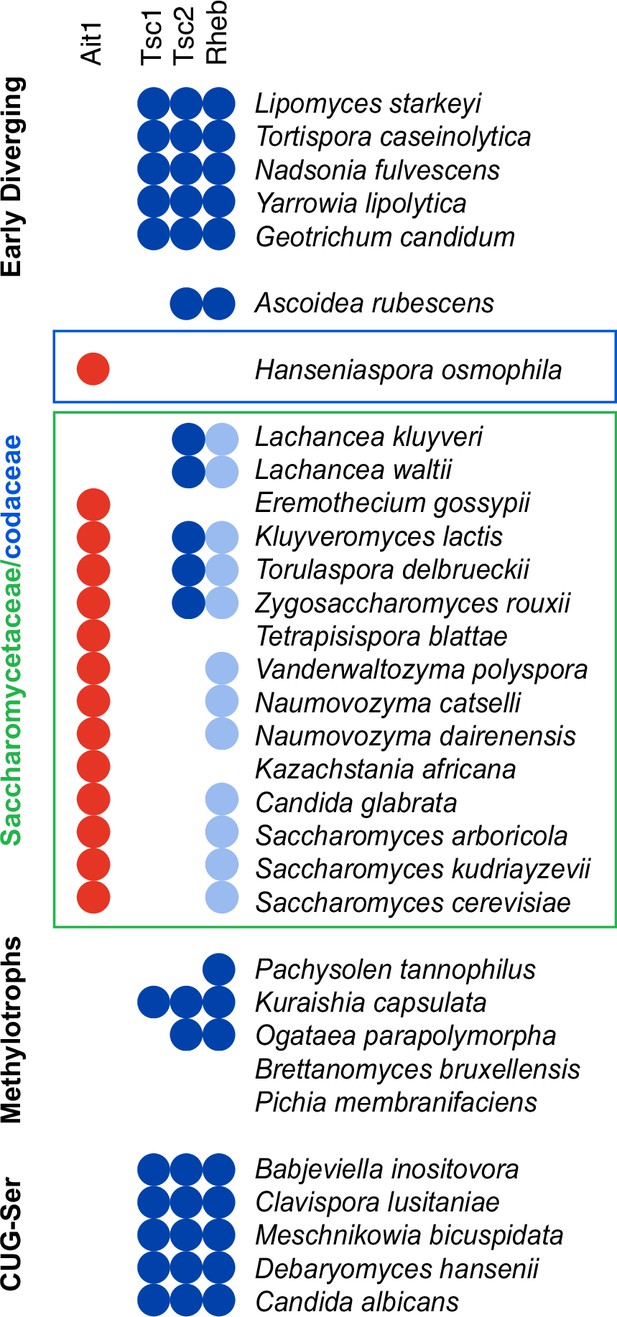
Evolution of the target of rapamycin complex I (TORC1) circuit in yeast.
Species identified as carrying Ait1 in a BLAST search (p < 0.001 cutoff) are marked with a red circle on a previously constructed map of Rheb and TSC1/2 conservation among the budding yeast, taken from Tatebe and Shiozaki, 2017. The light blue circles denote the presence of a highly divergent (nonfunctional) Rheb in species closely related to S. cerevisiae (Tatebe and Shiozaki, 2017). Ait1 was not detected in any of the yeasts outside the Saccharomycetaceae and Saccharomycodaceae.
Additional files
-
Supplementary file 1
Complete data for the Kog1 and Pib2 immunopurifications.
The IP-mock tab shows the number of background corrected peptide spectral counts (PSMs) for each protein detected (rows) across the different IPs (columns). The Filterpass tab shows the number of raw PSMs in each IP and mock IP. Proteins with at least twofold higher abundance in the true IP (Kog1-FLAG or GFP-Pib2) versus the mock IP (Kog1-HA or wild-type Pib2), and with at least seven peptide spectral maps in the true IP, were scored as potential interactors.
- https://cdn.elifesciences.org/articles/68773/elife-68773-supp1-v2.xlsx
-
Supplementary file 2
Results of a BLASTP search against Ydl180w from S. cerevisiae.
Subspecies/variants were removed from the table for clarity. No other families of yeast have species with significant E values (p < 0.01).
- https://cdn.elifesciences.org/articles/68773/elife-68773-supp2-v2.xlsx
-
Supplementary file 3
List of strains used in this study.
- https://cdn.elifesciences.org/articles/68773/elife-68773-supp3-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/68773/elife-68773-transrepform1-v2.docx
-
Source data 1
Labelled raw gel images.
The original labeled gel, from each panel in Figure 3—figure supplement 1, Figure 6, Figure 6—figure supplement 1, Figure 7, Figure 8, Figure 8—figure supplement 3, Figure 9, and Figure 9—figure supplement 1 are included in source data in two separate folders. In each case the gels are numbered as they are shown in the associated figure—from top to bottom. In the case of Figure 6, the two gels on the left are labeled 1 and 2 and the two gels on the right are labeled 3 and 4.
- https://cdn.elifesciences.org/articles/68773/elife-68773-data1-v2.zip
-
Source data 2
Labelled raw gel images.
- https://cdn.elifesciences.org/articles/68773/elife-68773-data2-v2.zip