Cardiac differentiation of human pluripotent stem cells using defined extracellular matrix proteins reveals essential role of fibronectin
Figures
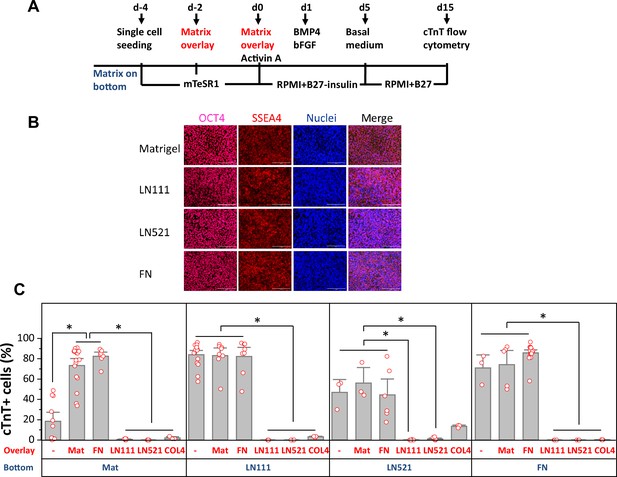
Defined ECM proteins support hPSC adhesion, growth and cardiac differentiation using the matrix sandwich protocol.
(A) Schematic method of the matrix sandwich protocol. Defined ECM proteins were tested as coating matrix (blue) and overlay matrix (red). (B) Fluorescence images of DF19-9-11T iPSCs growing on the ECM of Matrigel, LN111, LN521 and FN as confluent monolayer and immuno-labeled with antibodies against OCT4 and SSEA4. Scale bar is 100 µm. (C) cTnT+ cells measured by flow cytometry at 15 days of differentiation of DF19-9-11T iPSCs on different ECM proteins as substrate (bottom) and overlay (overlay). N≥3 biological replicates. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
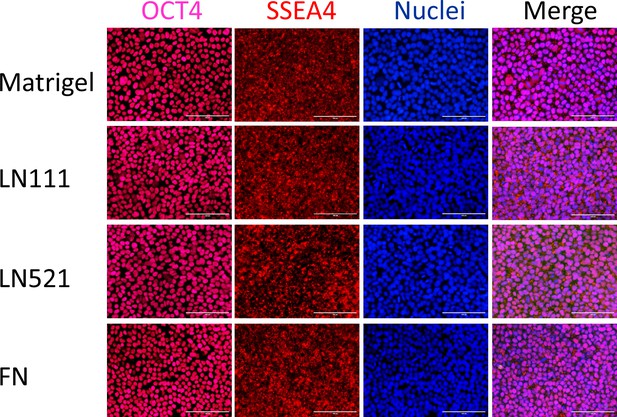
Fluorescence images of H1 ESCs growing on Matrigel, LN111, LN521 and FN coated surface as confluent monolayer immunolabeled with antibodies against OCT4 and SSEA4.
Scale bar is 100 µm.
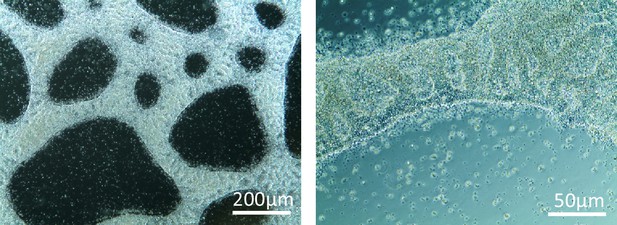

Morphology of DF19-9-11 iPSCs growing on human collagen IV coated surface.
The iPSCs were grown for 5 days on collagen IV coated surface and did not form confluent monolayer.
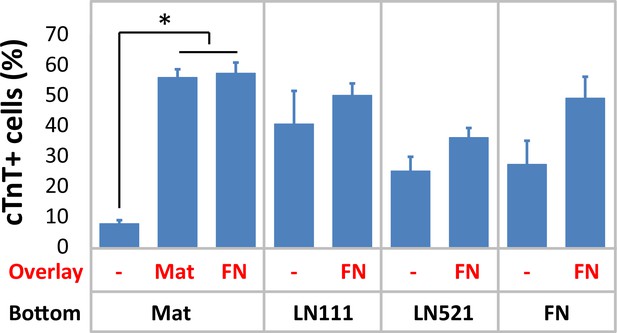
cTnT+ cells measured by flow cytometry at 15 days of differentiation of H1 ESCs on different ECM proteins as substrate (bottom) and overlay (overlay) using the matrix sandwich protocol.
N≥3 biological replicates. *p<0.05, one-way ANOVA with post-hoc Bonferroni and test.
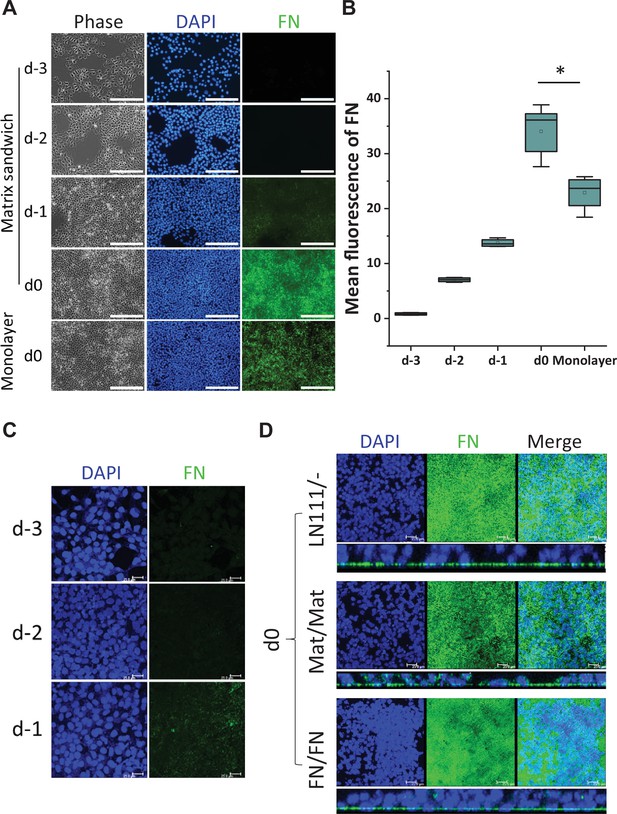
Production of endogenous FN in the hPSC-matrix sandwich culture and LN111 culture.
(A) Phase contrast and fluorescence images of the matrix sandwich culture of DF19-9-11T iPSCs grown for 4 days and immunolabeled using anti-FN antibody, compared with the monolayer culture. Scale bar is 200 µm. (B) Quantitative analysis of the FN fluorescence in A by Image J. N≥3 replicates. The box plots summarize the biological replicates with the box enclosing from first to third quartile and middle square indicating mean and line in box indicating median. *p<0.05, one-way ANOVA with post-hoc Bonferroni test. (C) Maximum projection view of the confocal z-scan of DF19-9-11T iPSCs growing on LN111 coated surface immunolabeled with FN antibody at day -3, -2 and -1. Scale bar is 25 µm. (D) The maximum projection view (upper panel) and side view (lower panel) of the confocal z-scan of DF19-9-11T iPSCs grown for 4 days (at day 0) on LN111 coated surface without matrix overlay immunolabeled with the anti-FN antibody. The multilayer growth and FN production are similar to Matrigel/Matrigel and FN/FN matrix sandwich cultures in parallel at the same time (day 0). Scale bar is 25 µm.
-
Figure 2—source data 1
Images and quantitative analysis for Figure 2B.
- https://cdn.elifesciences.org/articles/69028/elife-69028-fig2-data1-v1.zip
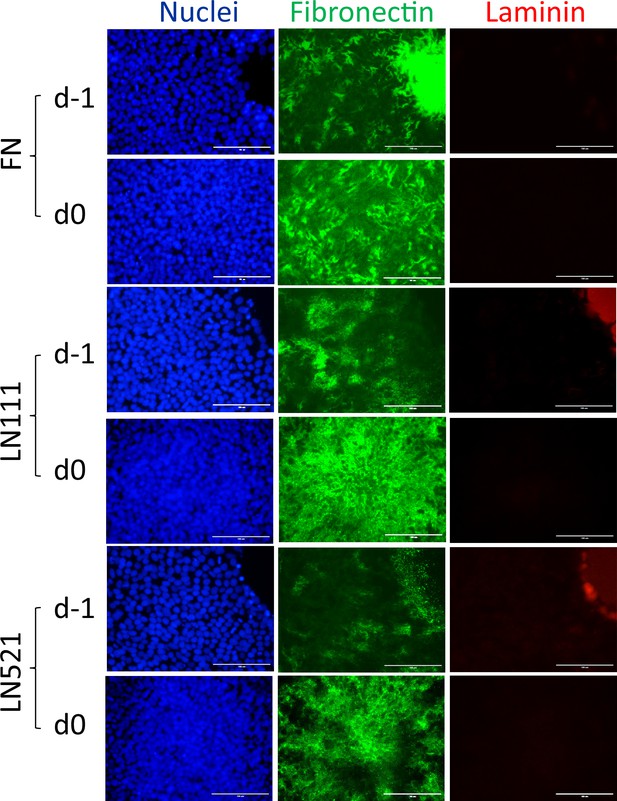
Fluorescence images of H1 ESCs growing on FN, LN111 and LN521 coated surface immunolabeled with antibodies against FN and laminin.
Scale bar is 100 µm.
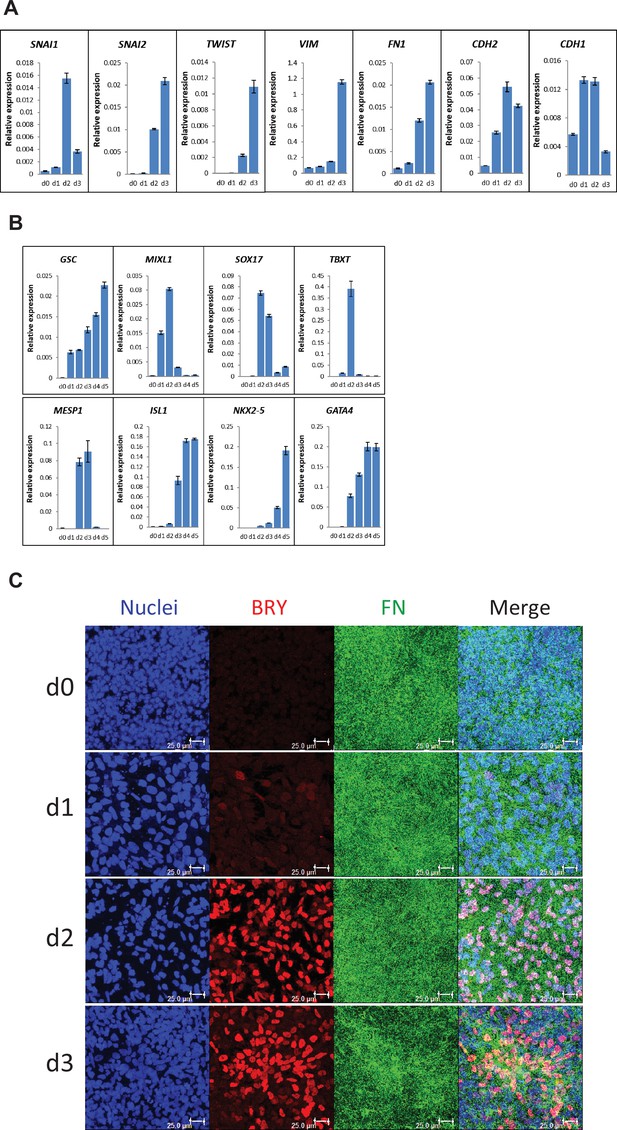
Expression of EMT, mesendoderm/mesoderm markers and cardiac transcription factors in the cardiac differentiation of hPSCs cultured on LN111 substrate by Activin A/BMP4/bFGF signaling.
(A) qRT-PCR for gene expression of EMT markers at days 0–3 of cardiac differentiation. (B) qRT-PCR for gene expression of mesendoderm/mesoderm and cardiac transcription factors at days 0–5 of cardiac differentiation. N=3 technical replicates for each point. (C) Maximum projection view of the confocal z-scan of DF19-9-11T iPSCs at days 0–3 of the cardiac differentiation co-labeled with antibodies against Brachyury (BRY) and FN. Scale bar is 25 µm. Error bars represent SEM.
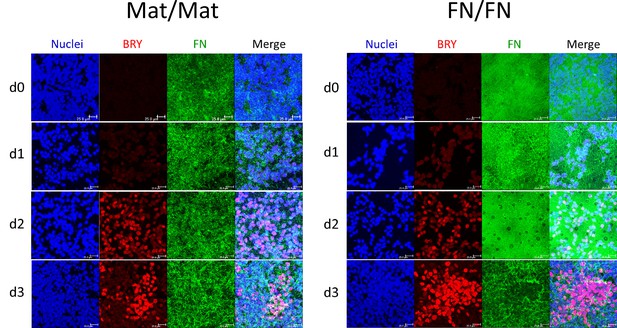
Maximum projection view of the confocal z-scan of DF19-9-11T iPSCs at days 0–3 of the cardiac differentiation using the Matrigel/Matrigel (Mat/Mat) and FN/FN matrix sandwich protocol.
The cells were co-labeled with antibodies against Brachyury (BRY) and FN. Scale bar is 25 µm.
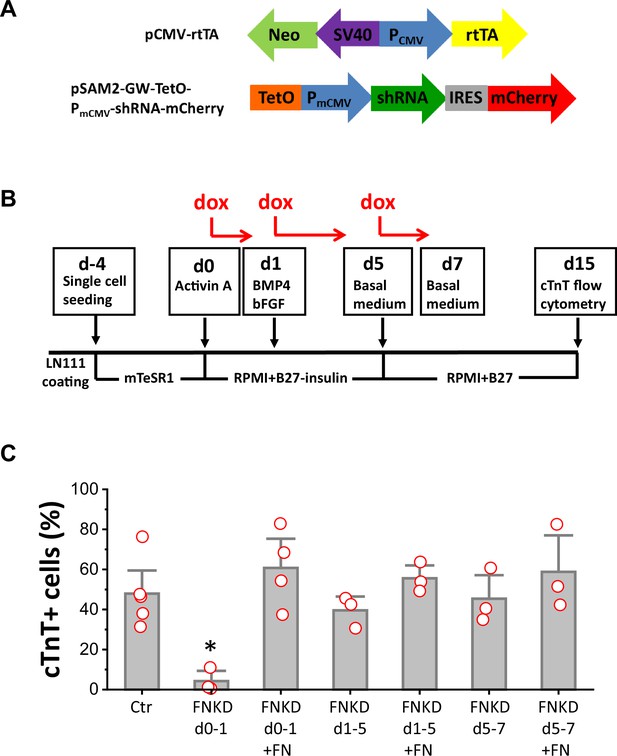
FN is essential at the initiation of cardiac differentiation of hPSCs.
(A) Schematic of the inducible shRNA construct for FN1 knockdown. (B) Schematic method of FN knockdown at differentiation stages of days 0–1, 1–5, and 5–7 in the cardiac differentiation protocol. (C) cTnT+ cells measured by flow cytometry at 15 days of differentiation of the H1 FN1 knockdown clone 34 using the protocol in (B). Dox concentration is 2 µg/ml, N≥3 biological replicates. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test. FNKD indicates FN knockdown by dox induction. +FN indicates exogenous FN added.

The FN1 shRNA oligos.
Yellow, CDS where shRNA binds; Green, palindrome. Oligo-1 binds in exon 8; Oligo-2 binds in 3’ UTR.
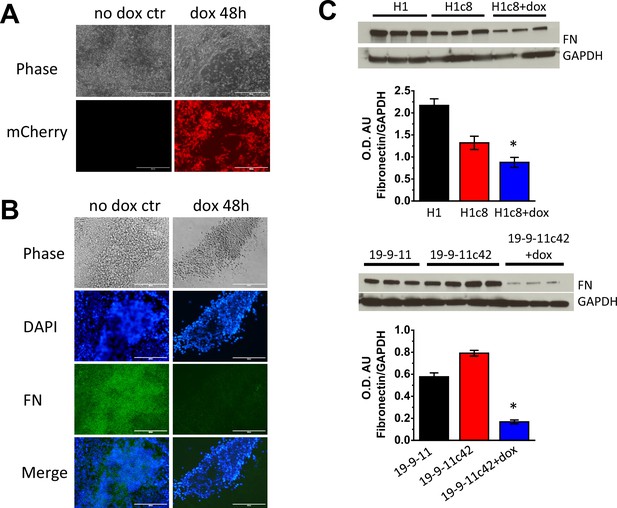
Characterization of hPSC FN1 knockdown clones.
(A) Doxycycline-induced mCherry expression in the hPSC culture of H1 FN1 knockdown clone. Scale bar is 200 µm. (B) Phase contrast and immunofluorescence images of FN expression in doxycycline hPSC culture of H1 FN1 knockdown clone. Scale bar is 200 µm. (C) Quantatitave western blots of FN from control of H1 and DF19-9-11T hPSCs and associated FN1 knockdown hPSC clones in the absence and presence of Dox (8 µg/ml). N=3 biological replicates. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
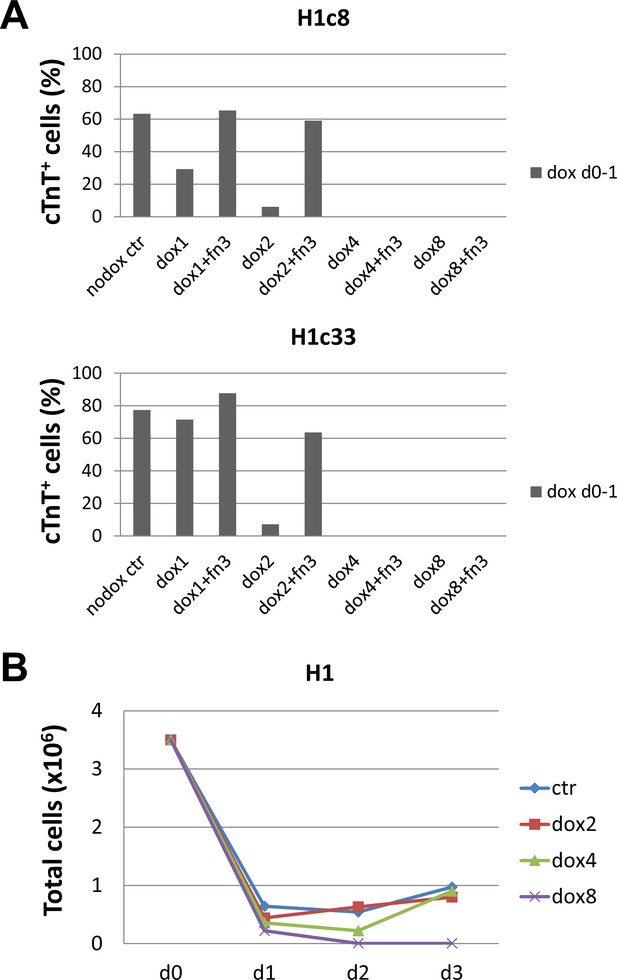
Effect of dox concentration-dependent FN knockdown at days 0–1 on cardiac differentiation using the protocol as shown in Figure 4A.
(A) cTnT+ cells measured by flow cytometry at 15 days of differentiation of the FN1 knockdown clones (H1c8 and H1c34) showing the effect of FN knockdown at days 0–1 at different concentrations of dox (1–8 μg/ml) and the effect of adding exogenous FN (3 μg/cm2) at each concentration of dox. Dox added at 2 μg/ml inhibited generation of cTnT+ cells which was rescued by added exogenous FN. Dox added at 4 µg/ml or higher inhibited cTnT+ cells and could not be rescued by exogenous FN under the conditions tested. (B) Total cells in culture for the unmodified control H1 hESC line undergoing cardiac differentiation measured at days 0–3 when dox was added at days 0–1. Dox concentrations of 0 (no dox ctr), 1 (dox1), 2 (dox2), 4 (dox4), and 8 (dox8) μg/ml were tested. At the highest concentration of dox tested (8 μg/ml) no cells survived.
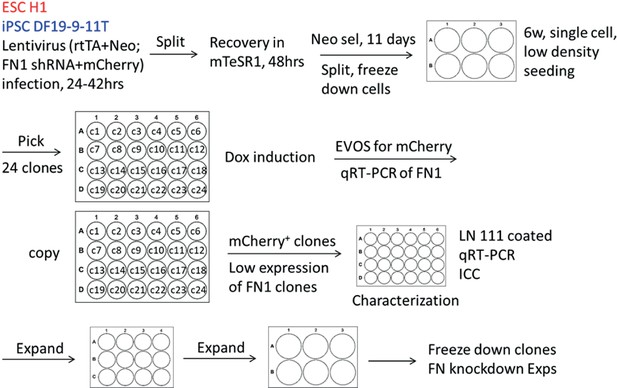
The cloning strategy in generation of inducible FN1 knockdown hPSC clones.
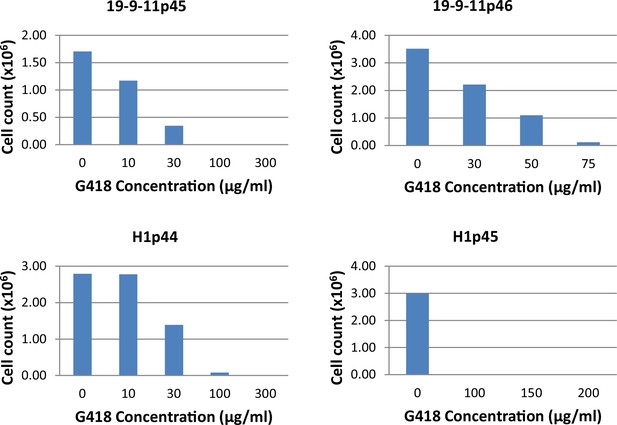
The concentrations of G418 tested for neo-resistant selection of hPSC clones.
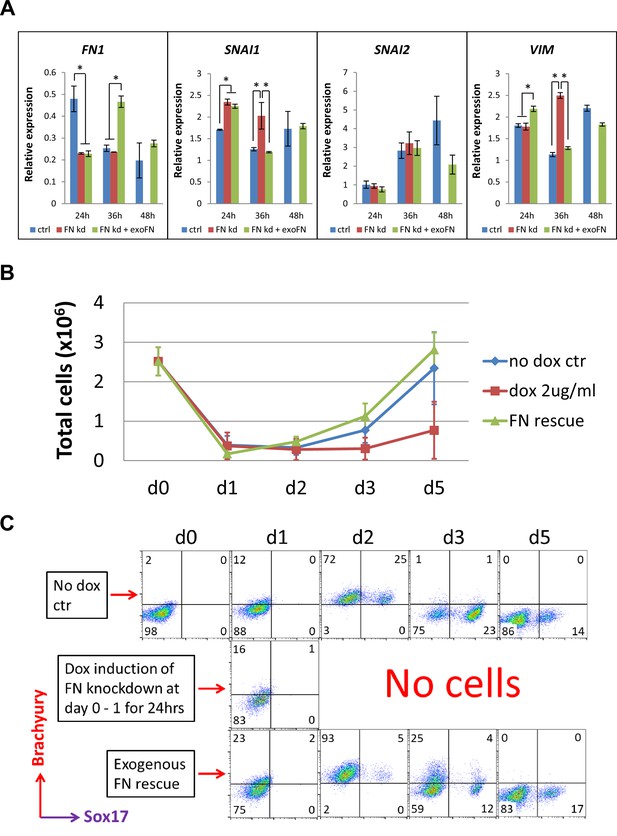
shRNA knockdown of FN results in loss of Brachyury+ cells.
(A) qRT-PCR for gene expression of EMT markers for the H1 FN knockdown clone in the cardiac differentiation time course of 0–48 hr at the no dox control, dox induction at day 0–1 and dox induction at day 0–1 with adding exogenous FN conditons. (B) Total cell number of the H1 FN knockdown clones in the time course of days 0–5 in the cardiac differentiation at the no dox control, dox induction at day 0–1 and dox induction at day 0–1 with adding exogenous FN conditions. (C) Flow cytometry of co-labeling the cells shown in B with Brachyury and Sox17 antibodies. The cardiac differentiation protocol is shown in Figure 4B. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
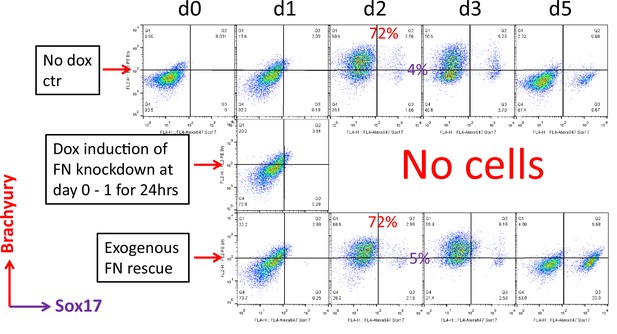
Flow cytometry co-labeling cells with antibodies to Brachyury and Sox17 using 19-9-11 FN knockdown clone 42 in the cardiac differentiation protocol at days 0–5, comparing no dox control, dox induction at day 0–1 and dox induction at day 0–1 with addition of exogenous FN conditions.
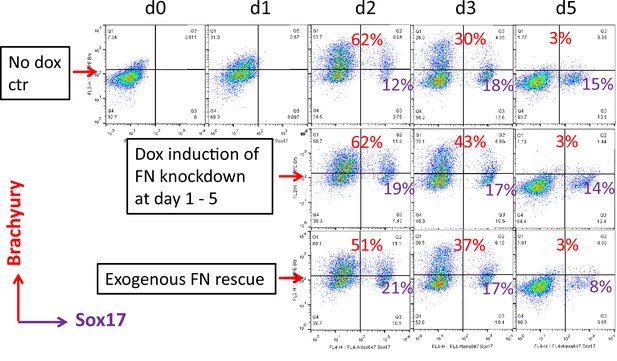
Flow cytometry co-labeling cells with antibodies to Brachyury and Sox17 using H1 FN knockdown clone 8 in the cardiac differentiation protocol at days 0–5, comparing no dox control, dox induction at days 1–5 and dox induction at days 1–5 with addition of exogenous FN conditions.
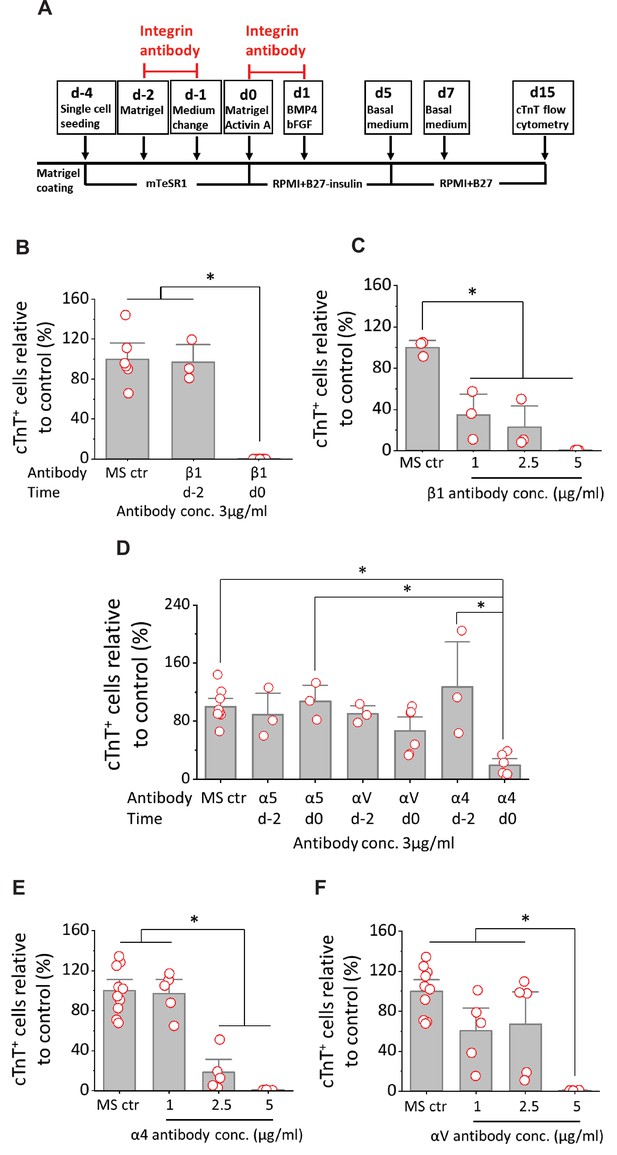
Cardiac differentiation is blocked by anti-integrin β1, α4 or αV antibodies when added at mesoderm formation in the matrix sandwich protocol.
(A) Schematic for testing monoclonal antibodies to block integrin β1 (P5D2), α5 (P1D6), αV (P3G8), and α4 (P4G9). (B) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin β1 antibody was added on day –2 or day 0 at 3 µg/ml in the matrix sandwich protocol as shown in A. (C) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin β1 antibody was added on day 0 at concentrations of 1, 2.5, and 5 µg/ml in the matrix sandwich protocol as shown in A. (D) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin α5, αV or α4 antibody was added on day –2 or day 0 at 3 µg/ml in the matrix sandwich protocol as shown in A. (E) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin α4 antibody was added on day 0 at concentrations of 1, 2.5, and 5 µg/ml in the matrix sandwich protocol. (F) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin αV antibody was added on day 0 at concentrations of 1, 2.5, and 5 µg/ml in the matrix sandwich protocol. The % of cTnT+ cells in each group was normalized to the control group to combine replicates from multiple differentiations and to compare across different antibodies blocking. N≥3 biological replicates. Data are from DF19-9-11T iPSC line. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
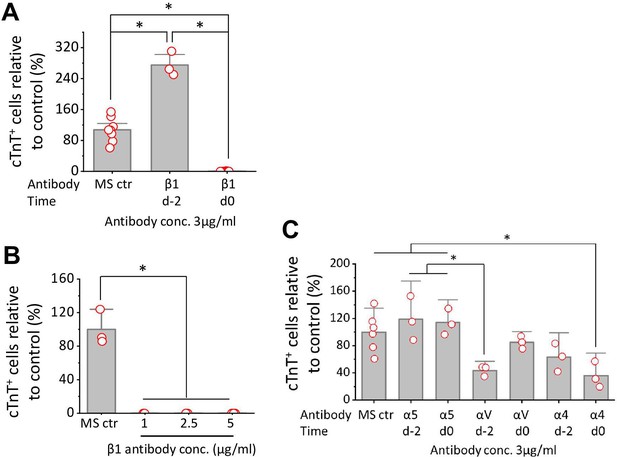
Cardiac differentiation of H1 ESCs is blocked by anti-integrin β1, α4, or αV antibody when added on day 0 in the matrix sandwich protocol.
(A) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin β1 antibody was added on day –2 or day 0 at 3 µg/ml in the matrix sandwich protocol. (B) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin β1 antibody was added on day 0 at concentrations of 0 (Ctr), 1, 2.5, and 5 µg/ml in the matrix sandwich protocol. (C) cTnT+ cells measured by flow cytometry at 15 days differentiation when anti-human integrin α5, αV, or α4 antibody was added on day –2 or day 0 at 3 µg/ml in the matrix sandwich protocol. The % of cTnT+ cells in each group was normalized to the control group without antibody blocking to combine replicates in multiple differentiation and to compare across different antibodies blocking. N≥3 biological replicates. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
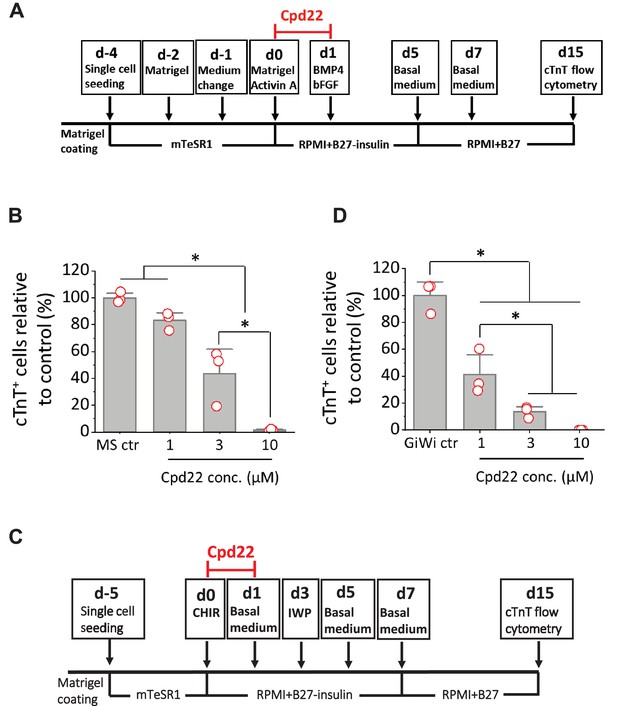
Cardiac differentiation is inhibited by ILK inhibitor in the matrix sandwich and GiWi protocols.
(A) Schematic to test inhibition of ILK by small molecule inhibitor Cpd22 in the matrix sandwich protocol. (B) cTnT+ cells measured by flow cytometry at 15 days differentiation when Cpd22 was added at day 0 at concentrations of 1, 3, and 10 µM in the matrix sandwich protocol as shown in A. (C) Schematic for testing ILK inhibitor Cpd22 in the GiWi protocol. (D) cTnT+ cells measured by flow cytometry at 15 days differentiation when Cpd22 was added on day 0 in the GiWi protocol as shown in C. The % of cTnT+ cells in each group was normalized to the control group to combine replicates in multiple differentiations and to compare across different antibodies. N≥3 biological replicates. Data are from DF19-9-11T iPSC line. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
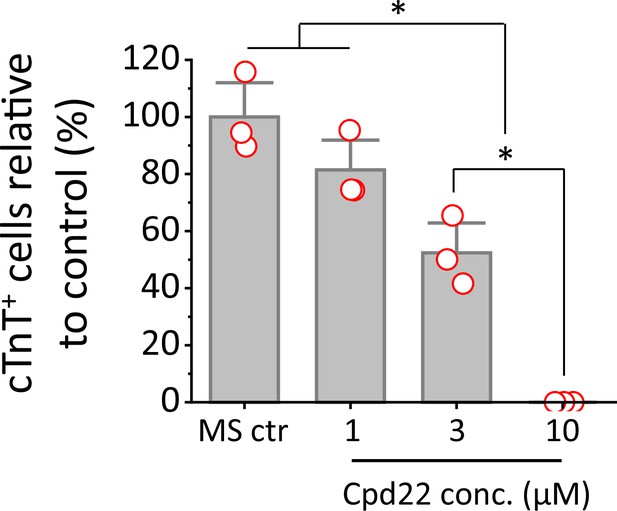
Cardiac differentiation of H1 ESCs is inhibited by ILK inhibitor, cpd22 when added on day 0 at concentrations of 0, 1, 3 and 10 uM in the matrix sandwich protocol.
The % of cTnT+ cells in each group was normalized to the control group to combine replicates in multiple differentiation. N=3 biological replicates. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
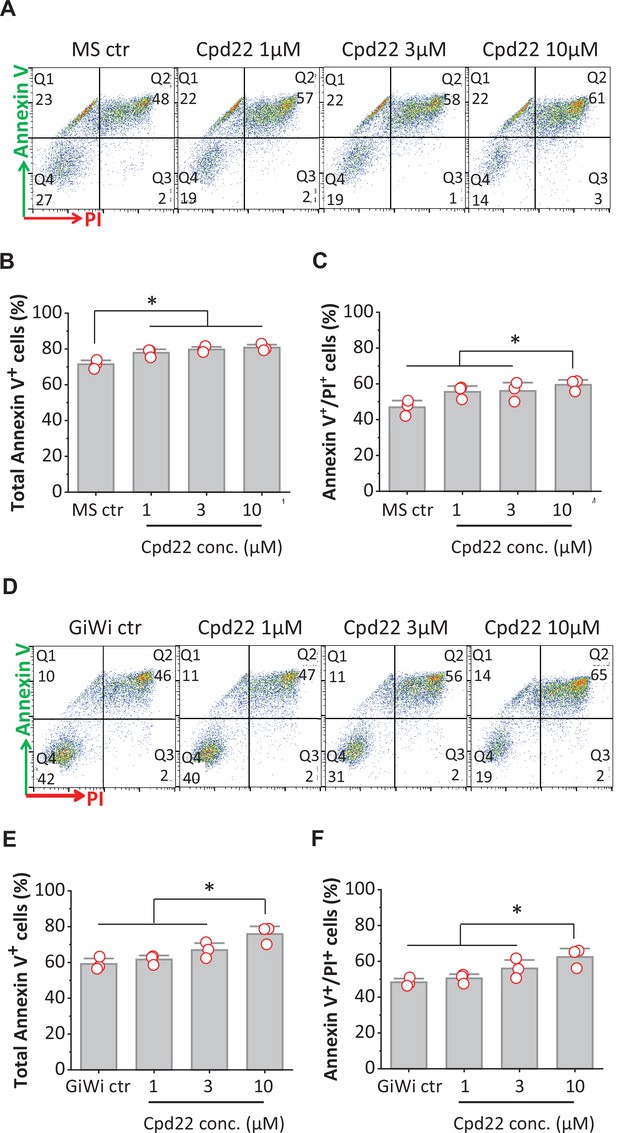
Inhibition of ILK promotes apoptosis of differentiating hPSCs at mesoderm formation in the matrix sandwich and GiWi protocols.
(A, D) Representative flow cytometry plots of cells collected at 18 hr of differentiation, labeled by an antibody recognizing AnnexinV and stained by propidium iodide (PI) in the Matrix Sandwich (MS) protocol (A) and GiWi protocol (D) in absence or presence of the ILK inhibitor cpd22. (B, C) Average total apoptotic cells (Annexin V+) and late apoptotic cells (Annexin V+/PI+) in the MS protocol in absence or presence of cpd22. (E, F) Average of total apoptotic cells of (Annexin V+) and late apoptotic cells (Annexin V+/PI+) in the GiWi protocol in absence or presence of cpd22. N=3 biological replicates. Data are from DF19-9-11T iPSC line. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
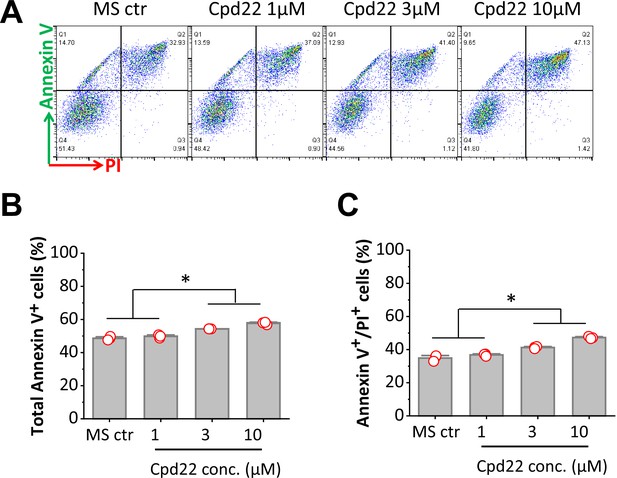
Inhibition of ILK promotes apoptosis of differentiating H1 ESCs at the mesoderm formation in the Matrix Sandwich protocol.
(A) Representative plots of flow cytometry of the cells collected at 18 hr of differentiation in absence or presence of cpd22 at concentrations of 1, 3, and 10 µM. (B, C) Average of total apoptotic cells of Annexin V+ (B) and late apoptotic cells of Annexin V+/PI+ (C) in absence (MS ctr) or presence of cpd22 at concentrations of 1, 3, and 10 µM. N=3 biological replicates. Error bars represent SEM. *p<0.05, one-way ANOVA with post-hoc Bonferroni test.
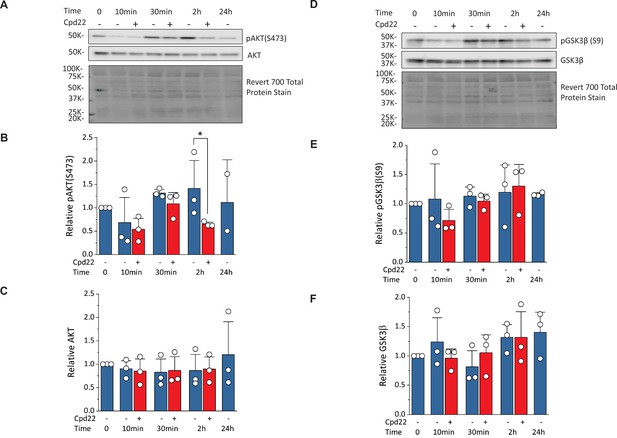
Inhibition of ILK reduces pAKT(Ser473) without changing pGSK3β(Ser9) at mesoderm formation in the matrix sandwich protocol.
(A) Representative western blot analysis of pAKT(Ser473) and total AKT as well as total protein staining from day 0 (Time 0) to day 1 (24 hr) of the matrix sandwich protocol with or without cpd22 treatment. (B, C) Densitometric analysis of pAKT(Ser473) (B) and AKT (C) normalized to total protein and plotted relative to time 0 for three separate experiments. (D) Representative Western blot analysis of pGSK3β(Ser9) and total GSK3β as well as total protein staining from day 0 (Time 0) to day 1 (24 hr) of the matrix sandwich protocol with or without cpd22 treatment. (E, F) Densitometric analysis of pGSK3β(Ser9) (E) and total GSK3β (F) normalized to total protein and plotted relative to time 0 for three separate experiments. Data in (B, C, E, F) are presented as mean ± SEM. *p<0.05, paired T-test comparing with and without cpd22.
-
Figure 9—source data 1
Raw unedited gels and blots and images with the uncropped gels and blots for all three experiements performed with the relevant bands labelled.
- https://cdn.elifesciences.org/articles/69028/elife-69028-fig9-data1-v1.zip
Additional files
-
Supplementary file 1
Tables of primary antibodies and qRT-PCR primers.
(a) Primary antibodies used in immunocytochemistry (ICC) and flow cytometry (FC). (b) Primers for quantitative RT-PCR. (c) Primary antibodies used for immunoblotting.
- https://cdn.elifesciences.org/articles/69028/elife-69028-supp1-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/69028/elife-69028-transrepform1-v1.pdf





