Thymocytes trigger self-antigen-controlling pathways in immature medullary thymic epithelial stages
Figures
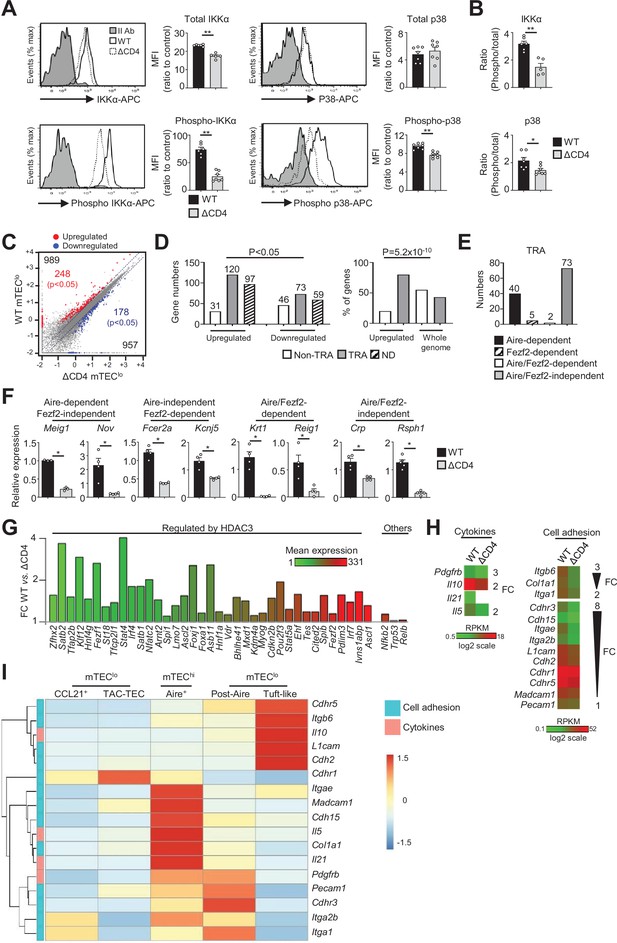
The transcriptional profile and IKKα and p38 MAPK signaling pathways are impaired in mTEClo of ΔCD4 mice.
(A, B) Total IKKα, p38 MAPK, phospho-IKKα(Ser180)/IKKβ(Ser181), and p38 MAPK (Thr180/Tyr182) (A) and the ratio of phospho/total proteins (B) analyzed by flow cytometry in mTEClo from WT and ΔCD4 mice. Data are representative of two independent experiments (n = 3–4 mice per group and experiment). (C) Scatter plot of gene expression levels (fragments per kilobase of transcript per million mapped reads [FPKM]) of mTEClo from WT versus ΔCD4 mice. Genes with fold difference ≥2 and p-adj<0.05 were considered as upregulated or downregulated genes (red and blue dots, respectively). RNA-seq was performed on two independent biological replicates with mTEClo derived from 3 to 5 mice. (D) Numbers of tissue-restricted self-antigens (TRAs) and non-TRAs in genes up- and downregulated (left panel) and the proportion of upregulated TRAs compared to those in the all genome (right panel). ND, not determined. (E) Numbers of induced Aire-dependent, Fezf2-dependent, Aire/Fezf2-dependent, and Aire/Fezf2-independent TRAs. (F) The expression of Aire-dependent (Meig1, Nov), Fezf2-dependent (Fcer2a, Kcnj5), Aire/Fezf2-dependent (Krt1, Reig1), and Aire/Fezf2-independent (Crp, Rsph1) TRAs measured by qPCR in WT (n = 3–4) and ΔCD4 (n = 3–4) mTEClo. (G) Expression fold change in HDAC3-induced transcriptional regulators and other transcription factors significantly upregulated in WT versus ΔCD4 mTEClo. The color code represents gene expression level. (H) Heatmaps of genes encoding for cell adhesion molecules and cytokines that were significantly downregulated in mTEClo from ΔCD4 mice. (I) Hierarchical clustering and heatmap of mean expression of these cell adhesion molecules and cytokines in mTEC subsets identified by scRNA-seq. Error bars show mean ± SEM, *p<0.05, **p<0.01 using two-tailed Mann–Whitney test for (A), (B) and (F) and chi-squared test for (D).
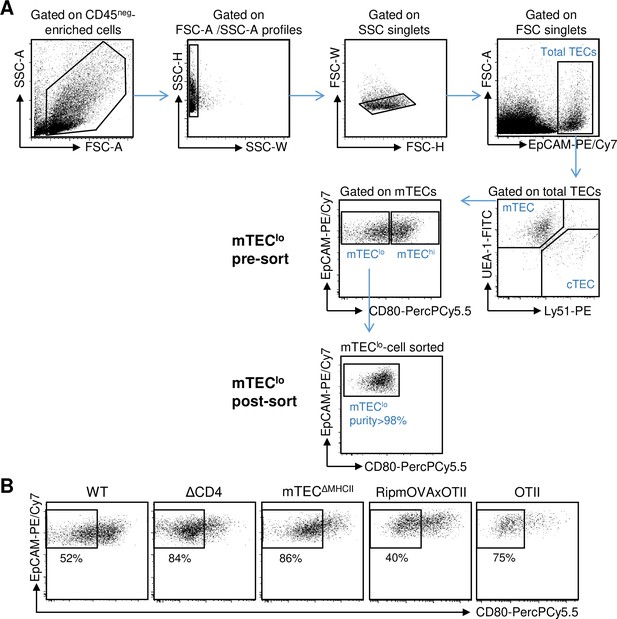
Gating strategy used to purify mTEClo cells.
(A) Total thymic epithelial cells (TECs) were defined as EpCAM+ in CD45-negative enriched thymic cells by autoMACS and were further divided into medullary TECs (mTECs) (UEA-1+Ly51lo) and cortical TECs (cTECs) (UEA-1−Ly51hi). mTEClo were identified and sorted based on low/intermediate levels of the CD80 co-stimulatory molecule. The purity of sorted mTEClo was >98%. (B) Gates used to sort mTEClo from WT, ΔCD4, mTECΔMHCII, RipmOVAxOTII-Rag2-/-, and OTII-Rag2-/- mice.
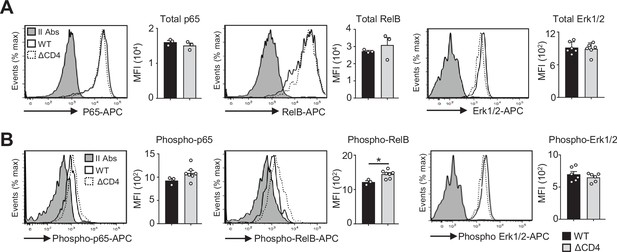
Normal total and phosphorylated p65, RelB, and Erk1/2 proteins in mTEClo from ΔCD4 mice.
Total p65, RelB, and Erk1/2 (A) and phospho-p65 (Ser536), phospho-RelB (Ser552), and phospho-Erk1/2 (Thr202/Tyr204) (B) proteins were analyzed by flow cytometry in mTEClo of WT and ΔCD4 mice. Histograms show the MFI. II Abs: secondary antibodies. Data are representative of two independent experiments (n = 2–3 mice per group and experiment). Error bars show mean ± SEM, *p<0.05 using the Mann–Whitney test.
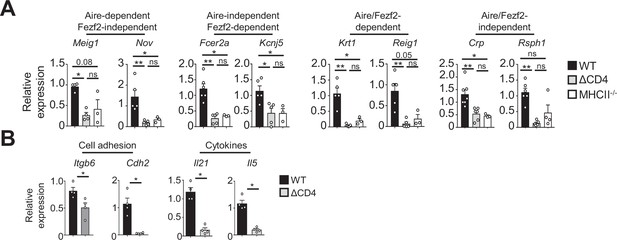
Impaired TRA expression in mTEClo from MHCII-/- mice.
(A) The expression of Aire-dependent (Meig1, Nov), Fezf2-dependent (Fcer2a, Kcnj5), Aire/Fezf2-dependent (Krt1, Reig1), and Aire/Fezf2-independent (Crp, Rsph1) tissue-restricted self-antigens (TRAs) was measured by qPCR in purified mTEClo from WT (n = 3), ΔCD4 (n = 3), and MHCII-/- (n = 3) mice. (B) Itgb6, Cdh2, Il21, and Il5 mRNAs were measured by qPCR in WT (n = 4) and ΔCD4 (n = 4) mTEClo.
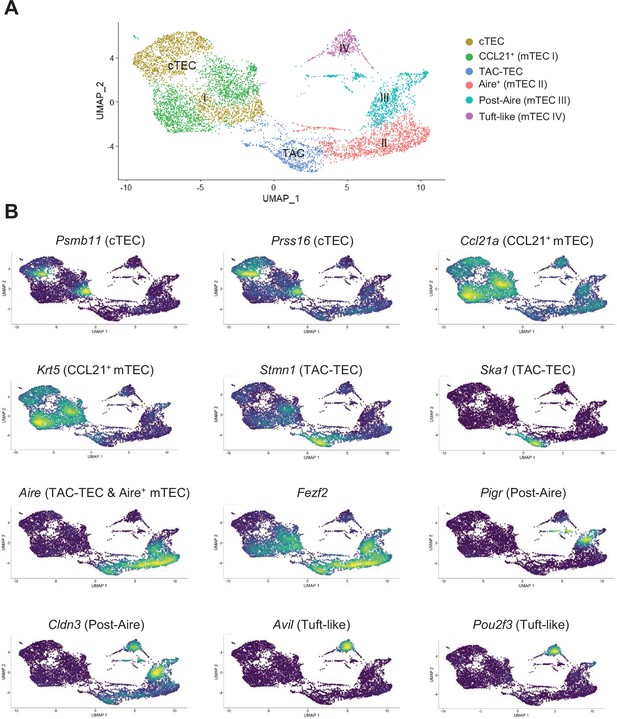
Identification of thymic epithelial cell (TEC) subsets by single-cell RNA-seq.
(A) UMAP visualization of single-cell RNA-seq data on TECs reanalyzed from Wells et al., 2020. Six clusters were identified corresponding to cortical TECs (cTECs), CCL21+ medullary TECs (mTECs) (mTEC I), TAC-TECs (transit-amplifying cells), Aire+ (mTEC II), post-Aire (mTEC III), and Tuft-like (mTEC IV) mTECs. (B) Expression of selected marker genes overlaid on UMAP visualization. Scale bars represent the log2 expression of the indicated genes.
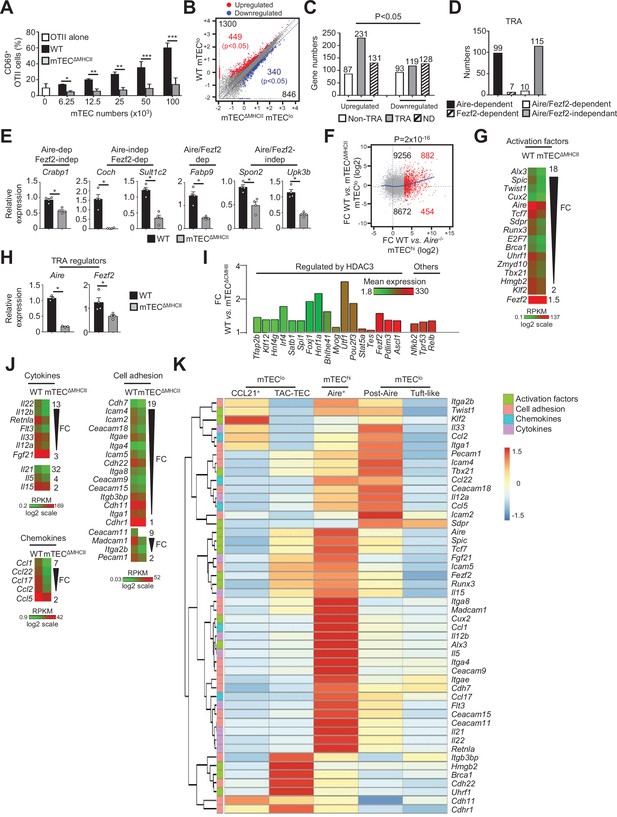
The transcriptional and functional properties of mTEClo are impaired in mTECΔMHCII mice.
(A) Percentages of CD69+ OTII CD4+ T cells cultured or not with variable numbers of OVA323-339-loaded WT or mTECΔMHCII mTECs derived from two independent experiments (n = 2–3 mice per group and experiment). (B) Scatter plot of gene expression levels (fragments per kilobase of transcript per million mapped reads [FPKM]) of mTEClo from WT versus mTECΔMHCII mice. Genes with fold difference ≥2 and p-adj<0.05 were considered as upregulated or downregulated genes (red and blue dots, respectively). RNA-seq was performed on two independent biological replicates with mTEClo derived from 3 to 5 mice. (C) Numbers of tissue-restricted self-antigens (TRAs) and non-TRAs in genes up- and downregulated in mTEClo from WT versus mTECΔMHCII mice. ND, not determined. (D) Numbers of induced TRAs regulated or not by Aire and/or Fezf2. (E) Aire-dependent (Crabp1), Fezf2-dependent (Coch, Sult1c2), Aire/Fezf2-dependent (Fabp9), and Aire/Fezf2-independent (Spon2, Upk3b) TRAs were measured by qPCR in mTEClo from WT (n = 4) and mTECΔMHCII (n = 4) mice. (F) Scatter plot of gene expression variation in mTEClo from WT versus mTECΔMHCII mice and in mTEChi from WT versus Aire-/- mice. The loess fitted curve is shown in blue and the induced Aire-dependent genes (fold change [FC] > 5) in red. (G) Heatmap of significantly downregulated activation factors in mTEClo from mTECΔMHCII mice. (H) Aire and Fezf2 mRNAs were measured by qPCR in mTEClo from WT (n = 3–4) and mTECΔMHCII (n = 4) mice. (I) FC in the expression of HDAC3-induced transcriptional regulators and other transcription factors significantly upregulated in WT versus mTECΔMHCII mice. The color code represents gene expression level. (J) Heatmap of significantly downregulated cytokines, chemokines, and cell adhesion molecules in mTEClo from mTECΔMHCII mice. (K) Hierarchical clustering and heatmap of mean expression of these activation factors, cell adhesion molecules, chemokines, and cytokines in mTEC subsets identified by scRNA-seq. Error bars show mean ± SEM, *p<0.05, **p<0.01, ***p<0.001 using two-tailed Mann–Whitney test for (A), (E) and (H) and chi-squared test for (C) and (F).
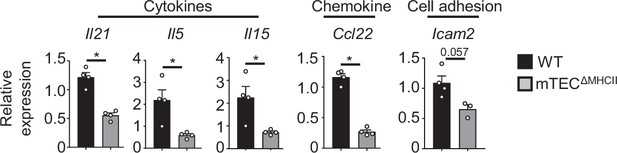
Altered expression of some cytokines, cell adhesion molecules, and chemokines in mTEClo from mTECΔMHCII mice.
Il21, Il5, Il15, Ccl22, and Icam2 mRNAs were measured by qPCR in mTEClo from WT (n = 4) and mTECΔMHCII (n = 3–4) mice. Error bars show mean ± SEM, *p<0.05 using two-tailed Mann–Whitney test.
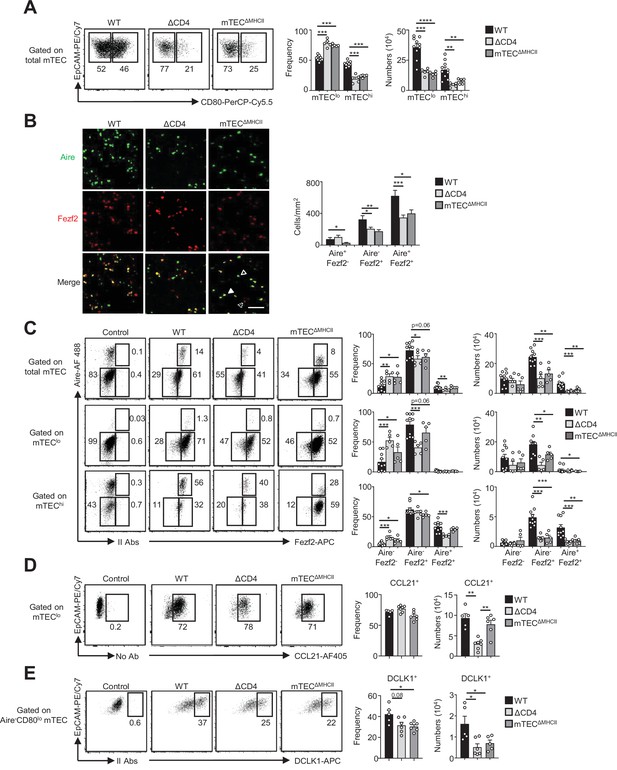
The composition in medullary thymic epithelial cell (mTEC) subsets is altered in ΔCD4 and mTECΔMHCII mice.
(A) Flow cytometry profiles, frequencies, and numbers of mTEClo and mTEChi in WT, ΔCD4, and mTECΔMHCII mice. Data are representative of 2–3 independent experiments (n = 2–5 mice per group and experiment). (B) Confocal images of thymic sections from WT, ΔCD4, and mTECΔMHCII mice stained for Aire (green) and Fezf2 (red). 12 and 20 sections derived from two WT, two ΔCD4, and two mTECΔMHCII mice were quantified. Scale bar, 50 μm. Unfilled, dashed and solid arrowheads indicate Aire+Fezf2-, Aire-Fezf2+, and Aire+Fezf2+ cells, respectively. The histogram shows the density of Aire+Fezf2-, Aire-Fezf2+, and Aire+Fezf2+ cells. (C–E) Flow cytometry profiles, frequencies, and numbers of Aire-Fezf2-, Aire-Fezf2+, and Aire+Fezf2+ cells in total mTECs, mTEClo, and mTEChi (C), of CCL21+ cells in mTEClo (D) and of DCKL1+ cells in Aire- mTEClo (E) from WT, ΔCD4, and mTECΔMHCII mice. II Abs: secondary antibodies. Data are representative of 2–3 independent experiments (n = 2–5 mice per group and experiment). Error bars show mean ± SEM, *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 using unpaired Student’s t-test for (B) and two-tailed Mann–Whitney test for (A) and (C-E).
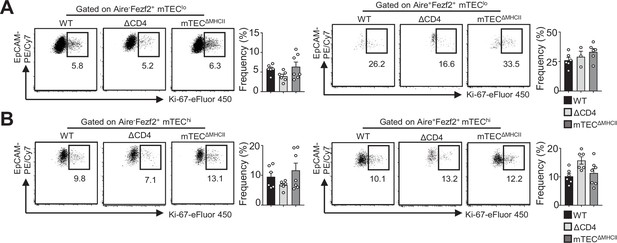
Normal proliferation of Aire-Fezf2+ and Aire +Fezf2+ medullary thymic epithelial cell (mTECs) in ΔCD4 and mTECΔMHCII mice.
(A, B) Flow cytometry profiles and frequencies of Ki-67+ proliferating Aire-Fezf2+ and Aire+Fezf2+ cells in mTEClo (A) and mTEChi (B) from WT, ΔCD4, and mTECΔMHCII mice. Data are representative of two independent experiments (n = 3–4 mice per group and experiment). Error bars show mean ± SEM.
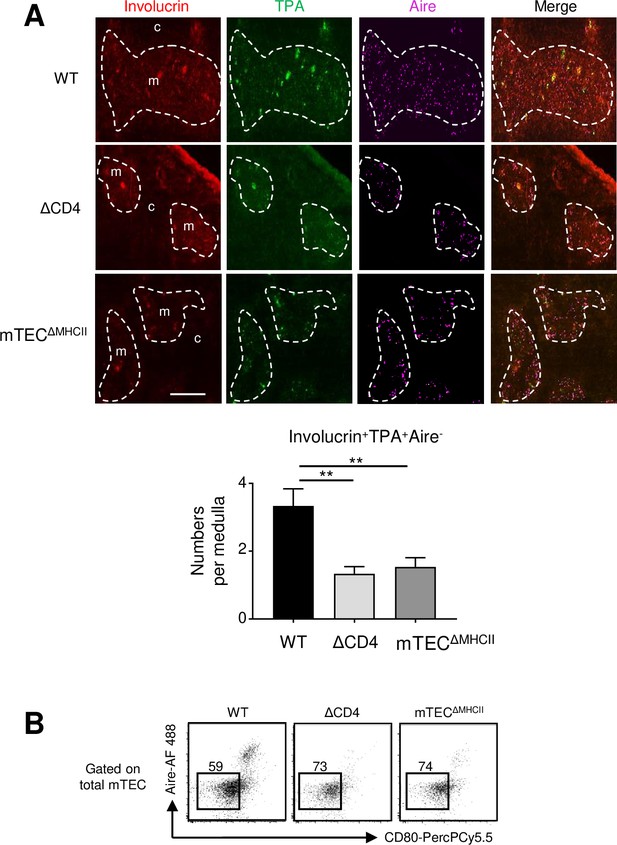
Reduced post-Aire medullary thymic epithelial cells (mTECs) in ΔCD4 and mTECΔMHCII mice.
(A) Representative thymic sections stained with antibodies against involucrin (red), TPA (green), and Aire (magenta). c and m denote the cortex and the medulla, respectively. The graph shows the number of involucrin+TPA+Aire- cells per medulla. 15 medullas derived from two WT, two ΔCD4, and two mTECΔMHCII mice were quantified, respectively. Scale bar: 200 μm. Error bars show mean ± SEM, **p<0.01 using unpaired Student’s t-test. (B) Gating strategy used to identify Aire- mTEClo cells in WT, ΔCD4, and mTECΔMHCII mice.
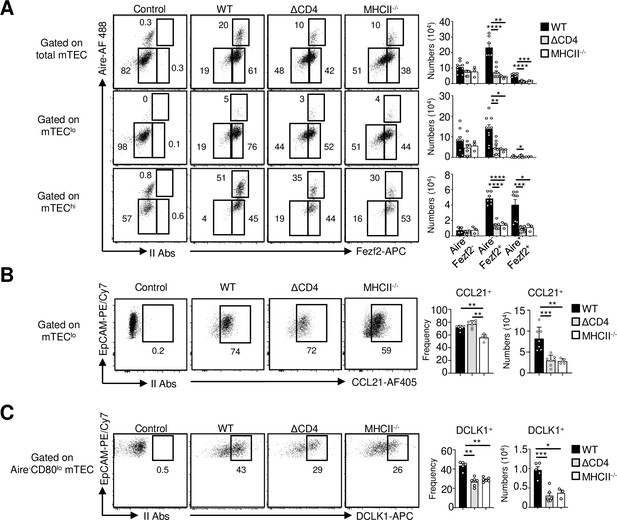
Analysis of medullary thymic epithelial cell (mTEC) subsets in MHCII-/- mice.
(A) Flow cytometry profiles and numbers of Aire-Fezf2-, Aire-Fezf2+, and Aire+Fezf2+ cells in total mTECs, mTEClo, and mTEChi from WT, ΔCD4, and MHCII-/- mice. II Abs: secondary antibodies. Data are representative of two independent experiments (n = 2–4 mice per group). (B) Flow cytometry profiles, frequencies, and numbers of CCL21+ cells in mTEClo from WT, ΔCD4, and MHCII-/- mice. (C) Flow cytometry profiles, frequencies, and numbers of DCKL1+ cells in Aire- mTEClo. Data are representative of two independent experiments (n = 2–3 mice per group). Error bars show mean ± SEM, *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 using the Mann–Whitney test.
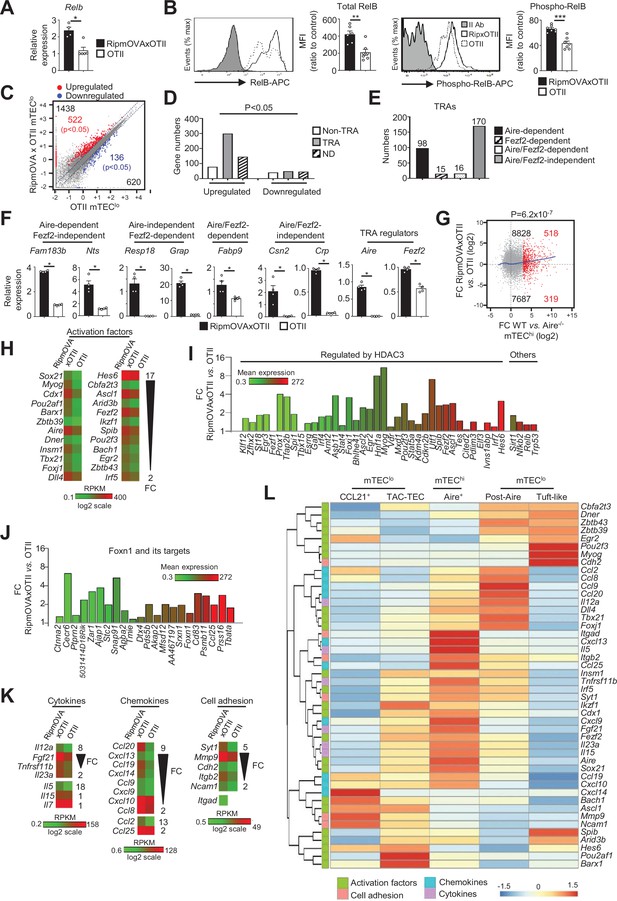
Highly self-reactive CD4+ thymocytes control the transcriptional and functional properties of mTEClo.
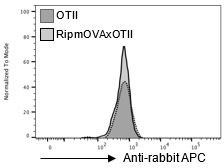
(A) Relb mRNA was measured by qPCR in mTEClo from RipmOVAxOTII-Rag2-/- (n = 4) and OTII-Rag2-/- (n = 5) mice. (B) Total and phospho-RelB (Ser552) were analyzed by flow cytometry in mTEClo from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. Data are representative of two independent experiments (n = 3–4 mice per group and experiment). (C) Scatter plot of gene expression levels (fragments per kilobase of transcript per million mapped reads [FPKM]) in mTEClo from RipmOVAxOTII-Rag2-/- versus OTII-Rag2-/- mice. Genes with fold difference ≥2 and p-adj<0.05 were considered as upregulated or downregulated genes (red and blue dots, respectively). RNA-seq was performed on two independent biological replicates with mTEClo derived from 5 to 8 mice. (D) Numbers of tissue-restricted self-antigens (TRAs) and non-TRAs in genes up- and downregulated in mTEClo from RipmOVAxOTII-Rag2-/- versus OTII-Rag2-/- mice. ND, not determined. (E) Numbers of induced Aire-dependent, Fezf2-dependent, Aire/Fezf2-dependent, and Aire/Fezf2-independent TRAs. (F) Aire-dependent (Fam183b, Nts), Fezf2-dependent (Resp18, Grap), Aire/Fezf2-dependent (Fabp9), Aire/Fezf2-independent (Csn2, Crp) TRAs, Aire and Fezf2 mRNAs were measured by qPCR in mTEClo from RipmOVAxOTII-Rag2-/- (n = 4) and OTII-Rag2-/- (n = 4) mice. (G) Scatter plot of gene expression variation in mTEClo from RipmOVAxOTII-Rag2-/- versus OTII-Rag2-/- mice and in mTEChi from WT versus Aire-/- mice. The loess fitted curve is shown in blue and induced Aire-dependent genes (fold change [FC] > 5) in red. (H) Heatmap of significantly upregulated activation factors in mTEClo from RipmOVAxOTII-Rag2-/- compared to OTII-Rag2-/- mice. (I, J) Expression FC in HDAC3-induced transcriptional regulators and other transcription factors (I) and in Foxn1 targets (J) in mTEClo from RipmOVAxOTII-Rag2-/- versus OTII-Rag2-/- mice. The color code represents gene expression level. (K) Heatmap of significantly upregulated cytokines, chemokines, and cell adhesion molecules in mTEClo from RipmOVAxOTII-Rag2-/- mice. (L) Hierarchical clustering and heatmap of mean expression of these activation factors, cell adhesion molecules, chemokines, and cytokines in mTEC subsets identified by scRNA-seq. Error bars show mean ± SEM, *p<0.05, **p<0.01, ***p<0.001 using two-tailed Mann–Whitney test for (A), (B) and (F) and chi-squared test for (D) and (G).
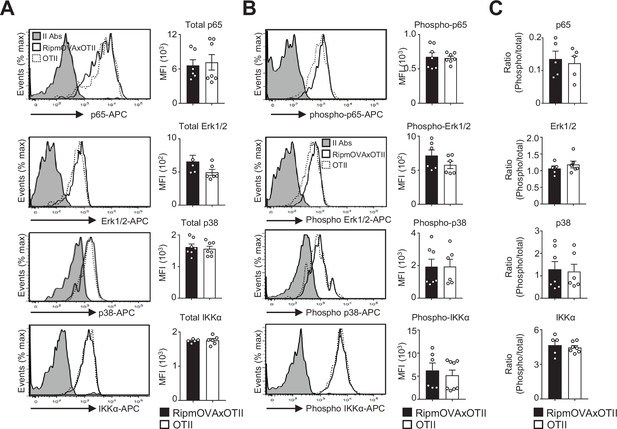
Similar levels of total and phosphorylated p65, Erk1/2, p38, and IKKα proteins in mTEClo from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice.
(A, B) Total p65, Erk1/2, p38, and IKKα (A) and phospho-p65 (Ser536), phospho-Erk1/2 MAPK (Thr202/Tyr204), phospho-p38 MAPK (Thr180/Tyr182), and phospho-IKKα(Ser180)/IKKβ(Ser181) (B) proteins were analyzed by flow cytometry in mTEClo from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. Histograms show the MFI. (C) Histograms represent the ratio of phospho-p65 (Ser536) to p65, phospho-Erk1/2 MAPK (Thr202/Tyr204) to Erk1/2, phospho-p38 MAPK (Thr180/Tyr182) to p38, and phospho-IKKα (Ser180)/IKKβ(Ser181) to IKKα. II Abs: secondary antibodies. Data are representative of two independent experiments (n = 2–4 mice per group and experiment). Error bars show mean ± SEM.
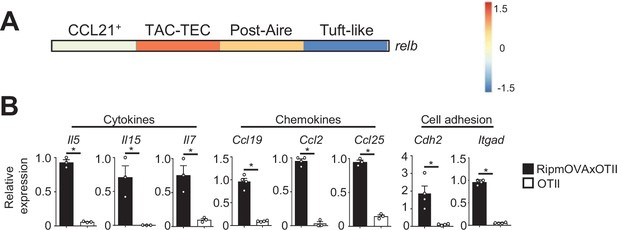
Expression of Relb, cytokines, chemokines, and cell adhesion molecules that was altered in mTEClo from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice.
(A) Mean expression of Relb in mTEClo subsets identified by scRNA-seq data derived from Wells et al., 2020. (B) Il5, Il15, Il7, Ccl19, Ccl2, Ccl25, Cdh2, and Itgad mRNAs were measured by qPCR in mTEClo from RipmOVAxOTII-Rag2-/- (n = 3–4) and OTII-Rag2-/- (n = 3–4) mice.
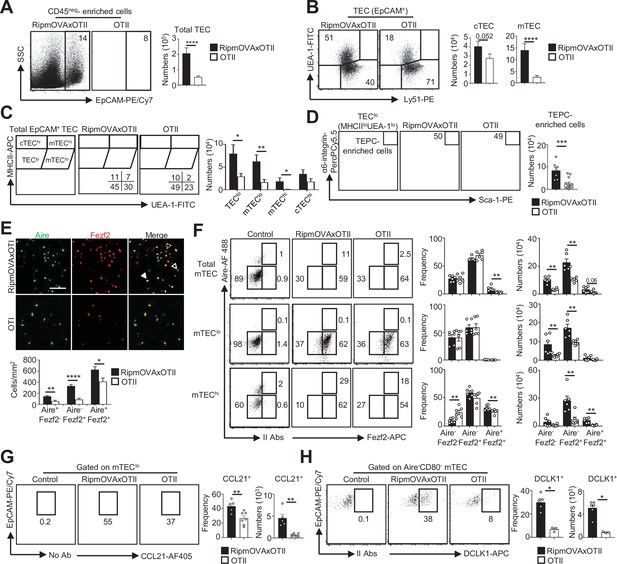
Highly self-reactive CD4+ thymocytes control medullary thymic epithelial cell (mTEC) development from an early progenitor stage.
(A–D) Flow cytometry profiles and numbers of total thymic epithelial cells (TECs) (EpCAM+) (A), cortical thymic epithelial cell (cTECs) (UEA-1-Ly51hi), mTECs (UEA-1+Ly51lo) (B), TEClo (MHCIIloUEA-1lo), cTEChi (MHCIIhiUEA-1lo), mTEClo (MHCIIloUEA-1hi), and mTEChi (MHCIIhiUEA-1hi) (C), α6-integrinhiSca-1hi TEPC-enriched cells in TEClo (D) in CD45neg-enriched cells from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. Data are representative of four experiments (n = 3 mice per group and experiment). (E) Confocal images of thymic sections from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice stained for Aire (green) and Fezf2 (red). 11 and 22 sections derived from two RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice were quantified, respectively. Scale bar, 50 μm. Unfilled, dashed and solid arrowheads indicate Aire+Fezf2lo, Aire-Fezf2+, and Aire+Fezf2+ cells, respectively. The histogram shows the density of Aire+Fezf2lo, Aire-Fezf2+, and Aire+Fezf2+ cells. (F–H) Flow cytometry profiles, frequencies, and numbers of Aire-Fezf2-, Aire-Fezf2+, and Aire+Fezf2+ cells in total mTECs, mTEClo, and mTEChi (F), of CCL21+ cells in mTEClo (G) and of DCKL1+ cells in Aire- mTECs (H) from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. II Abs: secondary antibodies. Data are representative of two independent experiments (n = 3–4 mice per group and experiment). Error bars show mean ± SEM, *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 using unpaired Student’s t-test for (A–E) and two-tailed Mann–Whitney test for (F–H).
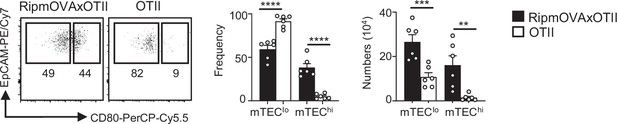
mTEClo and mTEChi cells are increased in RipmOVAxOTII-Rag2-/- compared to OTII-Rag2-/- mice.
Flow cytometry profiles, frequencies, and numbers of mTEClo and mTEChi from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. Data are representative of two independent experiments (n = 3 mice per group and experiment). Error bars show mean ± SEM. **p<0.01, ***p<0.001, ****p<0.0001 using the Mann–Whitney test.
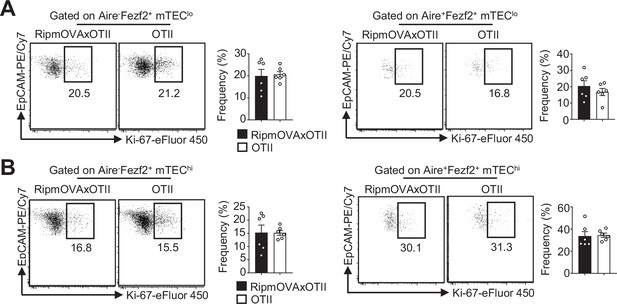
The proliferation of Aire-Fezf2+ and Aire+Fezf2+ medullary thymic epithelial cells (mTECs) is similar in RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice.
(A, B) Flow cytometry profiles and frequencies of proliferating Ki-67+ Aire-Fezf2+ and Aire+Fezf2+ in mTEClo (A) and mTEChi (B) from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. Data are representative of two independent experiments (n = 3 mice per group and experiment). Error bars show mean ± SEM.
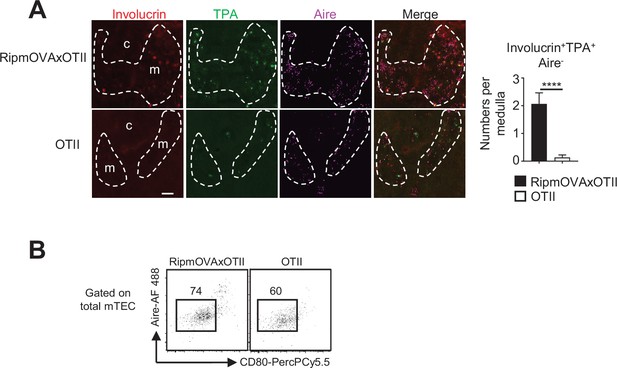
Post-Aire medullary thymic epithelial cells (mTECs) are increased in RipmOVAxOTII-Rag2-/- compared to OTII-Rag2-/- mice.
(A) Representative thymic sections from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice stained with antibodies against involucrin (red), TPA (green), and Aire (magenta). c and m denote the cortex and the medulla, respectively. The graph shows the number of involucrin+TPA+Aire- cells per medulla. 15 medullas derived from two RipmOVAxOTII-Rag2-/- and two OTII-Rag2-/- mice were quantified, respectively. Scale bar: 200 μm. Error bars show mean ± SEM, ****p<0.0001 using unpaired Student’s t-test. (B) Gating strategy used to identify Aire- mTEClo cells in RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice.
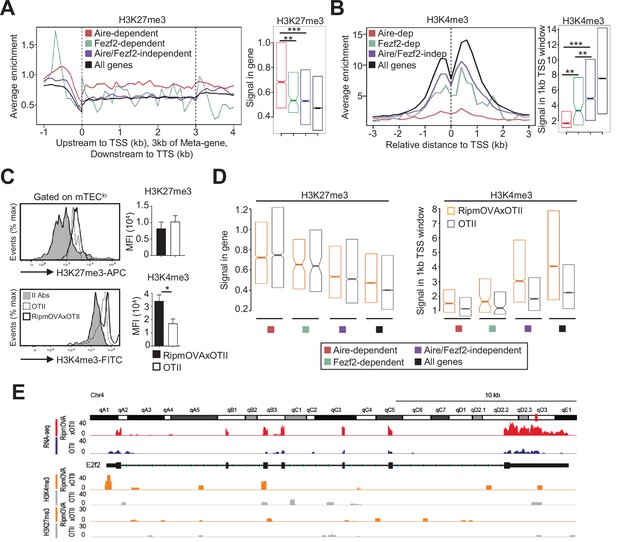
H3K27me3 and H3K4me3 landscape in tissue-restricted self-antigen (TRA) genes of mTEClo from WT, RipmOVAxOTII-Rag2-/-, and OTII-Rag2-/- mice.
(A, B) Metagene profiles of the average normalized enrichment of H3K27me3 (A) and H3K4me3 (B) against input for Aire-dependent, Fezf2-dependent, and Aire/Fezf2-independent TRAs as well as for all genes of WT mTEClo. Boxplots represent the median enrichment, the 95% CI of the median (notches), and the 75th and 25th percentiles of H3K27me3 and H3K4me3. (C) H3K27me3 and H3K4me3 levels were analyzed by flow cytometry in mTEClo from RipmOVAxOTII-Rag2-/- (n = 4) and OTII-Rag2-/- (n = 5) mice. Histograms show the MFI. (D) Boxplots represent the median enrichment of H3K27me3 and H3K4me3 of Aire-dependent, Fezf2-dependent, and Aire/Fezf2-independent TRAs and in all genes of mTEClo from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. (E) Expression (RNA-seq) and H3K27me3 and H3K4me3 chromatin state (ChIP-seq) of the Aire/Fezf2-independent TRA, E2f2, in mTEClo from RipmOVAxOTII-Rag2-/- and OTII-Rag2-/- mice. *p<0.05, **p<10–3, ***p<10–7, using the Mann–Whitney test for (A) and (B) and unpaired Student’s t-test for (C).
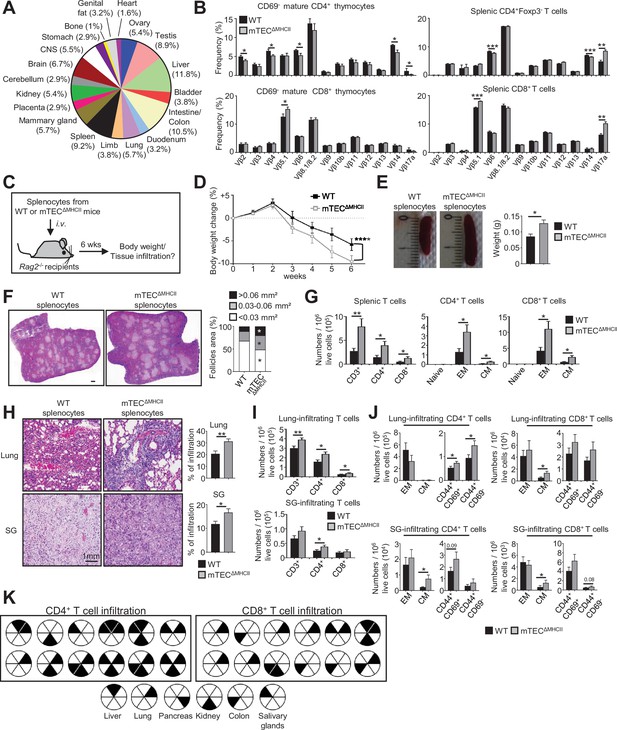
The adoptive transfer of T cells from mTECΔMHCII mice into Rag2-/- recipients induces autoimmunity.
(A) Tissue-restricted self-antigens (TRAs) underexpressed in mTEClo from mTECΔMHCII mice were assigned to their peripheral expression. (B) TCRVβ usage by CD69- mature CD4+ and CD8+ thymocytes (left panel) and CD4+Foxp3- and CD8+ splenic T cells (right panel) from WT and mTECΔMHCII mice. (C) Body weight of Rag2-/- recipients transferred with splenic T cells from WT or mTECΔMHCII was monitored during 6 weeks, and tissue infiltration was examined. (D) Weight loss relative to the initial weight. (E, F) Representative spleen pictures and their weights (E) and hematoxylin/eosin counterstained splenic sections (F). Scale bar, 1 mm. The histogram shows follicle areas. (G) Numbers of splenic CD3+, CD4+, and CD8+ T cells and of naive (CD44loCD62Lhi), effector memory (EM; CD44hiCD62Llo) and central memory (CM; CD44hiCD62Lhi) phenotype. (H) Lung and salivary gland (SG) immune infiltrates detected by hematoxylin/eosin counterstaining. Scale bar, 1 mm. (I, J) Numbers of T cells (I) and of naive, effector and central memory phenotype as well as CD44+CD69+ and CD44+CD69- T cells (J) in lungs and SG. (K) Schematic of T-cell infiltrates in mice transferred with mTECΔMHCII T cells relative to those transferred with WT T cells. Each circle and black triangles represent an individual mouse and T-cell infiltration in a specific tissue, respectively. Data are representative of two independent experiments (n = 5–7 mice per group and experiment). Error bars show mean ± SEM, ****p<0.0001 using two-way ANOVA for (D) and unpaired Student’s t-test for (B) and (E-J). *p<0.05, **p<0.01, ***p<0.001.
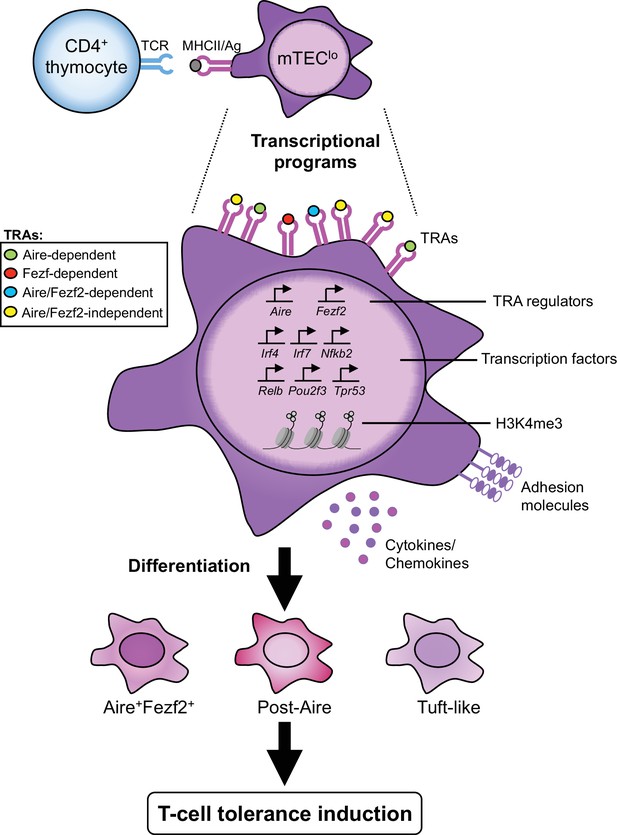
CD4+ thymocytes through MHCII/TCR-mediated interactions control transcriptional programs of mTEClo that drive their differentiation and function.
Antigen-specific interactions between CD4+ thymocytes and mTEClo lead to the upregulation of tissue-restricted self-antigen (TRA) regulators, TRAs, key medullary thymic epithelial cell (mTEC)-specific transcription factors, adhesion molecules, cytokines, and chemokines, which correlates with an increased level of the active H3K4me3 histone mark. These interactions enhance the transcriptional activity of TAC-TECs accompanying the transition to Aire+Fezf2+, as well as post-Aire cells and tuft-like mTECs. They are thus essential to generate a self-tolerant T-cell repertoire and prevent the development of autoimmunity.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Mus musculus) | C57BL/6J background | Charles River | RRID:IMSR_JAX:000664 | |
| Genetic reagent (M. musculus) | Ciitatm2Wrth/Ciitatm2Wrth | LeibundGut-Landmann et al., 2004 | RRID:MGI:3052466 | C57BL/6 background, ΔCD4 mice |
| Genetic reagent (M. musculus) | H2dlAb1-Ea/H2dlAb1-Ea | Madsen et al., 1999 | RRID:MGI:4436873 | C57BL/6 background, MHCII-/- mice |
| Genetic reagent (M. musculus) | K14xCiitaIII+IV-/- | Irla et al., 2008 | C57BL/6 background, mTECΔMHCII mice | |
| Genetic reagent (M. musculus) | Tg(TcraTcrb)425Cbn | Barnden et al., 1998 | RRID:MGI:3762632 | C57BL/6 background, OTII mice |
| Genetic reagent (M. musculus) | Tg(Ins2-TFRC/OVA)296Wehi | Kurts et al., 1996 | RRID:MGI:3623748 | C57BL/6 background, Rip-mOVA mice |
| Genetic reagent (M. musculus) | Rag2tm1Fwa/Rag2tm1Fwa | Shinkai et al., 1992 | RRID:MGI:2174910 | C57BL/6 background, Rag2-/- mice |
| Antibody | Anti-IKKα (rabbit polyclonal) | Cell Signaling Technology | Cat# 2682; RRID:AB_331626 | FACS (1:500) |
| Antibody | Anti-phospho IKKα (Ser180)/IKKβ(Ser181) (rabbit polyclonal) | Cell Signaling Technology | Cat# 2681S; RRID:AB_331624 | FACS (1:500) |
| Antibody | Anti-p38 MAPK (rabbit polyclonal) | Cell Signaling Technology | Cat# 9212; RRID:AB_330713 | FACS (1:500) |
| Antibody | Anti-phospho p38 MAPK (Thr180/Tyr182) (rabbit polyclonal) | Cell Signaling Technology | Cat# 9211S; RRID:AB_331641 | FACS (1:500) |
| Antibody | Anti-Erk1/2(rabbit polyclonal) | Cell Signaling Technology | Cat# 9102; RRID:AB_330744 | FACS (1:500) |
| Antibody | Anti-phospho Erk1/2 (Thr202/Tyr204) (rabbit polyclonal) | Cell Signaling Technology | Cat# 9101S;RRID:AB_331646 | FACS (1:500) |
| Antibody | Anti-NF-κB p65 (clone D14E12, rabbit monoclonal) | Cell Signaling Technology | Cat# 8242S; RRID:AB_10859369 | FACS (1:500) |
| Antibody | Phospho-NF-κB p65 (Ser536) (clone 93H1, rabbit monoclonal) | Cell Signaling Technology | Cat# 3033S; RRID:AB_331284 | FACS (1:3000) |
| Antibody | Anti-RelB (clone C-19, rabbit polyclonal) | Santa Cruz Biotechnology | Cat# sc-226; RRID:AB_632341 | FACS (1:200) |
| Antibody | Anti-phospho RelB (ser552) (clone D41B9, rabbit monoclonal) | Cell Signaling Technology | Cat# 5025S; RRID:AB_10622001 | FACS (1:1000) |
| Antibody | Anti-DCLK1 (clone D2U3L, rabbit monoclonal) | Cell Signaling Technology | Cat# 62257; RRID:AB_2799622 | FACS (1:200) |
| Antibody | Anti-H3K4me3 (rabbit polyclonal) | Abcam | Cat# ab8580; RRID:AB_306649 | FACS(1:1000)ChIP-seq (2 µg:25 µg chromatin) |
| Antibody | Anti-H3K27me3 (clone C36B11, rabbit monoclonal) | Cell Signaling Technology | Cat# 9733; RRID:AB_2616029 | FACS(1:1000)ChIP-seq(1:50) |
| Antibody | PE-Cy7 anti-CD326 (EpCAM) (clone G8.8, rat monoclonal) | eBioscience | Cat# 25-5791-80; RRID:AB_1724047 | FACS(1:3000) |
| Antibody | Alexa Fluor 488 anti-Aire (clone 5H12, rat monoclonal) | eBioscience | Cat# 53-5934-82; RRID:AB_10854132 | FACS, IF(1:200) |
| Antibody | PE anti-Ly51 (clone BP-1, mouse monoclonal) | BD Biosciences | Cat# 553735; RRID:AB_395018 | FACS(1:3000) |
| Antibody | PerCP-Cy5.5 anti-CD80 (clone 16-10A1, Armenian hamster monoclonal) | BioLegend | Cat# 104722; RRID:AB_2291392 | FACS(1:200) |
| Antibody | eFluor 450 anti-Ki-67 (clone SolA15, rat monoclonal) | eBioscience | Cat# 48-5698-82; RRID:AB_11149124 | FACS(1:200) |
| Antibody | Anti-Fezf2 (clone F441, rabbit polyclonal) | IBL Tecan | Cat# JP18997; RRID:AB_2341444 | FACS, IF(1:200) |
| Antibody | Anti-Involucrin (clone Poly19244, rabbit polyclonal) | BioLegend | Cat# 924401; RRID:AB_2565452 | IF(1:100) |
| Antibody | PE anti-Ly-6A/E (Sca-1) (clone D7, rat monoclonal) | BD Biosciences | Cat# 553108; RRID:AB_394629 | FACS(1:600) |
| Antibody | Biotin anti-CD49f (α6-integrin) (clone GoH3, rat monoclonal) | BioLegend | Cat# 313604; RRID:AB_345298 | FACS(1:200) |
| Antibody | Alexa Fluor 647 anti-I-Ab (MHCII) (clone AF6-120.1, mouse monoclonal) | BioLegend | Cat# 116412; RRID:AB_493141 | FACS(1:200) |
| Antibody | Brilliant Violet 421 anti-CD4 (clone RM4-5, rat monoclonal) | BioLegend | Cat# 100544; RRID:AB_11219790 | FACS(1:200) |
| Antibody | PerCP-Cy5.5 anti-CD4 (clone RM4-5, rat monoclonal) | BD Biosciences | Cat# 550954; RRID:AB_393977 | FACS(1:200) |
| Antibody | Pacific Blue anti-CD8α (clone 53-6.7, rat monoclonal) | BD Biosciences | Cat# 558106; RRID:AB_397029 | FACS(1:200) |
| Antibody | PE/Cy7 anti-CD8α (clone 53-6.7, rat monoclonal) | BioLegend | Cat# 100722; RRID:AB_312761 | FACS(1:600) |
| Antibody | Alexa Fluor 488 anti-CD44 (clone IM7, rat monoclonal) | BioLegend | Cat# 103016; RRID:AB_493679 | FACS(1:200) |
| Antibody | PE anti-CD69 (clone H1.2F3, rat monoclonal) | BioLegend | Cat# 104508; RRID:AB_313111 | FACS(1:400) |
| Antibody | PE anti-CD62L (clone MEL-14, rat monoclonal) | BD Biosciences | Cat# 553151; RRID:AB_394666 | FACS(1:300) |
| Antibody | PerCP-Cy5.5 anti-CD3ε (clone 17A2, rat monoclonal) | BD Biosciences | Cat# 560527;RRID:AB_1727463 | FACS(1:200) |
| Antibody | Alexa Fluor 405 anti-CCL21 (clone 59106, rat monoclonal) | R&D Systems | Cat# IC457V | FACS(1:100) |
| Antibody | CD45 MicroBeads, mouse (clone 30F11.1, rat monoclonal) | Miltenyi | Cat# 130052301; RRID:AB_2877061 | |
| Antibody | Cy5 anti-rabbit IgG (goat polyclonal) | Invitrogen | Cat# A10523; RRID:AB_2534032 | FACS(1:500) |
| Antibody | Cyanine 3 anti-rabbit IgG (goat polyclonal) | Invitrogen | Cat# A10520; RRID:AB_2534029 | IF(1:500) |
| Antibody | FITC anti-TCR Vβ2 (clone B20.6, rat monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | FITC anti-TCR Vβ3 (clone KJ25, Armenian hamster monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | FITC anti-TCR Vβ4 (clone KT4, rat monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | FITC anti-TCR Vβ5.1, 5.2 (clone MR9-4, mouse monoclonal) | BD Biosciences | Cat# 553189; RRID:AB_394697 | FACS(1:100) |
| Antibody | FITC anti-TCR Vβ6 (clone RR4-7, rat monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | PE anti-TCR Vβ8.1, 8.2 (clone MR5-2, mouse monoclonal) | BioLegend | Cat# 140103; RRID:AB_10641144 | FACS(1:300) |
| Antibody | FITC anti-TCR Vβ9 (clone MR10-2, mouse monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | PE anti-TCR Vβ10b (clone B21.5, rat monoclonal) | BD Biosciences | Cat# 553285; RRID:AB_394757 | FACS(1:300) |
| Antibody | Biotin anti-TCR Vβ11 (clone RR3-15, rat monoclonal) | BD Biosciences | Cat# 553196; RRID:AB_394702 | FACS(1:300) |
| Antibody | FITC anti-TCR Vβ12 (clone MR11-1, mouse monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | FITC anti-TCR Vβ13 (clone MR12-3, mouse monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | FITC anti-TCR Vβ14 (clone 14-2, rat monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | FITC anti-TCR Vβ17a (clone KJ23, rat monoclonal) | BD Biosciences | Cat# 557004; RRID:AB_647180 | FACS(20 µl per 106 cells) |
| Antibody | CD4+ T cell isolation kit, mouse | Miltenyi Biotec | Cat# 130-104-454 | |
| Peptide, recombinant protein | PerCP-Cy5.5 Streptavidin | BioLegend | Cat# 405214; RRID:AB_2716577 | FACS(1:400) |
| Peptide, recombinant protein | Alexa Fluor 488 Streptavidin | Invitrogen | Cat# S11223 | IF(1:1000) |
| Peptide, recombinant protein | Ovalbumin (323–339) | PolyPeptide | Cat# SC1303 | 5 μM |
| Chemical compound, drug | Liberase TM | Roche | Cat# 05401127001 | 50 μg/ml |
| Chemical compound, drug | DNase I | Roche | Cat# 10104159001 | 100 μg/ml |
| Chemical compound, drug | TRIzol | Thermo Fisher Scientific | Cat# 15596018 | |
| Software, algorithm | GraphPad Prism | GraphPad Software | RRID:SCR_002798 | |
| Software, algorithm | FlowJo | FlowJo | https://www.flowjo.com/RRID:SCR_008520 | |
| Software, algorithm | Fiji/ImageJ software | Fiji-ImageJ | https://imagej.nih.gov/ij/RRID:SCR_003070 | |
| Software, algorithm | 7500 Real-Time PCR Software | Thermo Fisher | https://www.thermofisher.com/us/en/home/technical-resources/software-downloads/applied-biosystems-7500-real-time-pcr-system.htmlRRID:SCR_014596 | |
| Software, algorithm | Pheatmap 0.2 | https://github.com/raivokolde/pheatmap (Kolde, 2018) | RRID:SCR_016418 | |
| Software, algorithm | Seurat | Hao et al., 2021 | RRID:SCR_016341 | |
| Sequence-based reagent | Actin-FW | Sigma-Aldrich | PCR primers | CAGAAGGAGATTACTGCTCTGGCT |
| Sequence-based reagent | Actin-RV | Sigma-Aldrich | PCR primers | GGAGCCACCGATCCACACA |
| Sequence-based reagent | Aire-FW | Sigma-Aldrich | PCR primers | GCATAGCATCCTGGACGGCTTCC |
| Sequence-based reagent | Aire-RV | Sigma-Aldrich | PCR primers | CTGGGCTGGAGACGCTCTTTGAG |
| Sequence-based reagent | Ccl19-FW | Sigma-Aldrich | PCR primers | GCTAATGATGCGGAAGACTG |
| Sequence-based reagent | Ccl19-RV | Sigma-Aldrich | PCR primers | ACTCACATCGACTCTCTAGG |
| Sequence-based reagent | Ccl2-FW | Sigma-Aldrich | PCR primers | TGGAGCATCCACGTGTTG |
| Sequence-based reagent | Ccl2-RV | Sigma-Aldrich | PCR primers | ACTCATTGGGATCATCTTGCT |
| Sequence-based reagent | Ccl22-FW | Sigma-Aldrich | PCR primers | CTGATGCAGGTCCCTATGGT |
| Sequence-based reagent | Ccl22-RV | Sigma-Aldrich | PCR primers | GGAGTAGCTTCTTCACCCAG |
| Sequence-based reagent | Ccl25-FW | Sigma-Aldrich | PCR primers | GCCTGGTTGCCTGTTTTGTT |
| Sequence-based reagent | Ccl25-RV | Sigma-Aldrich | PCR primers | ACCCAGGCAGCAGTCTTCAA |
| Sequence-based reagent | Cdh2-FW | Sigma-Aldrich | PCR primers | AGCGCAGTCTTACCGAAGG |
| Sequence-based reagent | Cdh2-RV | Sigma-Aldrich | PCR primers | TCGCTGCTTTCATACTGAACTTT |
| Sequence-based reagent | Coch-FW | Sigma-Aldrich | PCR primers | GTGCAGCAAAACCTGCTACAA |
| Sequence-based reagent | Coch -RV | Sigma-Aldrich | PCR primers | AGCTAGGACGTTCTCTTTGGT |
| Sequence-based reagent | Crabp1-FW | Sigma-Aldrich | PCR primers | CAGCAGCGAGAATTTCGACGA |
| Sequence-based reagent | Crabp1-RV | Sigma-Aldrich | PCR primers | CGCACAGTAGTGGATGTCTTGA |
| Sequence-based reagent | Crp-FW | Sigma-Aldrich | PCR primers | CATAGCCATGGAGAAGCTAC |
| Sequence-based reagent | Crp-RV | Sigma-Aldrich | PCR primers | CAGTGGCTTCTTTGACTCTG |
| Sequence-based reagent | Csn2-FW | Sigma-Aldrich | PCR primers | CTCCACTAAAGGACTTGACAG |
| Sequence-based reagent | Csn2-RV | Sigma-Aldrich | PCR primers | ACCTTCTGAAGTTTCTGCTC |
| Sequence-based reagent | Fabp9-FW | Sigma-Aldrich | PCR primers | CACTGCAGACAACCGAAAAG |
| Sequence-based reagent | Fabp9-RV | Sigma-Aldrich | PCR primers | TCTGTTTGCCAAGCCATTTT |
| Sequence-based reagent | Fam183b-FW | Sigma-Aldrich | PCR primers | CGTGTGGGGCAGATGAAGAAT |
| Sequence-based reagent | Fam183b-RV | Sigma-Aldrich | PCR primers | GGTGAATGAGGTTCAGGAACTTG |
| Sequence-based reagent | Fcer2a-FW | Sigma-Aldrich | PCR primers | CCAGGAGGATCTAAGGAACGC |
| Sequence-based reagent | Fcer2a-RV | Sigma-Aldrich | PCR primers | TCGTCTTGGAGTCTGTTCAGG |
| Sequence-based reagent | Fezf2-FW | Sigma-Aldrich | PCR primers | CAGCACTCTCTGCAGACACAA |
| Sequence-based reagent | Fezf2-RV | Sigma-Aldrich | PCR primers | TGCCGCACTGGTTACACTTA |
| Sequence-based reagent | Grap-FW | Sigma-Aldrich | PCR primers | GATCAGGGAGAGTGAGAGTTCC |
| Sequence-based reagent | Grap-RV | Sigma-Aldrich | PCR primers | CAGCTCGTTGAGGGAGTTGA |
| Sequence-based reagent | Icam2-FW | Sigma-Aldrich | PCR primers | ATCAACTGCAGCACCAACTG |
| Sequence-based reagent | Icam2-RV | Sigma-Aldrich | PCR primers | ACTTGAGCTGGAGGCTGGTA |
| Sequence-based reagent | Il15-FW | Sigma-Aldrich | PCR primers | AGCAGATAACCAGCCTACAGGA |
| Sequence-based reagent | Il15-RV | Sigma-Aldrich | PCR primers | TGTTGAAGATGAGCTGGCTATGG |
| Sequence-based reagent | Il21-FW | Sigma-Aldrich | PCR primers | CGCCTCCTGATTAGACTTCG |
| Sequence-based reagent | Il21-RV | Sigma-Aldrich | PCR primers | TGGAGCTGATAGAAGTTCAGGA |
| Sequence-based reagent | Il5-FW | Sigma-Aldrich | PCR primers | CCGCCAAAAAGAGAAGTGTGGCGA |
| Sequence-based reagent | Il5-RV | Sigma-Aldrich | PCR primers | GCCTCAGCCTTCCATTGCCCA |
| Sequence-based reagent | Il7-FW | Sigma-Aldrich | PCR primers | GGGTCCTGGGAGTGATTATGG |
| Sequence-based reagent | Il7-RV | Sigma-Aldrich | PCR primers | CGGGAGGTGGGTGTAGTCAT |
| Sequence-based reagent | Itgad-FW | Sigma-Aldrich | PCR primers | CGAAAGGGTTCAGACTTTGC |
| Sequence-based reagent | Itgad-RV | Sigma-Aldrich | PCR primers | ACACCTCCACGGATAGAAGTC |
| Sequence-based reagent | Itgb6-FW | Sigma-Aldrich | PCR primers | GCTGGTCTGCCTGTTTCTGC |
| Sequence-based reagent | Itgb6-RV | Sigma-Aldrich | PCR primers | TGAGCAGCTTTCTGCACCAC |
| Sequence-based reagent | Kcnj5-FW | Sigma-Aldrich | PCR primers | AAAACCTTAGCGGCTTTGTATCT |
| Sequence-based reagent | Kcnj5-RV | Sigma-Aldrich | PCR primers | AAGGCATTAACAATCGAGCCC |
| Sequence-based reagent | Krt1-FW | Sigma-Aldrich | PCR primers | TGGGAGATTTTCAGGAGGAGG |
| Sequence-based reagent | Krt1-RV | Sigma-Aldrich | PCR primers | GCCACACTCTTGGAGATGCTC |
| Sequence-based reagent | Meig1-FW | Sigma-Aldrich | PCR primers | CTTCAGCGGAGGGACAATAC |
| Sequence-based reagent | Meig1-RV | Sigma-Aldrich | PCR primers | CAAGGTTTCAAGGTGGGTGT |
| Sequence-based reagent | Nov-FW | Sigma-Aldrich | PCR primers | AGACCCCAACAACCAGACTG |
| Sequence-based reagent | Nov-RV | Sigma-Aldrich | PCR primers | CGGTAAATGACCCCATCGAAC |
| Sequence-based reagent | Nts-FW | Sigma-Aldrich | PCR primers | GCAAGTCCTCCGTCTTGGAAA |
| Sequence-based reagent | Nts-RV | Sigma-Aldrich | PCR primers | TGCCAACAAGGTCGTCATCAT |
| Sequence-based reagent | Reig1-FW | Sigma-Aldrich | PCR primers | ATGGCTAGGAACGCCTACTTC |
| Sequence-based reagent | Reig1-RV | Sigma-Aldrich | PCR primers | CCCAAGTTAAACGGTCTTCAGT |
| Sequence-based reagent | Resp18-FW | Sigma-Aldrich | PCR primers | CCAGCCAAGATGCAGAGTTCGTTAAAG |
| Sequence-based reagent | Resp18-RV | Sigma-Aldrich | PCR primers | TCAGTCAGCAACAAGGTTGAGGCCCAC |
| Sequence-based reagent | Rsph1-FW | Sigma-Aldrich | PCR primers | ACGGGGACACATATGAAGGA |
| Sequence-based reagent | Rsph1-RV | Sigma-Aldrich | PCR primers | GGCCGTGCTTTTTATTTTTG |
| Sequence-based reagent | Spon2-FW | Sigma-Aldrich | PCR primers | ATGGAAAACGTGAGTCTTGCC |
| Sequence-based reagent | Spon2-RV | Sigma-Aldrich | PCR primers | TGATGCTGTATCTAGCCAGAGG |
| Sequence-based reagent | Sult1c2-FW | Sigma-Aldrich | PCR primers | ATGGCCTTGACCCCAGAAC |
| Sequence-based reagent | Sult1c2-RV | Sigma-Aldrich | PCR primers | TCGAAGGTCTGAATCTGCCTC |
| Sequence-based reagent | Upk3b-FW | Sigma-Aldrich | PCR primers | CATCTGGCTAGTGGTGGCTTT |
| Sequence-based reagent | Upk3b-RV | Sigma-Aldrich | PCR primers | GGTAATGTCATATAGTGGCCGTC |
| Other | Biotinylated Lotus Tetragonolobus Lectin (LTL) | Vector Laboratories | Cat# B-1325; RRID:AB_2336558 | IF(1:500) |
| Other | FITC Ulex Europaeus Agglutinin I (UEA I) | Vector Laboratories | Cat# FL-1061; RRID:AB_2336767 | FACS(1:600) |
| Other | SuperScript II Reverse Transcriptase | Thermo Fisher | Cat# 18064022 | |
| Other | SYBR Premix Ex Taq master mix | Takara | Cat# RR390A | |
| Other | miRNeasy Micro Kit | QIAGEN | Cat# 217084 | |
| Other | TruSeq ChIP Library Preparation Kit | Illumina | Cat# IP-202-2012 |
Additional files
-
Supplementary file 1
List of tissue-restricted self-antigens (TRAs) differentially expressed in mTEClo from WT and ΔCD4 mice.
- https://cdn.elifesciences.org/articles/69982/elife-69982-supp1-v1.pdf
-
Supplementary file 2
List of tissue-restricted self-antigens (TRAs) differentially expressed in mTEClo from WT and mTECΔMHCII mice.
- https://cdn.elifesciences.org/articles/69982/elife-69982-supp2-v1.pdf
-
Supplementary file 3
List of tissue-restricted self-antigens (TRAs) differentially expressed in mTEClo from OTII-Rag2-/- and RipmOVAxOTII-Rag2-/- mice.
- https://cdn.elifesciences.org/articles/69982/elife-69982-supp3-v1.pdf
-
Supplementary file 4
Main target organs and fold change associated with the expression of tissue-restricted self-antigens (TRAs) differentially expressed in mTEClo from WT and mTECΔMHCII mice.
- https://cdn.elifesciences.org/articles/69982/elife-69982-supp4-v1.pdf
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/69982/elife-69982-transrepform1-v1.pdf