A light-gated transcriptional recorder for detecting cell-cell contacts
Figures
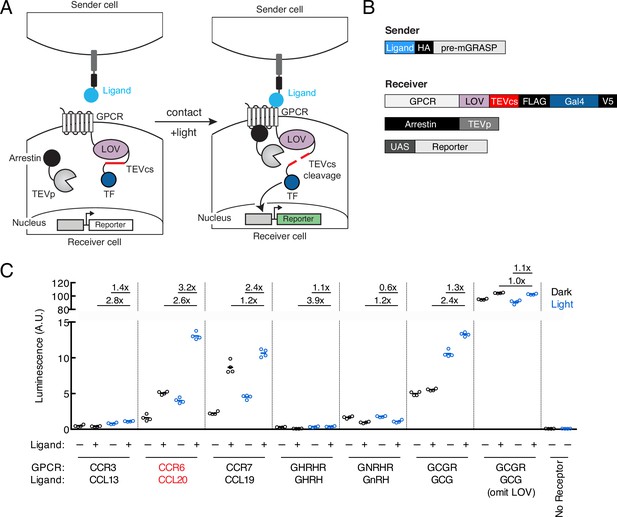
Design of TRACC (Transcriptional Readout Activated by Cell-cell Contacts).
(A) Schematic of TRACC. A ligand is presented on sender cells; a GPCR specifically activated by the selected ligand is expressed in receiver cells. The GPCR is fused to the light-oxygen-voltage-sensing (LOV) domain, TEV protease cleavage site (TEVcs), and transcription factor (TF). Upon both cell-cell contact and exposure to blue light, the GPCR is activated and recruits arrestin fused to TEV protease (TEVp); blue light uncages the LOV domain, allowing cleavage of the TEVcs and subsequent release of the TF, which translocates to the nucleus and drives expression of a reporter of interest. (B) Constructs used in TRACC. The sender construct is comprised of a peptide ligand fused to pre-mGRASP (Kim et al., 2011) and the HA epitope tag. Receiver constructs include the corresponding GPCR fused to the LOV domain, TEVcs, and TF (Gal4), arrestin fused to TEVp, and a reporter construct. (C) Luciferase assay to screen a panel of GPCR-ligand pairs in cis. HEK293T cells were transfected with both sender and receiver constructs corresponding to each GPCR-ligand pair indicated, using the reporter UAS-luciferase. Approximately 8 hr after 10 min blue light exposure, the UAS-luciferase luminescence was recorded using a plate reader (n = 4 replicates per condition). The CCR6-CCL20 pair (red) showed the highest ±light and ±ligand signal ratios of 2.6-fold and 3.2-fold, respectively. A receiver construct using the glucagon receptor (GCGR), but omitting the LOV domain, analogous to that of previously published trans-Tango (Talay et al., 2017), was included as a control.
-
Figure 1—source data 1
Primary data for luminescence graphs in Figure 1C.
- https://cdn.elifesciences.org/articles/70881/elife-70881-fig1-data1-v1.xlsx
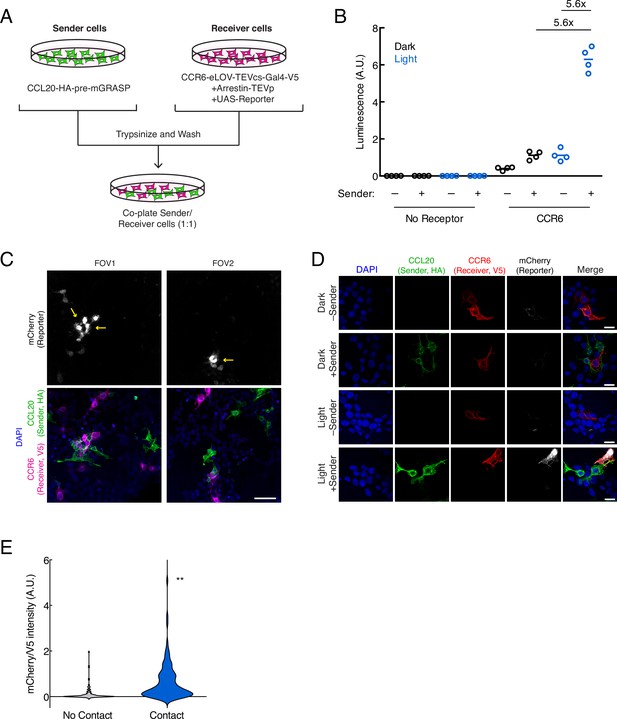
Using TRACC (Transcriptional Readout Activated by Cell-cell Contacts) to detect cell-cell contacts in HEK293T culture.
(A) Experimental design for co-plating sender and receiver cells. (B) Luciferase assay using the CCR6-CCL20 GPCR-ligand pair in trans. HEK293T cells transfected with the CCL20 sender construct were co-plated with HEK293T cells transfected with receiver constructs and UAS-luciferase. Approximately 8 hr after 10 min blue light exposure, the UAS-luciferase luminescence was recorded (n = 4 replicates per condition). (C) Confocal fluorescence imaging of sender cells co-plated with receiver cells, using UAS-mCherry reporter. Approximately 8 hr after 10 min blue light exposure, cells were fixed and immunostained. mCherry activation occurs in receiver cells (V5-positive) that contact sender cells (HA-positive). Yellow arrowheads denote examples in which receiver cells are in contact with sender cells. Scale bar, 60 μm. (D) Confocal fluorescence imaging of sender cells co-plated with receiver cells, using UAS-mCherry at higher magnification. Cells were treated as in (C). Scale bars, 20 μm. (E) Quantification of mCherry/V5 intensity ratios for all V5-positive cells in the +light condition. The mCherry/V5 ratio was significantly higher in V5-positive cells that were in contact with HA-positive sender cells. (no contact, n = 108 cells; contact, n = 106 cells; two-tailed t-test, **p < 0.005).
-
Figure 2—source data 1
Primary data for luminescence and cell count graphs in Figure 2.
- https://cdn.elifesciences.org/articles/70881/elife-70881-fig2-data1-v1.xlsx
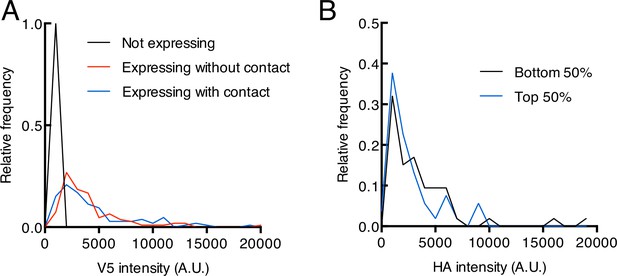
Additional quantification of HEK293T imaging in trans.
(A) Distribution of V5 intensity in V5-positive cells in Figure 2E. The distributions of sender-contacting cells and non-contacting cells are similar, suggesting that mCherry activation is sender-dependent and not a result of differential V5 expression (not expressing, n = 50 cells, expressing without contact, n = 108 cells, expressing with contact, n = 106 cells). (B) Distribution of HA (sender) intensity within the same ROI (region of interest) for each V5-positive cell in contact with a sender cell. Cells were ranked based on their mCherry/V5 intensity ratios and split into groups corresponding to the top and bottom 50 percentiles. Distributions of HA intensity are similar between the two groups, suggesting factors other than differential HA (sender) expression may explain why ~ 19.8% of contacting cells have low mCherry (reporter) expression.
-
Figure 2—figure supplement 1—source data 1
Primary data for graphs in Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/70881/elife-70881-fig2-figsupp1-data1-v1.xlsx
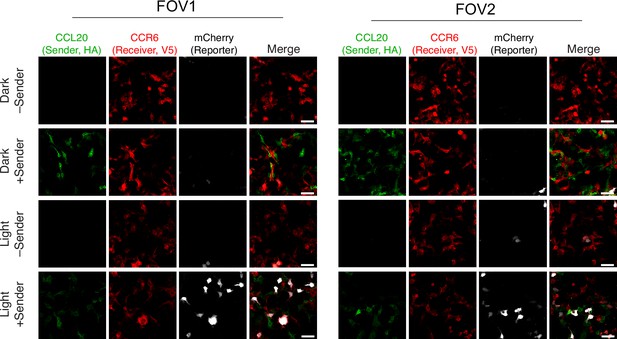
Evaluation of TRACC (Transcriptional Readout Activated by Cell-cell Contacts) in HEK293T cells using lentiviral transduction.
Confocal fluorescence imaging of sender cells co-plated with receiver cells using UAS-mCherry. TRACC components were introduced by lentivirus transduction. Approximately 8 hr after 10 min blue light exposure, cells were fixed and immunostained. Scale bars, 30 μm. FOV, field of view.
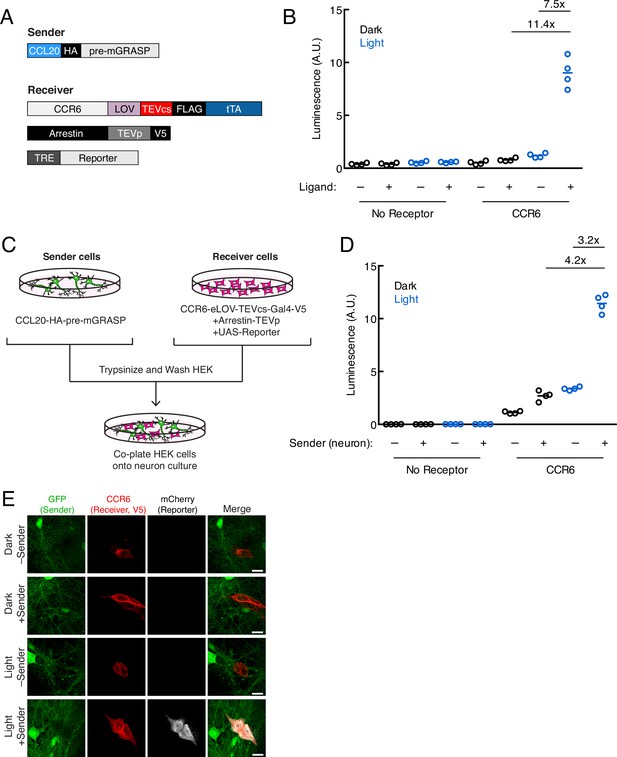
Using TRACC (Transcriptional Readout Activated by Cell-cell Contacts) to detect contacts in neuron culture and HEK293T-neuron co-culture.
(A) Constructs used in TRACC in neuron culture. CCR6 is the GPCR and CCL20 is its activating peptide ligand. For expression in neurons, the transcription factor (TF) is changed from Gal4 to tTA and the reporter gene is driven by TRE rather than a UAS promoter. (B) Luciferase assay using TRACC constructs expressed in cis in neuron culture. Primary rat cortical neurons were infected with AAV1/2 viruses encoding both sender and receiver constructs, including the reporter TRE-luciferase, on DIV5 and light-stimulated on DIV10. Approximately 24 hr after 10 min blue light exposure, the TRE-luciferase luminescence was recorded using a plate reader (n = 4 replicates per condition). (C) Experimental design for co-plating sender neurons and receiver HEK293T cells. (D) Luciferase assay using sender neurons co-cultured with receiver HEK293T cells. Primary rat cortical neurons were infected with AAV1/2 viruses encoding the sender construct on DIV5. HEK293T cells expressing receiver constructs and UAS-luciferase were co-plated onto sender neurons on DIV9, and the resulting co-culture was light-stimulated on DIV10. Approximately 8 hr after 10 min blue light exposure, the UAS-luciferase luminescence was recorded (n = 4 replicates per condition). (E) Confocal fluorescence imaging of receiver HEK293T cells co-cultured with sender neurons, using UAS-mCherry. GFP driven by the synapsin promoter was included as an infection marker to visualize transduced neurons. Cells were treated as in (D). Scale bars, 20 μm.
-
Figure 3—source data 1
Primary data for luminescence graphs in Figure 3.
- https://cdn.elifesciences.org/articles/70881/elife-70881-fig3-data1-v1.xlsx
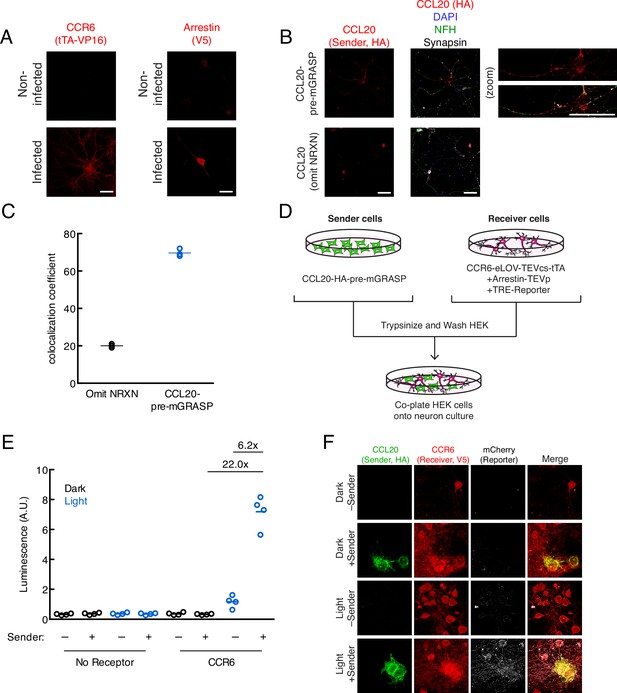
Additional characterization of TRACC (Transcriptional Readout Activated by Cell-cell Contacts) constructs in neuron culture.
(A) Confocal imaging of receiver constructs expressed in primary rat cortical neurons. Neurons were fixed and immunostained with anti-VP16 and anti-V5 to detect the GPCR and arrestin components, respectively. Scale bars, 20 μm. (B) Confocal imaging of the CCL20 sender construct expressed in neuron culture. Neurons were fixed and immunostained with anti-HA to detect the CCL20 sender construct. Neurofilament (NFH) and synapsin were co-stained to visualize neurons and pre-synaptic terminals, respectively. A sender construct omitting the intracellular neurexin domain was included as a control. Scale bars, 20 μm. (C) Quantification of colocalization between HA (sender) and synapsin signals (n = 3 fields of view [FOVs] per condition). (D) Experimental design for co-plating receiver neurons and sender HEK293T cells. (E) Luciferase assay using receiver neurons co-cultured with sender HEK293T cells. Primary rat cortical neurons were infected with AAV1/2 viruses encoding the receiver constructs, using TRE-luciferase on DIV5. HEK293T cells expressing the sender construct were co-plated onto sender neurons on DIV9, and the resulting co-culture was light-stimulated on DIV10. Approximately 24 hr after 10 min blue light exposure, the TRE-luciferase luminescence was recorded using a plate reader (n = 4 replicates per condition). (F) Confocal fluorescence imaging of sender HEK293T cells co-cultured with receiver neurons, using TRE-mCherry. Cells were treated as in (E). Scale bars, 20 μm.
-
Figure 3—figure supplement 1—source data 1
Primary data for colocalization and luminescence graphs in Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/70881/elife-70881-fig3-figsupp1-data1-v1.xlsx
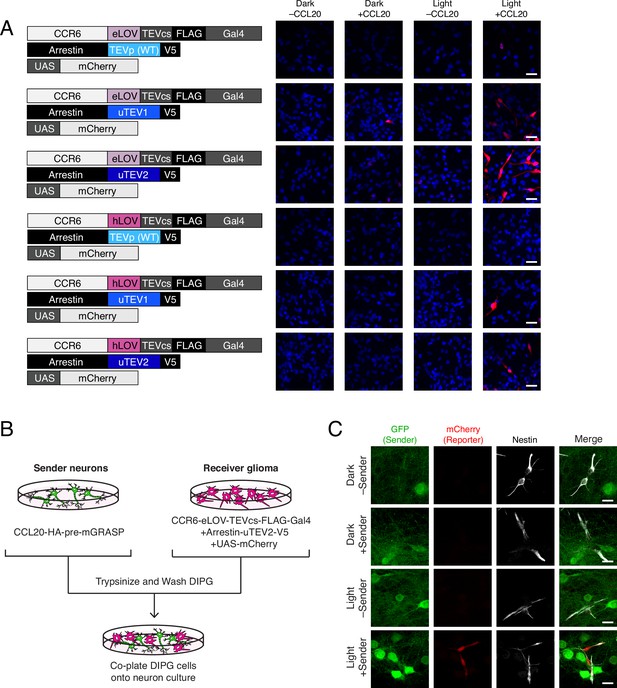
Using TRACC (Transcriptional Readout Activated by Cell-cell Contacts) to detect contacts in DIPG (diffuse intrinsic pontine glioma) culture.
(A) Optimization of TRACC components in transposon-integrated SU-DIPG-VI stable cell lines. We compared TRACC constructs containing eLOV or hLOV, and WT TEVp, uTEV1, or uTEV2. Cells were plated and treated with 0.2 μg/mL recombinant CCL20 and 10 min blue light. Approximately 24 hr after blue light exposure, cells were fixed and immunostained. Scale bar, 30 μm. (B) Experimental design for co-plating sender neurons and receiver DIPG cells. (C) Confocal fluorescence imaging of DIPG glioma expressing receiver constructs containing eLOV and uTEV2 co-plated with sender neurons. Primary rat cortical neurons were infected with AAV1/2 viruses encoding the sender construct on DIV5. DIPG cells were co-plated onto sender neurons on DIV9, and the resulting co-culture was light-stimulated on DIV11. Approximately 24 hr after 10 min blue light exposure, cells were fixed and immunostained. GFP driven by the synapsin promoter was included as an infection marker to visualize transduced neurons; Nestin is a marker for DIPG cells. Scale bar, 20 μm.
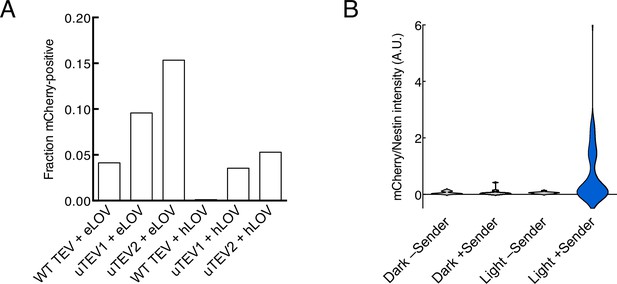
Quantification of activation in diffuse intrinsic pontine glioma (DIPG) stable lines.
(A) Quantification of mCherry activation upon recombinant CCL20 addition and light stimulation for each transposon-integrated DIPG stable line. 97–208 cells were analyzed for each condition. (B) Quantification of mCherry/Nestin intensity ratios for all Nestin-positive DIPG cells in Figure 4C. 10–35 cells were analyzed for each condition.
-
Figure 4—figure supplement 1—source data 1
Primary data for cell count graphs in Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/70881/elife-70881-fig4-figsupp1-data1-v1.xlsx





