A cellular and molecular atlas reveals the basis of chytrid development
Figures
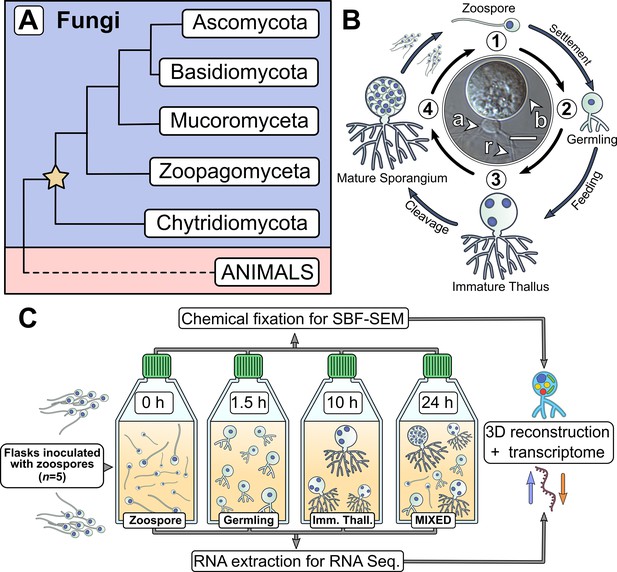
Chytrids are a major fungal phylum with a dimorphic life cycle.
(A) Chytrids (phylum Chytridiomycota) are a major fungal lineage, many members of which exhibit cellular characteristics retained from the last common ancestor of branching (rhizoidal and hyphal) fungi (star). Simplified phylogenetic tree from Laundon and Cunliffe, 2021; Tedersoo et al., 2018. (B) Rhizoclosmatium globosum exhibits an archetypal chytrid life cycle and cell plan delineated here into four discrete major stages. Labelled is the apophysis (a), cell body (b), and rhizoids (r). Scale bar = 10 µm. (C) Diagrammatic workflow of the experimental setup used in this study for comparative cellular serial block face scanning electron microscopy (SBF-SEM) and molecular (transcriptome) analysis.
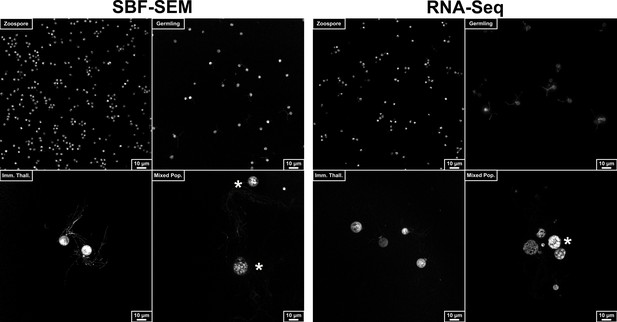
Representative images from confocal surveys conducted to assess the synchronicity of cell cultures for serial block face scanning electron microscopy (SBF-SEM) and RNA-Seq harvesting.
Cells diluted 1:1000, fixed in 0.2% formaldehyde, and stained with FM 1–43 FX to visualise cell membranes. Asterisks mark mature zoosporangia in mixed populations.
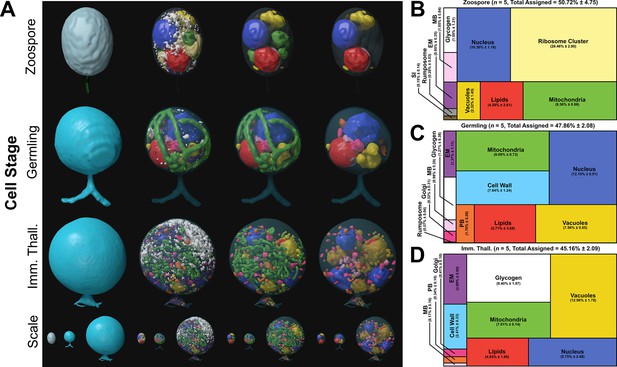
Serial block face scanning electron microscopy (SBF-SEM) reconstructions of the Rhizoclosmatium globosum life cycle.
(A) Representative SBF-SEM reconstructions of the first three life stages of the R. globosum life cycle. Bottom row shows the stages to scale. Organelle colours as in (B–D) and conserved throughout. (B–D) Volumetric composition of assigned organelles in SBF-SEM reconstructions (n = 5) of zoospores (B), germlings (C), and immature thalli (D). EM = endomembrane, MB = microbodies, PB = peripheral bodies, SI = striated inclusion.
-
Figure 2—source data 1
Data associated with Figure 2.
- https://cdn.elifesciences.org/articles/73933/elife-73933-fig2-data1-v1.xlsx
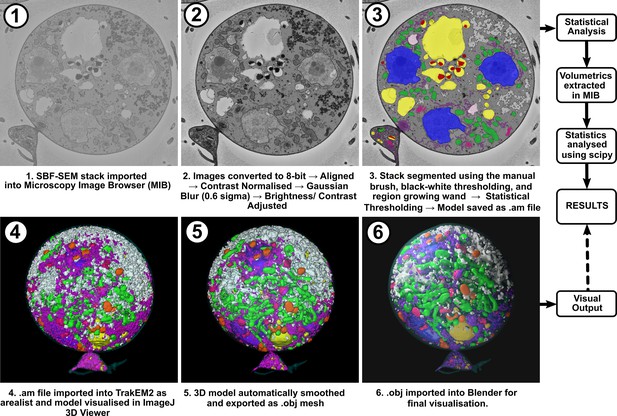
Workflow of the image analysis protocol used to generate and visualise 3D reconstructions of chytrid cells from serial block face scanning electron microscopy (SBF-SEM) stacks.
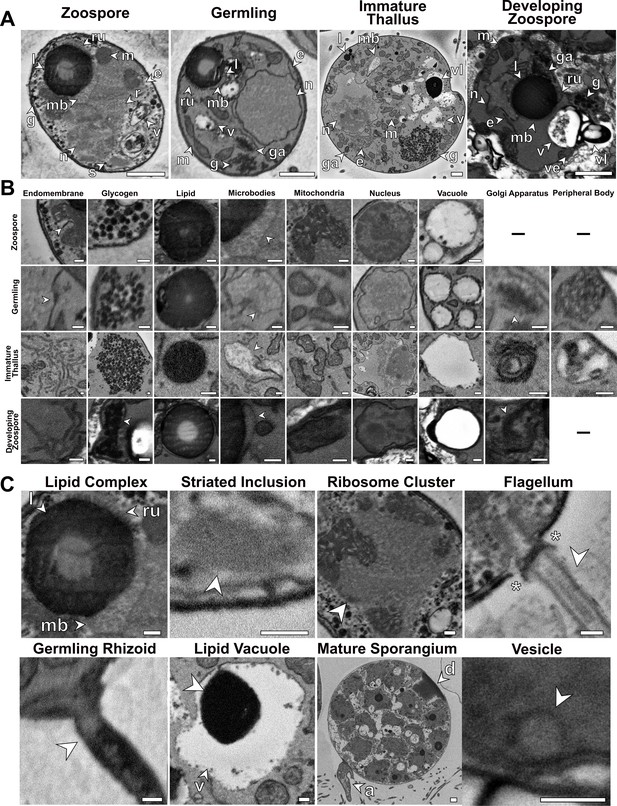
Examples of subcellular components identified in this study taken from single serial block face scanning electron microscopy (SBF-SEM) slices.
(A) Whole cell slices of each chytrid life stage showing the localisation and orientation of subcellular structures in context. (B) High magnification images of individual subcellular structures identified across life stages, where present. (C) Individual subcellular structures largely unique to individual life stages. a = apophysis, d = discharge plug, e = endomembrane, g = glycogen, ga = Golgi apparatus, l = lipid globule, m = mitochondria, mb = microbodies, n = nucleus, r = ribosomal cluster, ru = rumposome, s = striated inclusion, v = vacuoles. Asterisks in (C) show electron-dense plate at the base of the zoospore flagella. Scale bars = 1 µm (A) and 0.2 µm (B–C).
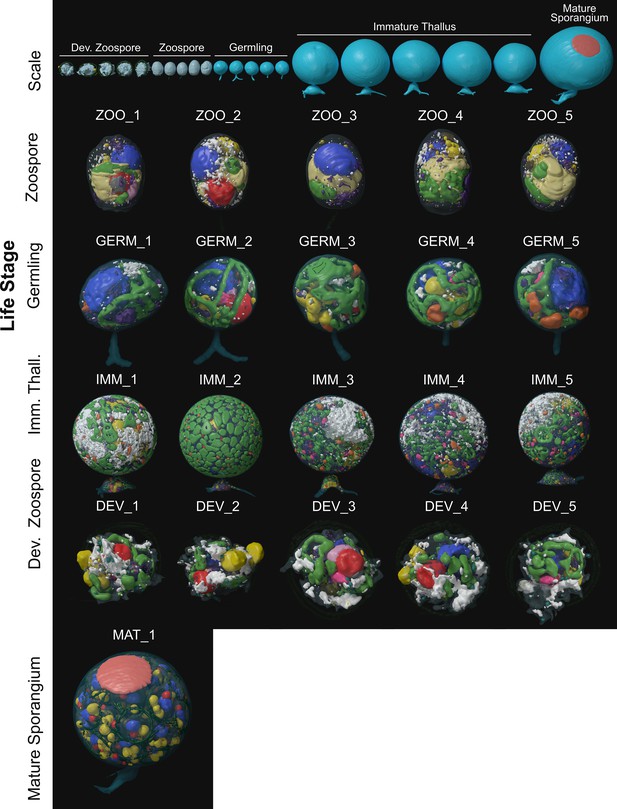
Individual 3D serial block face scanning electron microscopy (SBF-SEM) reconstructions of Rhizoclosmatium globosum cells (not to scale) across life stages labelled with replicate IDs.
Organelle colours as in Figure 2A–D. Top row shows all replicates to scale.
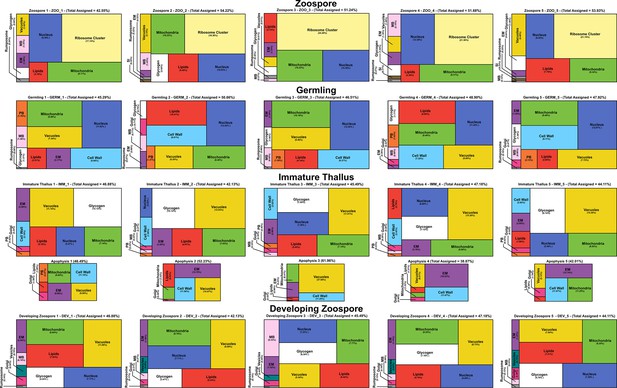
Individual volumetric compositions of assigned organelles from Rhizoclosmatium globosum serial block face scanning electron microscopy (SBF-SEM) reconstructions across life stages labelled with replicate IDs.
Organelle colours as in Figure 2A–D.
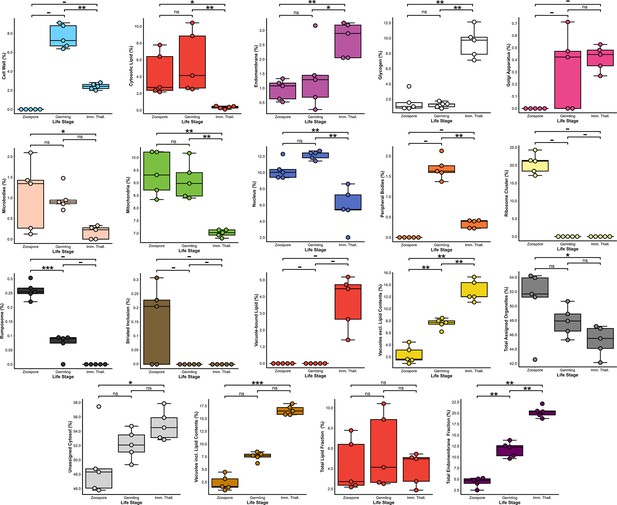
Comparisons of volumetric proportions of subcellular structures across chytrid life stages (n = 5).
n.s p > 0.05 (not significant), *p < 0.05, **p < 0.01, ***p < 0.001.
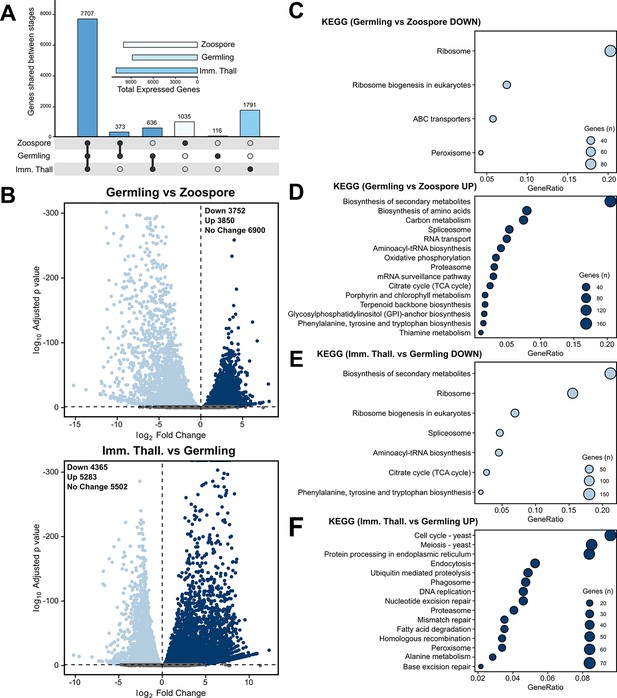
Transcriptome analysis of the Rhizoclosmatium globosum life cycle.
(A) Shared and unique gene mRNA counts between life stages. Inset shows total gene mRNA counts per life stage. (B) Pairwise comparison of differentially expressed genes (DEGs) between germlings and zoospores, and immature thalli and germlings. (C–F) Pairwise comparison of significant (n = 3, p < 0.05) Kyoto Encyclopedia of Genes and Genomes (KEGG) categories between germlings and zoospores (C–D), and immature thalli and germlings (E–F).
-
Figure 3—source data 1
Data associated with Figure 3.
- https://cdn.elifesciences.org/articles/73933/elife-73933-fig3-data1-v1.xlsx
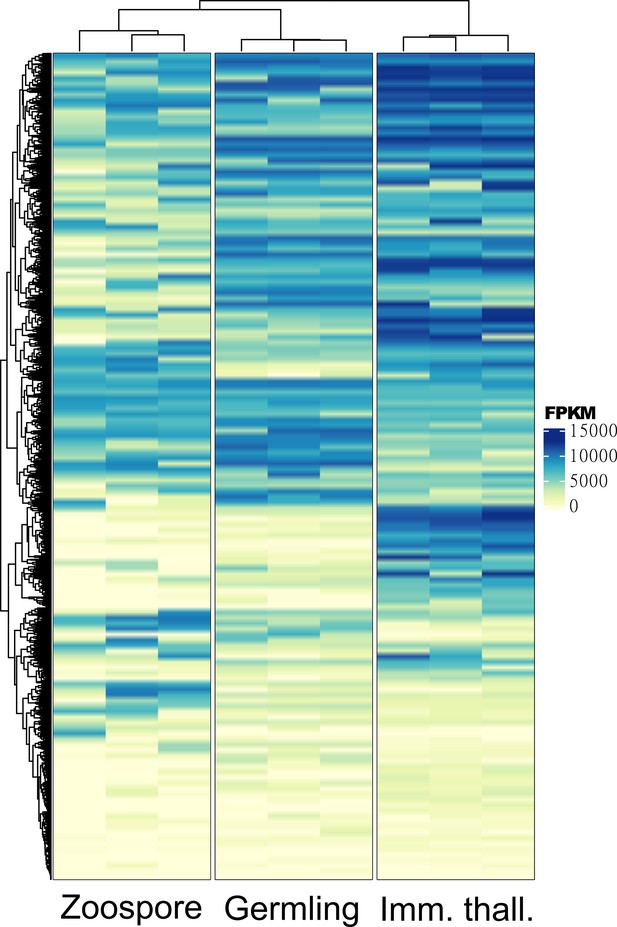
Heatmap clustering of all differentially expressed genes (DEGs) between zoospore, germling, and immature thallus replicates.
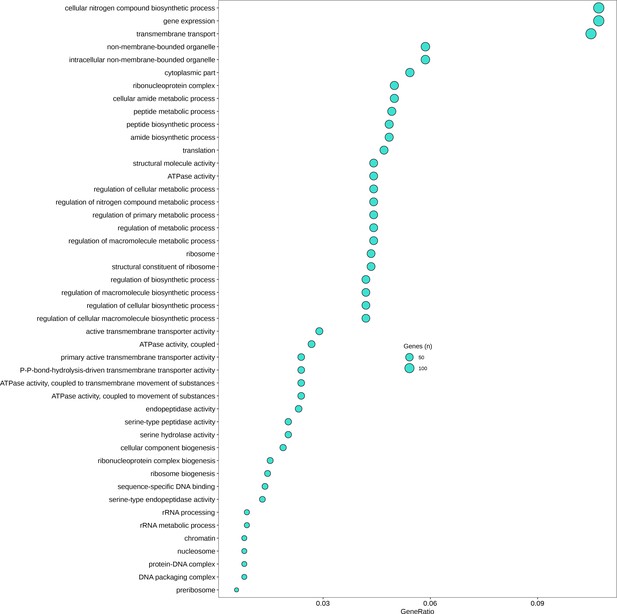
Gene Ontology (GO) categories significantly downregulated (p < 0.05) in germlings relative to zoospores.
GeneRatio is number of genes significantly upregulated/total number of genes. Circle size represents numbers of genes.
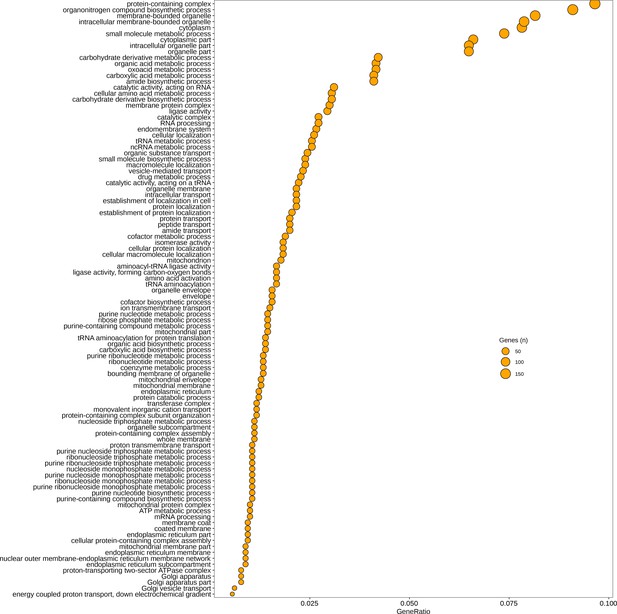
Gene Ontology (GO) categories significantly upregulated (p < 0.05) in germlings relative to zoospores.
GeneRatio is number of genes significantly upregulated/total number of genes. Circle size represents numbers of genes.
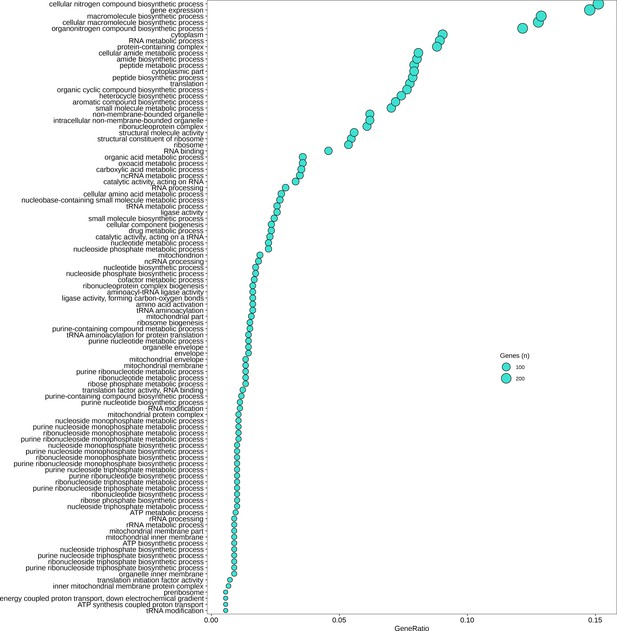
Gene Ontology (GO) categories significantly downregulated (p < 0.05) in immature thalli relative to germlings.
GeneRatio is number of genes significantly upregulated/total number of genes. Circle size represents numbers of genes.
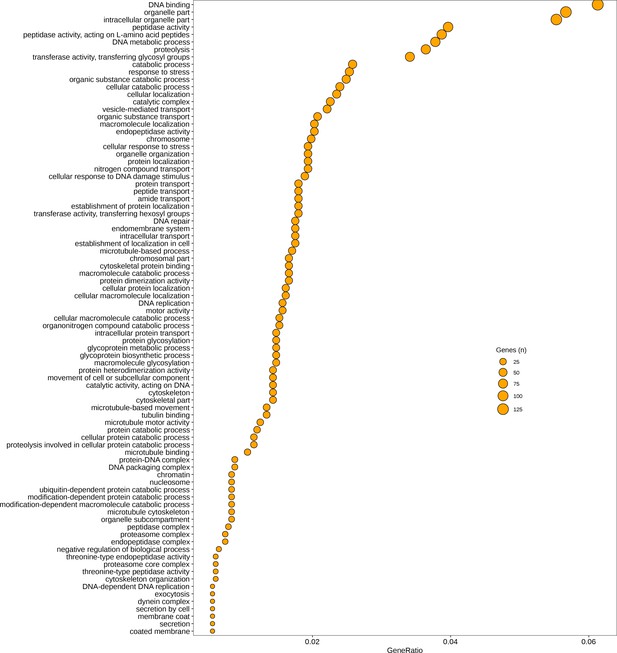
Gene Ontology (GO) categories significantly upregulated (p < 0.05) in immature thalli relative to germlings.
GeneRatio is number of genes significantly upregulated/total number of genes. Circle size represents numbers of genes.
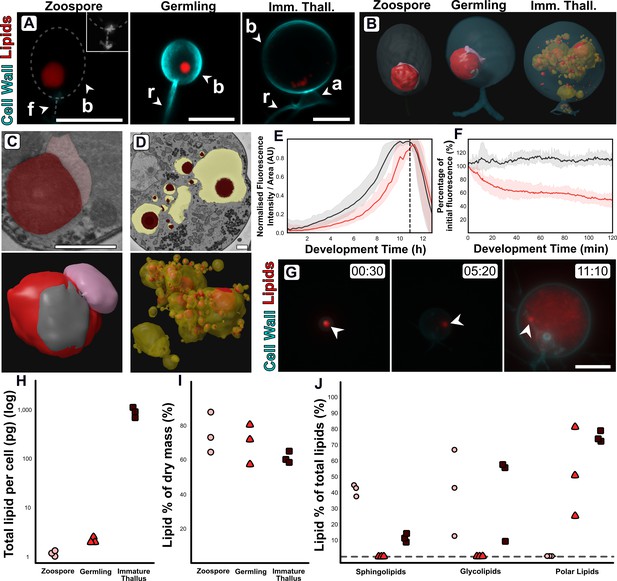
Changes in lipid and lipid-associated cell structures occur with transitions between life stages.
(A) Fluorescent labelling of Rhizoclosmatium globosum shows distinct shifts in lipid structures across the chytrid life cycle and cell wall. Dashed line demarks cell boundary where not labelled in the zoospore. Zoospore inset shows precursory cell wall material at the flagellar base contrast-brightness adjusted for visualisation. Apophysis (a), cell body (b), flagellum (f), rhizoid (r). Scale bars = 5 µm. (B) Representative serial block face scanning electron microscopy (SBF-SEM) reconstructions of lipid globules and lipid-associated structures across chytrid life stages. (C–D) Representative single false-coloured SBF-SEM slices (top) and SBF-SEM reconstructions (bottom) of the lipid-rumposome-microbody (LRM) complex from zoospores (also seen in germlings) (C) and intravacuolar lipid globules (D) from immature thalli. Scale bars = 1 µm. (E) Live-cell imaging (n = 5) of R. globosum population-level Nile red-stained lipid dynamics. Red = mean lipid fluorescence (±min/max), black = mean total cell area (±min/max), dashed line = mean sporulation time of population. (F) Immediately following zoospore settlement, the population-level (n = 5) lipid fluorescence (red) decreases relative to fixed photobleaching control populations (black). (G) Live-cell imaging revealed differential lipid dynamics across the chytrid life cycle. Note that the original zoospore lipid globule (arrowhead) remains intact up to the point of lipid anabolism in the immature thallus. Timestamp = HH:MM. Scale bar = 10 µm. (H–J) Lipid analysis shows shifts in lipid composition of the chytrid life cycle. Lipid quantities as total mass per cell (H) and as a percentage of total dry mass (I) between chytrid life stages. Changes in lipid fractions were found between chytrid life stages (J). Dashed line = below analytical detection.
-
Figure 4—source data 1
Data associated with Figure 4.
- https://cdn.elifesciences.org/articles/73933/elife-73933-fig4-data1-v1.xlsx
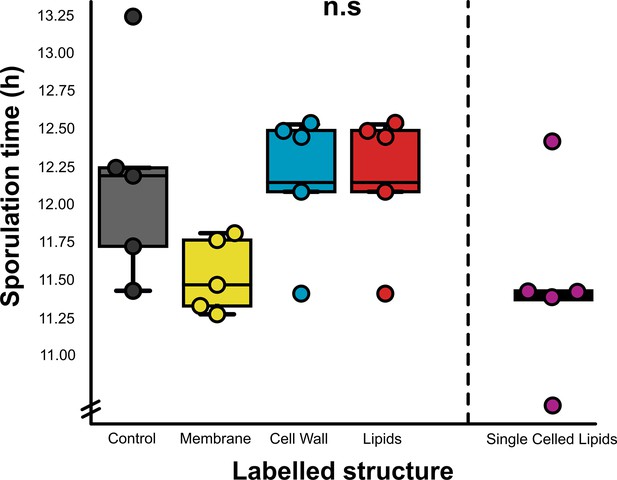
Comparison of sporulation times (as a proxy for normal cell development) for dye-labelled chytrid populations (n = 5) imaged by live-cell microscopy, relative to no-dye controls.
n.s. p > 0.05 (not significant).
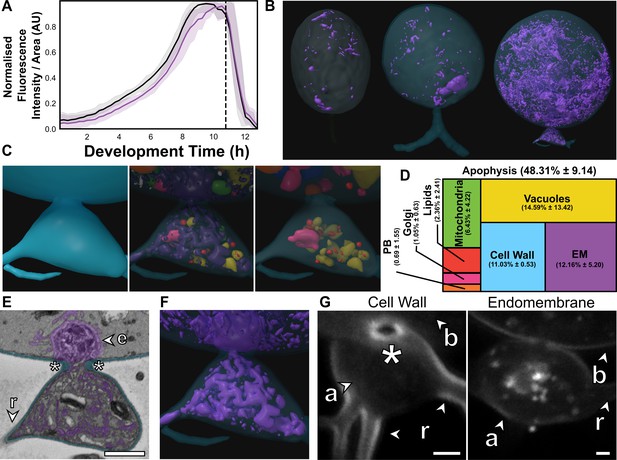
The apophysis is a distinct subcellular structure characterised by increased endomembrane trafficking.
(A) Live-cell imaging (n = 5) of Rhizoclosmatium globosum population-level FM1–43-stained endomembrane dynamics. Purple = mean endomembrane fluorescence (±min/max), black = mean total cell area (±min/max), dashed line = mean sporulation time of population. (B–C) Representative serial block face scanning electron microscopy (SBF-SEM) reconstructions of endomembrane across chytrid life stages (B) and the apophysis from immature thalli (C). Volumetric composition of SBF-SEM reconstructions (n = 5) of immature thallus apophyses (D). Representative single false-coloured SBF-SEM slice (E) and reconstruction (F) of the endomembrane and thickened cell wall (asterisk) at the apophysis-cell body junction. Fluorescent labelling of the chitin-rich wall around the apophysis-cell body connecting pore and associated endomembrane structures (G). Labels as in Figure 4A. All scale bars = 1 µm.
-
Figure 5—source data 1
Data associated with Figure 5.
- https://cdn.elifesciences.org/articles/73933/elife-73933-fig5-data1-v1.xlsx
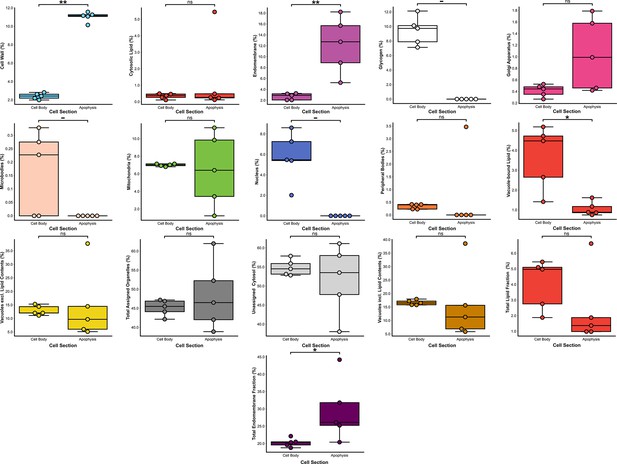
Comparisons of volumetric proportions of subcellular structures between immature thalli cell bodies and their corresponding apophyses (n = 5).
n.s. p > 0.05 (not significant), *p < 0.05, **p < 0.01, ***p < 0.001.
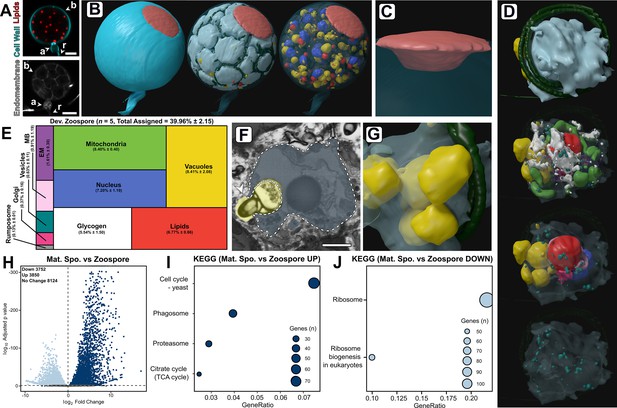
Developing zoospores in the zoosporangium displayed an amoeboid morphology with putative endocytotic activity.
(A) Fluorescent labelling of lipids, cell wall, and endomembrane in a Rhizoclosmatium globosum mature zoosporangium. Scale bar = 5 µm. (B–C) Serial block face scanning electron microscopy (SBF-SEM) reconstructions of an 82-zoospore containing mature zoosporangium (B) highlighting the discharge plug, shown in coral (C). (D) Representative SBF-SEM reconstructions of a developing zoospore. Organelle colours as in E. (E) Volumetric composition of SBF-SEM reconstructions of developing zoospores (n = 5). (F–G) Representative single false-coloured SBF-SEM slice (F) and reconstruction (G) of the putative endocytotic vacuoles in developing zoospores. Dashed line delineates the zoospore cell boundary in (F). Scale bar = 1 µm. (H) Pairwise comparison of differentially expressed genes (DEGs) between mature zoosporangia and the free-swimming zoospore life stage. (I–J) Pairwise comparison of significant Kyoto Encyclopedia of Genes and Genomes (KEGG) categories between mature zoosporangia and the free-swimming zoospore life stage.
-
Figure 6—source data 1
Data associated with Figure 6.
- https://cdn.elifesciences.org/articles/73933/elife-73933-fig6-data1-v1.xlsx
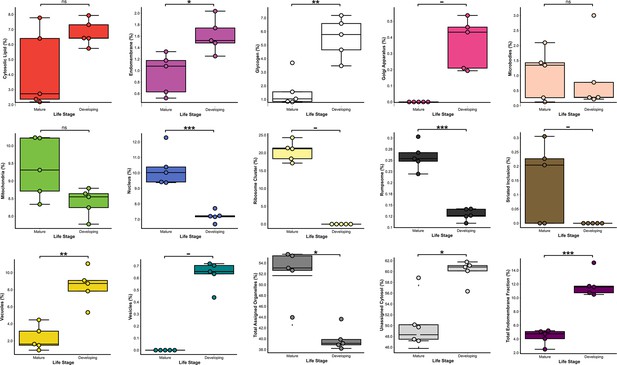
Comparisons of volumetric proportions of subcellular structures between developing and mature zoospores (n = 5).
n.s. p > 0.05 (not significant), *p < 0.05, **p < 0.01, ***p < 0.001.
Videos
Serial block face scanning electron microscopy (SBF-SEM) reconstructions allowed the structural comparison of life stages in Rhizoclosmatium globosum.
Representative SBF-SEM reconstructions of the zoospore, germling, and immature thallus life stages for comparison. Zoospore and germling cells shown to scale at the beginning of the video, and later enlarged.
Structural shifts in lipid globules were observed across Rhizoclosmatium globosum life stages, associated with the change from catabolism/conversion to anabolism.
Representative serial block face scanning electron microscopy (SBF-SEM) reconstructions of the zoospore, germling, and immature thallus lipid structures for comparison.
The zoospore lipid globule remained as an intact structure across the Rhizoclosmatium globosum life cycle.
Automated particle tracking of lipid globules (red) across the chytrid life cycle. Magenta circles mark individual lipid globules. Yellow track shows particle tracking of the initial lipid globule into the period of lipid anabolism. Cell wall shown in cyan. Timestamp = HH:MM.
The Rhizoclosmatium globosum apophysis is structurally dominated by endomembrane structures.
Representative serial block face scanning electron microscopy (SBF-SEM) reconstruction of a chytrid apophysis from an immature thallus.
The Rhizoclosmatium globosum apophysis links intracellular trafficking between the rhizoids and cell body.
Live-cell imaging of endomembrane dynamics in the chytrid apophysis. The apophysis links endomembrane dynamics between the rhizoid system and thallus. Shown are DIC (left), endomembrane (centre), and overlay (right) channels. Timestamp = MM:SS.
Serial block face scanning electron microscopy (SBF-SEM) reconstruction of a Rhizoclosmatium globosum mature zoosporangium.
Developingidual 3D serial block face scanning electron micr zoospores were more amoeboid than mature zoospores in Rhizoclosmatium globosum, putatively due to elevated endocytosis and trafficking.
Representative serial block face scanning electron microscopy (SBF-SEM) reconstructions of the ‘mature’ zoospore and developing zoospore life stages for comparison.
Additional files
-
Supplementary file 1
Mean volumetric quantities of cellular structures recorded across chytrid life stages.
- https://cdn.elifesciences.org/articles/73933/elife-73933-supp1-v1.docx
-
Supplementary file 2
Mean numerical quantities of cellular structures recorded across chytrid life stages.
- https://cdn.elifesciences.org/articles/73933/elife-73933-supp2-v1.docx
-
Supplementary file 3
Mean volumetric percentages and statistical comparisons of cellular structures recorded across chytrid life stages.
- https://cdn.elifesciences.org/articles/73933/elife-73933-supp3-v1.docx
-
Supplementary file 4
Mean volumetric percentages and statistical comparisons of cell bodies and their corresponding apophyses in immature thalli.
- https://cdn.elifesciences.org/articles/73933/elife-73933-supp4-v1.docx
-
Supplementary file 5
Mean volumetric percentages and statistical comparisons of free-swimming and developing zoospores.
- https://cdn.elifesciences.org/articles/73933/elife-73933-supp5-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/73933/elife-73933-transrepform1-v1.docx
-
Source code 1
Python script used to quantify population-level fluorescence of developing chytrid cells (Figures 4 and 5).
- https://cdn.elifesciences.org/articles/73933/elife-73933-code1-v1.zip
-
Source code 2
Python script used to quantify single-cell Nile Red fluorescence of settled chytrid zoospores (Figure 4).
- https://cdn.elifesciences.org/articles/73933/elife-73933-code2-v1.zip






