C/EBPδ-induced epigenetic changes control the dynamic gene transcription of S100a8 and S100a9
Figures
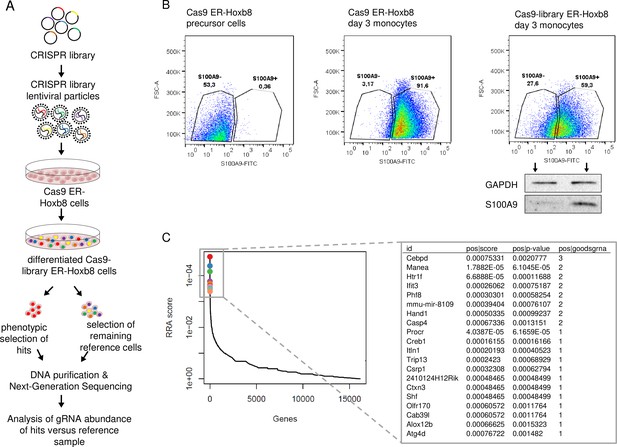
Genome-Scale CRISPR Knockout lentiviral pooled library screen to identify S100A9 regulators.
(A) For genome-wide screen, over 100,000 plasmids, each containing a guide RNA towards different early consecutive exons, were packaged into lentiviral particles. Cas9-expressing ER-Hoxb8 cells were pool-transduced, selected, and differentiated to induce S100A9 expression. Hits and reference cells were collected by sorting according to their phenotypes of interest. DNA of both samples was purified for next-generation sequencing and subsequent analysis. (B) Precursor and differentiated Cas9 and Cas9-library ER-Hoxb8 cells were stained intracellularly for S100A9 using a FITC-labelled antibody. Cas9-library ER-Hoxb8 day 3 monocytes with no or lower S100A9 expression were sorted as hits, the remaining cells served as reference cells. (C) Data was analysed using the Model-based Analysis of Genome-wide CRISPR-Cas9 Knockout (MAGeCK) software for identification of enriched guide RNAs in the hit sample. Corresponding genes were rank-ordered by robust rank aggregation (RRA) scores. The list states the top 20 genes according to RRA scores, arranged after the number of guides that are enriched in the hit sample. See also Figure 1—figure supplement 1 and Figure 1—source data 1.
-
Figure 1—source data 1
Gene summary of Model-based Analysis of Genome-wide CRISPR-Cas9 Knockout (MaGECK) analysis.
- https://cdn.elifesciences.org/articles/75594/elife-75594-fig1-data1-v2.xlsx
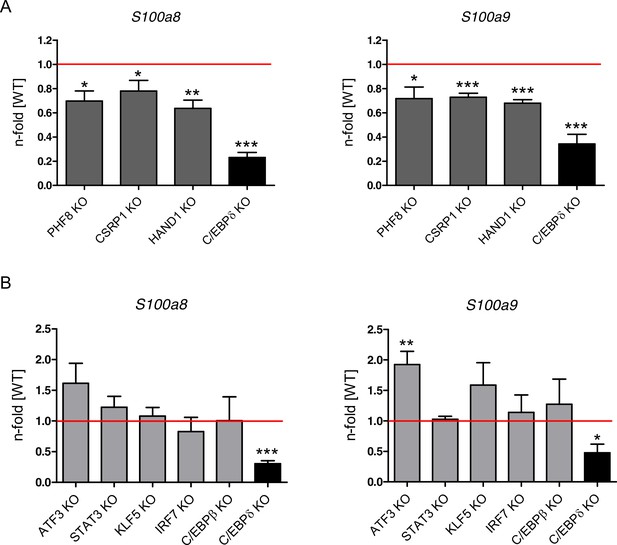
Relative S100a8 and S100a9 expression in differentiated single knockout (KO) ER-Hoxb8 monocytes.
Cell lines generated by CRISPR/Cas9 (A: PHF8, CSRP1, and HAND1 KO, dark grey bars = candidates from Genome-Scale CRISPR/Cas9 Knockout (GeCKO) screen, B: ATF3, STAT3, KLF5, IRF7, and C/EBPβ KO, light grey bars = candidates from previous studies) were compared to a non-targeting CRISPR/Cas9 control cell line. Cell lines generated from transgenic mice (C/EBPδ KO, black bars = hit from GeCKO screen) were compared to wildtype (WT) counterparts. All cells were differentiated to until day 2 or day 3 and relative S100a8 and S100a9 mRNA levels were analysed using quantitative reverse transcription polymerase chain reaction (qRT-PCR). Red lines indicate relative s100 level of controls (WT). Values are the means ± SEM of four experiments. *p<0.05, **p<0.01, ***p<0.001 by two-tailed Student’s t test.
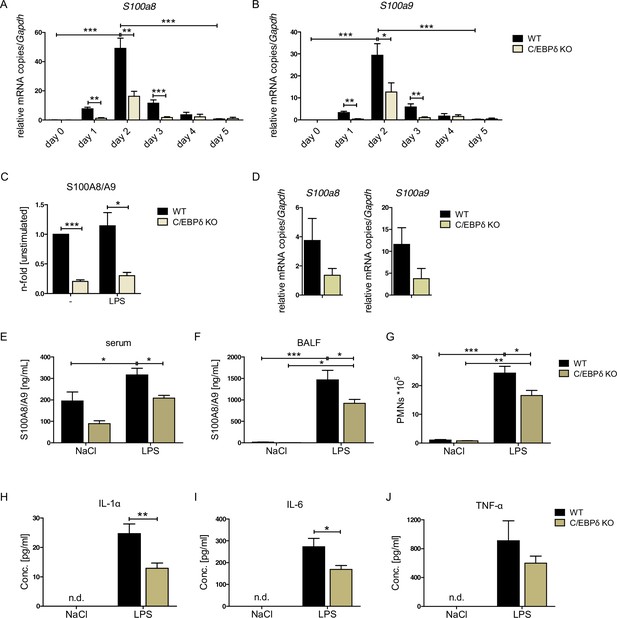
S100A8 and S100A9 expression in wildtype (WT) and C/EBPδ knockout (KO) monocytes.
(A) Relative S100a8 and (B) S100a9 mRNA level during differentiation of WT and C/EBPδ KO ER-Hoxb8 monocytes were measured using quantitative reverse transcription polymerase chain reaction (qRT-PCR) (n=3–8). (C) S100A8/A9 concentrations in supernatant of differentiation day 4 of WT and C/EBPδ KO ER-Hoxb8 monocytes stimulated with 10 ng LPS for 4 hr or left untreated (n=3) were quantified using our in-house mouse S100A8/S100A9 sandwich enzyme-linked immunosorbent assay (ELISA) (n=6). (D) Relative S100a8 and S100a9 mRNA levels of bone marrow-derived mouse monocytes were measured using qRT-PCR (n=3). (E) Serum and (F) bronchoalveolar lavage fluid (BALF) of LPS- (WT: n=9, C/EBPδ KO: n=11) or NaCl-only (WT: n=6, C/EBPδ KO: n=3) exposed mice were harvested 4 hr after onset of lung inflammation and were analysed for S100A8/A9 expression, (G) for amount of recruited polymorphonuclear leukocytes (PMNs) and for (H) IL-1α, (I) IL-6, and (J) TNF-α production in BALF. Values are the means ± SEM. *p<0.05, **p<0.01, ***p<0.001, by two-tailed Student’s t test (A–C, H–J) and by one-way ANOVA with Bonferroni’s correction (E–G). See also Figure 2—figure supplements 1 and 2.
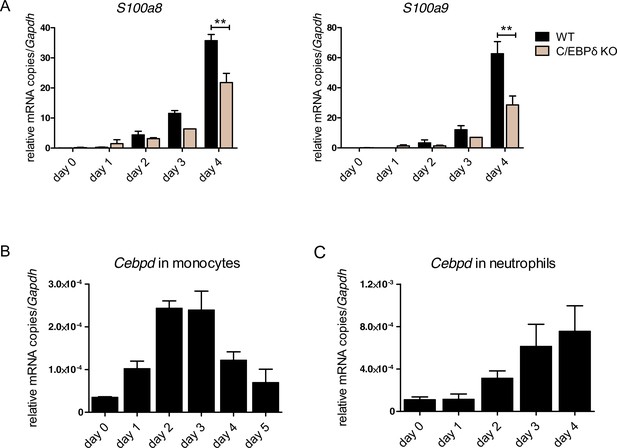
S100a8, S100a9, and Cebpd expression kinetics in ER-Hoxb8 cells.
(A) S100a8 and S100a9 mRNA levels were measured using qPCR (left, middle) and protein levels by using our in-house mouse S100A8/S100A9 sandwich enzyme-linked immunosorbent assay (ELISA) (right) in precursor and differentiated wildtype (WT) and C/EBPδ knockout (KO) ER-Hoxb8 neutrophils were analysed. Cebpd mRNA level was measured using qPCR in precursor and differentiated WT ER-Hoxb8 monocytes (B) and neutrophils (C). Values are the means ± SEM of three experiments. **p<0.01, by Student’s t test.
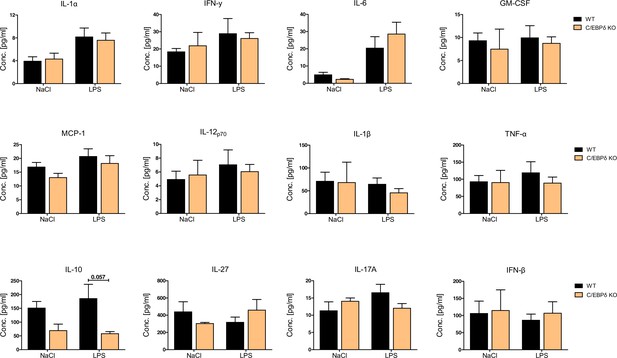
Baseline (NaCl) and LPS-induced cytokine production in sera of wildtype (WT) and C/EBPδ knockout (KO) mice.
Serum of LPS- (WT: n=9, C/EBPδ KO: n=11) or NaCl-only (WT: n=6, C/EBPδ KO: n=3) exposed mice was harvested 4 hr after onset of lung inflammation and was analysed for various cytokines using the bead-based immunoassay LEGENDplex. Values are the means ± SEM. p=0.057 by one-way ANOVA with Bonferroni’s correction.
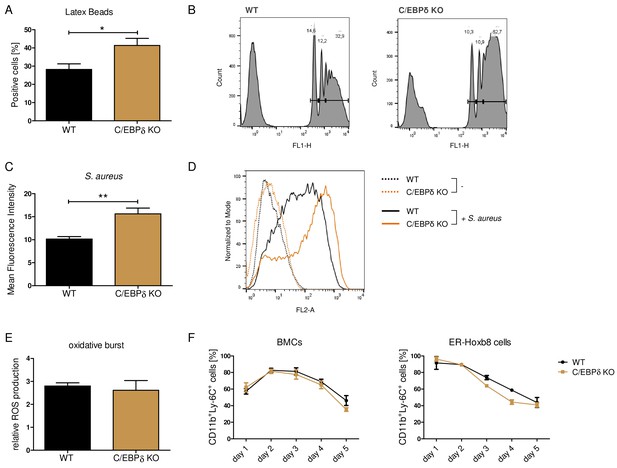
Functional properties of wildtype (WT) and C/EBPδ knockout (KO) ER-Hoxb8 monocytes.
Differentiated WT and C/EBPδ KO ER-Hoxb8 cells were incubated with green fluorescent latex beads (A, B) or with pHrodo Red Staphylococcus aureus Bioparticles Conjugate (C, D) for 2 hr at a ratio of 10 beads per cells. The phagocytosis was measured using flow cytometry. (A) Percentages of phagocytosis of ≥3 beads or (C) gMFI shifts of S. aureus -treated cells in relation to control cells were used for quantification. Representative histogram plots of Latex Beads- (B) and S. aureus Bioparticles- (D) mediated phagocytosis are presented. (B) Histogram plots, showing gates for phagocytosis of one (.), two (...), or ≥3 or more than three (…) beads. (E) Differentiated WT and C/EBPδ KO ER-Hoxb8 cells were treated with 10 nM PMA for 15 min and 15 µM DHR123 to measure oxidative burst was measured using DHR123. The fluorescence of the cells was measured using flow cytometry. Relative ROS production as = gMFI shifts of PMA-treated cells in relation to non-treated cells is shown. (F) Proportion of CD11b+Ly-6C+ cells during differentiation of bone marrow cells (BMCs) and ER-Hoxb8 cells was analysed using flow cytometry. Values are the means ± SEM of three to four experiments. *p<0.05, **p<0.01 by two-tailed Student’s t test. See also Figure 3—figure supplements 1 and 2.
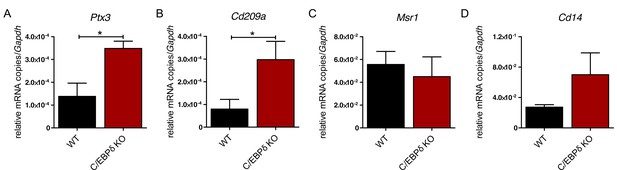
Differential expression of phagocytosis-related genes in wildtype (WT) and C/EBPδ knockout (KO) monocytes.
(A) Ptx3, (B) Cd209a, (C) Msr1, and (D) Cd14 mRNA levels were analysed using qPCR in differentiated WT and C/EBPδ KO ER-Hoxb8 monocytes. Values are the means ± SEM of three to five experiments. *p<0.05, by two-tailed Student’s t test.
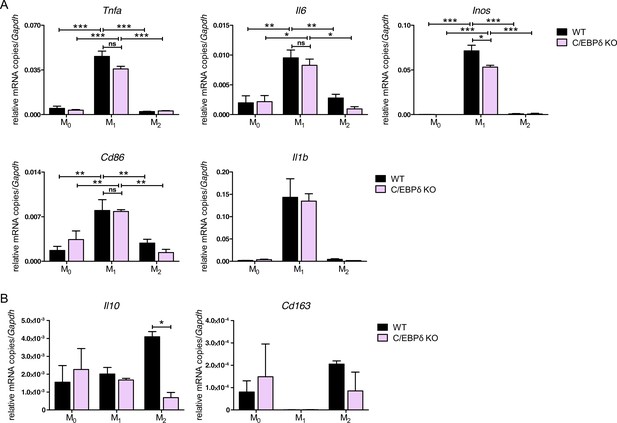
Effect of C/EBPδ deletion on polarization of bone marrow-derived monocytes.
The mRNA levels of genes (A) associated with M1-polarization (Tnfa, Il6, Inos, Cd86, and Il1b) and (B) associated with M2-polarization (Il10 and Cd163) were analysed. For this, wildtype (WT) and C/EBPδ knockout (KO) bone marrow-derived mouse monocytes (BMDMs) were either stimulated for 24 hr with 50 ng/ml IFN-γ and 10 ng/ml LPS (M1), with 20 n/ml IL-4 (M2) or left untreated (M0). Values are the means ± SEM of three experiments. *p<0.05, **p<0.01, ***p<0.001, ns = not significant, by one-way ANOVA with Bonferroni’s correction.
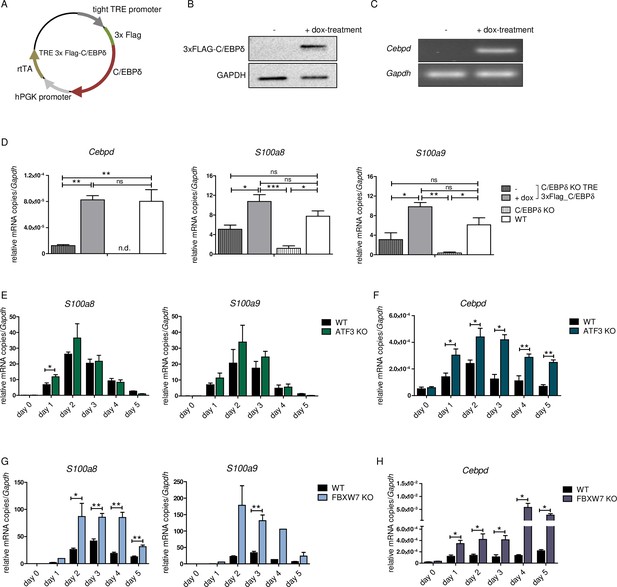
S100a8 and S100a9 expression in differentiated ER-Hoxb8 cells is dependent on C/EBPδ abundancy.
(A) Tet-On construct of inducible 3xFlag-C/EBPδ expression due to constitutively expressed rtTA (reverse tetracycline-controlled transactivator) that binds to TRE promoter upon doxycycline treatment was transduced into C/EBPδ knockout (KO) ER-Hoxb8 cells. (B) Induction of 3xFlag-C/EBPδ upon doxycycline treatment (2 µg/ml, 24 hr) was analysed by western blot and (C) quantitative reverse transcription polymerase chain reaction (qRT-PCR) in comparison to untreated cells. (D) Induction of 3xFlag-C/EBPδ was also analysed by qRT-PCR (Cebpd), as well as expression of S100a8 and S100a9 mRNAs, in untreated and dox-treated C/EBPδ KO TRE_3xFlag-C/EBPδ monocytes and in comparison to wildtype (WT) and C/EBPδ KO monocytes on differentiation day 1 (n=3). (E, G) S100a8 and S100a9, (F, H) and Cebpd mRNA levels were measured using qRT-PCR in precursor and differentiated WT and ATF3 KO (E, F) and in WT and FBXW7 KO (G, H) ER-Hoxb8 monocytes (n=3–4). Values are the means ± SEM. *p<0.05, **p<0.01, by one-way ANOVA with Bonferroni’s correction (D) and by two-tailed Student’s t test (E–H).
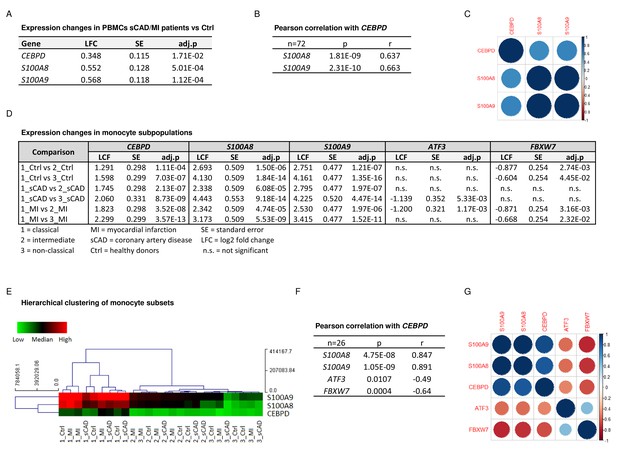
CEBPD expression positively correlates with S100A8 and S100A9 expression in proinflammatory monocytes of myocardial infarction/stable coronary artery disease (MI/sCAD) patients.
(A) Gene expression changes detected by RNA-sequencing (RNA-seq) in peripheral blood mononuclear cells (PBMCs) of BioNRW participants (n=72, sCAD/MI vs. Ctrl). LFC = log2 fold change, SE = standard error, and adj.p = adjusted p-value. (B) Pearson correlation coefficient = r, p-value = p in PBMCs and (C) corresponding correlation matrix. (D) Gene expression changes of CEBPD, S100A8, S100A9, ATF3, and FBXW7 detected by RNA-seq in monocyte subpopulations of BioNRW participants (n=26, from three individuals in each of the sCAD, MI, and Ctrl diagnostic groups). (E) Hierarchical clustering of S100A8-, S100A9-, and CEBPD normalized counts (using Euclidean distance metric with complete linkage). Shown are classical (1), intermediate (2), and non-classical (3) monocytes of healthy donors (Ctrl), MI, and sCAD patients. (F) Pearson correlation coefficient = r, p-value = p in monocytes and (G) corresponding correlation matrix. See also Figure 5—source data 1.
-
Figure 5—source data 1
Expression changes in the BioNRW monocytes dataset (RNA-sequencing [RNA-seq], n=26).
- https://cdn.elifesciences.org/articles/75594/elife-75594-fig5-data1-v2.xlsx
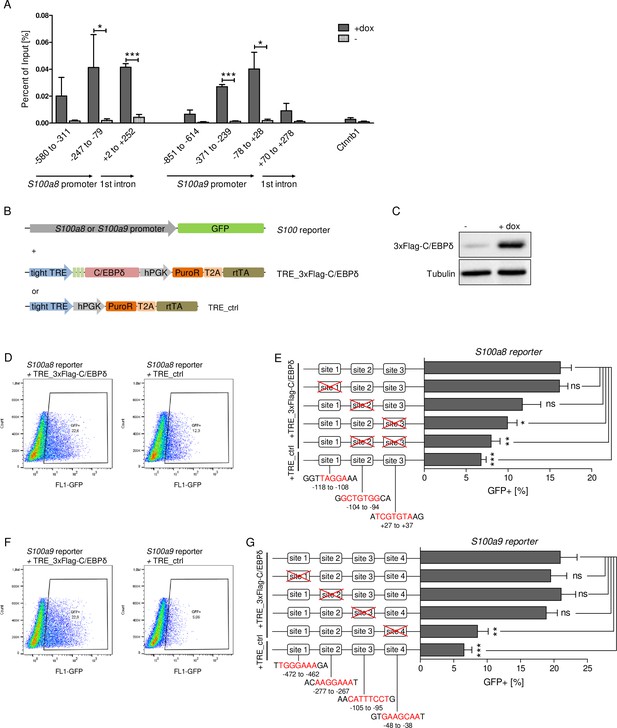
C/EBPδ binds to regions within the S100a8 and S100a9 promoters.
(A) Chromatin immunoprecipitation was performed in untreated (-) and dox-treated (+dox) TRE_3xFlag-C/EBPδ monocytes on differentiation day 1 using a Flag-antibody. Purified DNA was analysed using primer pairs flanking different S100a8 and S100a9 promoter and intronic regions and a negative control primer pair flanking a random genomic region (n=3). (B) Co-transfection of vectors carrying constructs for GFP under the S100a8 or S100a9 promoter (reporter), together with the doxycycline-dependent 3xFlag-C/EBPδ expression cassette (TRE_3xFlag-C/EBPδ) or a corresponding control vector lacking the 3xFlag-C/EBPδ expression cassette (TRE_ctrl) in HEK293T cells, was performed. (C) Induction of 3xFlag-C/EBPδ upon doxycycline treatment (2 µg/ml, 24 hr) was analysed by western blot. Representative dot plots from flow cytometry analysis show GFP+ gates of co-transfected HEK293T cells, either using TRE_3xFlag-C/EBPδ or TRE_ctrl together with S100a8 reporter (D) and with S100a9 reporter (F) upon doxycycline treatment (2 µg/ml, 24 hr). Co-transfection of TRE_3xFlag-C/EBPδ and S100a8 (E) and S100a9 (G) reporter plasmids carrying different mutated possible binding sites was performed, analysed 24 hr post-transfection and compared to co-transfection of TRE_ctrl and S100 reporter plasmid activities. Suggested C/EBP-binding sites targeted by depletion are indicated by nucleic acids marked in red (n=4–5). Values are the means ± SEM. *p<0.05, **p<0.01, ***p<0.001, ns = not significant, by two-tailed Student’s t test.
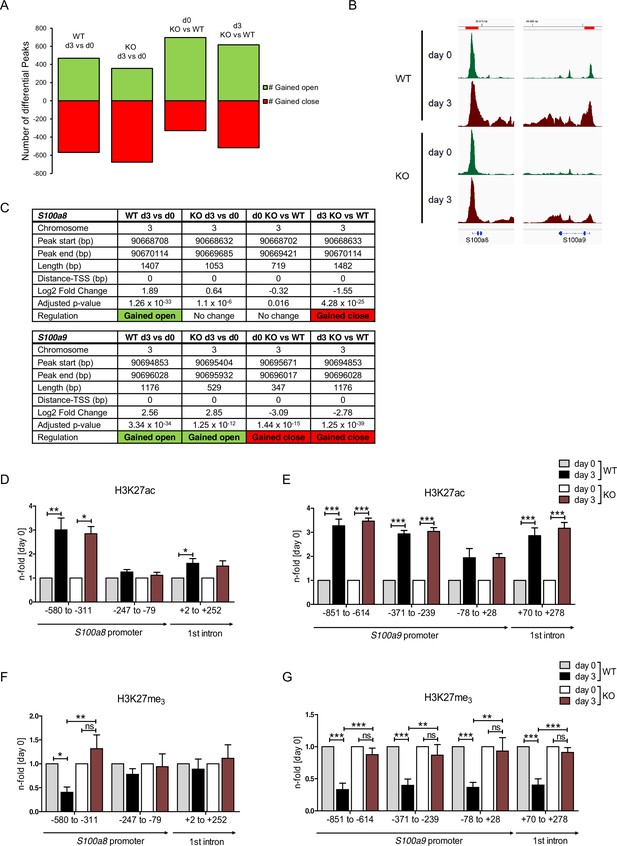
Analysis of chromatin accessibility and epigenetic features within S100a8 and S100a9 promoter regions.
Assay for transposase-accessible chromatin using sequencing (ATAC-seq) was executed in precursor (day 0=d0) and differentiated (day 3=d3) wildtype (WT) and C/EBPδ knockout (KO) (KO) ER-Hoxb8 monocytes. (A) Differential peak analysis between all conditions was performed (n=3). (B) Combined gene tracks showing ATAC-seq reads of precursor (day 0) and differentiated (day 3) WT and C/EBPδ KO cells at the S100a8 and S100a9 gene regions. (C) Differential accessibility at S100a8 and S100a9 loci was analysed for all comparisons. Chromatin immunoprecipitation was performed using anti-H3K27ac (D, E), anti-H3K27me3 (F, G) in chromatin of precursor (day 0) and differentiated (day 3) WT and C/EBPδ KO (KO) ER-Hoxb8 monocytes. Purified DNA was analysed using primer pairs flanking different S100a8 (D, F) and S100a9 (E, G) promoter regions (n=3–6). N-folds are based on percent of input values of respective day 0 ChIP-PCR samples. Values are the means ± SEM. *p<0.05, **p<0.01, ***p<0.001, ns = not significant, by one-way ANOVA with Bonferroni’s correction. See also Figure 7—figure supplement 1 and Figure 7—source data 1.
-
Figure 7—source data 1
Differential peak analysis based on assay for transposase-accessible chromatin using sequencing (ATAC-seq) data of wildtype (WT) and C/EBPδ knockout (KO) day 0 and day 3 ER-Hoxb8 cells.
- https://cdn.elifesciences.org/articles/75594/elife-75594-fig7-data1-v2.xlsx
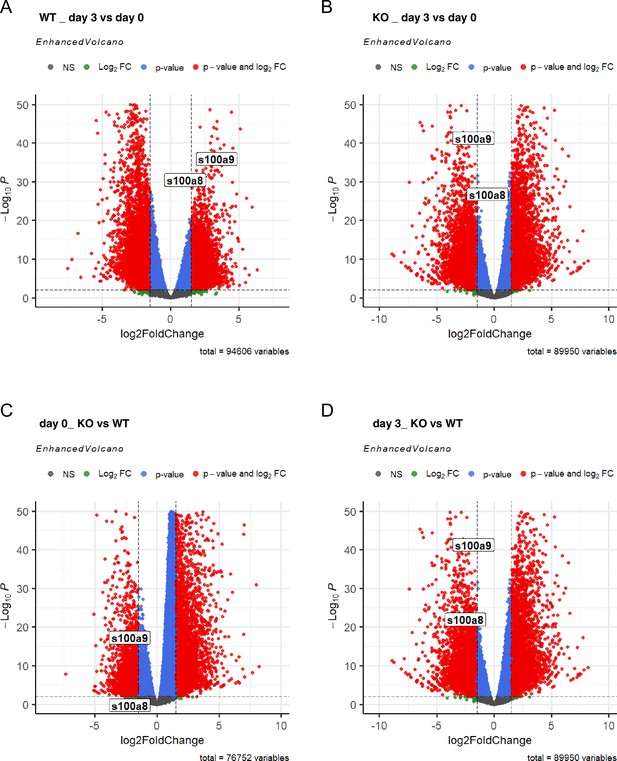
Differential peak analysis based on assay for transposase-accessible chromatin using sequencing (ATAC-seq) data of wildtype (WT) and C/EBPδ knockout (KO) day 0 and day 3 ER-Hoxb8 cells.
Volcano plots show differentially accessible regions between day 3 and day 0 (A) in WT and (B) in C/EBPδ KO cells, and between C/EBPδ KO and WT (C) at day 0 and (D) at day 3. NS = not significant; FC = fold change.
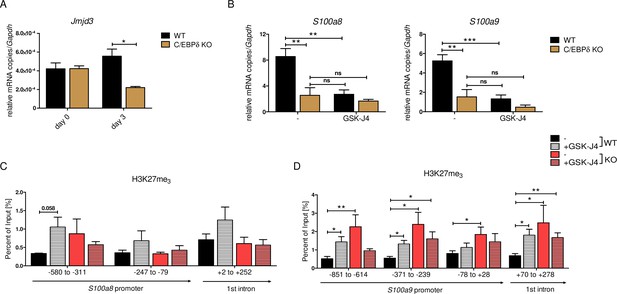
JMJD3-mediated demethylation of H3K27me3 is crucial for S100a8 and S100a9 expression.
(A) Jmjd3 mRNA levels of precursor and differentiated wildtype (WT) and C/EBPδ knockout (KO) ER-Hoxb8 cells were analysed using quantitative reverse transcription polymerase chain reaction (qRT-PCR) (n=3). (B) WT and C/EBPδ KO ER-Hoxb8 cells were treated with 5 µM GSK-J4 for 3 days during differentiation and S100a8 and S100a9 mRNA levels were analysed using qRT-PCR (n=5). Chromatin immunoprecipitation was performed using anti-H3K27me3 and appropriate IgG control antibodies in chromatin of vehicle controls (-) and treated (+GSK-J4) WT and C/EBPδ KO (KO) ER-Hoxb8 monocytes. Purified DNA was analysed using primer pairs flanking different S100a8 (C) and S100a9 (D) promoter regions (n=3–5). Values are the means ± SEM. *p<0.05, **p<0.01, ***p<0.001, ns = not significant, by one-way ANOVA with Bonferroni’s correction (A, B) or by two-tailed Student’s t test in comparison to WT (-) (C, D).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| gene (Mus musculus) | S100a8 | NCBI | Gene ID:20201 | |
| gene (Mus musculus) | S100a9 | NCBI | Gene ID:20202 | |
| Strain, strain background (Escherichia coli) | Subcloning Efficiency DH5α Competent Cells | Thermo Fisher Scientific | Cat# 18265017 | |
| Strain, strain background (Staphylococcus aureus) | pHrodo Red S. aureus Bioparticles Conjugate | Thermo Fisher Scientific | Cat# A10010 | 5 × 106 per 0.5 × 106 cells |
| Genetic reagent (Mus musculus) | Cebpdtm1Pfj/Cebpdtm1Pfj | PMID:9724803 | RRID: MGI:3606309 | Esta Sterneck (CCR, NIH, MD) |
| Genetic reagent (Mus musculus) | B6J.129(Cg)-Igs2tm1.1(CAG-cas9*)Mmw/J | Jackson Laboratory | Stock# 028239 RRID:IMSR_JAX:028239 | PMID:26178787 |
| Cell line (Mus musculus) | WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Cell line (Mus musculus) | Cas9-expressing ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Cell line (Mus musculus) | C/EBPδ KO ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Cell line (Homo sapiens) | HEK293T | ATCC | RRID:CVCL_0063 | |
| Transfected construct (synthetic) | Cas9+PHF8 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+CSRP1 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+HAND1 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+FBXW7 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+ATF3 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+STAT3 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+KLF5 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+IRF7 gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | Cas9+C/EBPβ gRNA in WT ER-Hoxb8 cells | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a8 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a8 s1 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a8 s2 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a8 s3 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a8 s2+3 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a9 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a9 s1 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a9 s2 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a9 s3 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | MSCV S100a9 s4 prom_GFP in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | TRE_3xFlag-C/EBPδ in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Transfected construct (synthetic) | TRE_ctrl in HEK293T | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Biological sample (Homo sapiens) | Patient-derived PBMCs | PMID:35379829 | N/A | BioNRW Study |
| Antibody | Anti-mouse S100A8 (rabbit polyclonal) | PMID:25098555 | N/A | ELISA (1:460) WB (1:500) |
| Antibody | Anti-mouse S100A9 (rabbit polyclonal) | PMID:25098555 | N/A | ELISA (1:2300) WB (1:1000) |
| Antibody | Anti-mouse S100A9-FITC (rabbit polyclonal) | This paper | N/A | FACS (5 µg/ml) |
| Antibody | Anti-mouse Ly-6B.2 Alloantigen (7/4) (rat monoclonal) | BioRad | Cat# MCA771F, RRID: AB_322951 | Flow cytometry (1:200) |
| Antibody | Anti-mouse Gr-1 (RB6-8C5) from hybridoma cells (recombinant monoclonal) | This paper | N/A | Flow cytometry (1:200) AG Zarbock (Department of Anesthesiology, UKM) |
| Antibody | Anti-mouse CD11b-Pb (M1/70) (rat monoclonal) | BioLegend | Cat# 101224, RRID: AB_755986 | FACS (1:100) Flow cytometry (1:200) |
| Antibody | Anti-mouse Ly-6C-PE (HK1.4) (rat monoclonal) | BioLegend | Cat# 128008, RRID: AB_1186132 | FACS (1:100) Flow cytometry (1:200) |
| Antibody | Anti-IgG2b, κ Isotype control- Pb(RTK4530) (rat monoclonal) | BioLegend | Cat# 400627, RRID: AB_493561 | Flow cytometry (1:200) |
| Antibody | Anti-IgG2c, κ Isotype control-PE (RTK4174) (rat monoclonal) | BioLegend | Cat# 400707, RRID: AB_326573 | Flow cytometry (1:200) |
| Antibody | Anti-GAPDH (14 C10) (rabbit monoclonal) | Cell Signaling Technology | Cat# 2118, RRID: AB_561053 | WB (1:1000) |
| Antibody | Anti-alpha/beta-Tubulin (rabbit polyclonal) | Cell Signaling Technology | Cat# 2148, RRID: AB_2288042 | WB (1:1000) |
| Antibody | Anti-H3K27me3 (C36B11) (rabbit monoclonal) | Cell Signaling Technology | Cat# 9733, RRID: AB_2616029 | ChIP(3 µg) |
| Antibody | Normal IgG control (rabbit polyclonal) | Cell Signaling Technology | Cat# 2729, RRID: AB_1031062 | ChIP(3 µg) |
| Antibody | Anti-H3K27ac (rabbit polyclonal) | Abcam | Cat# ab4729, RRID: AB_2118291 | ChIP(3 µg) |
| Antibody | Anti-FLAG M2 (mouse monoclonal) | Sigma-Aldrich | Cat# F1804, RRID: AB_262044 | ChIP(3 µg) WB (1:1500) |
| Antibody | IgG1 Isotype control (MOPC-21) (mouse monoclonal) | BD Biosciences | Cat# 554121, RRID: AB_395252 | ChIP(3 µg) |
| Antibody | Anti-Rabbit Immunoglobulins/HRP (goat polyclonal) | Agilent | Cat# P0448, RRID: AB_2617138 | WB (1:4000) |
| Antibody | Anti-Mouse Immunoglobulins/ HRP (rabbit polyclonal) | Agilent | Cat# P0260, RRID: AB_2636929 | WB (1:2000) |
| Antibody | Anti-CD2-PE (RPA-2.10) (mouse monoclonal) | BD Biosciences | Cat# 555327, RRID: AB_395734 | FACS (1:20) |
| Antibody | Anti-CD14-APC (M5E2) (mouse monoclonal) | BD Biosciences | Cat# 555399, RRID: AB_398596 | FACS (1:20) |
| Antibody | Anti-CD15-PE (HI98) (mouse monoclonal) | BD Biosciences | Cat# 555402, RRID: AB_395802 | FACS (1:5) |
| Antibody | Anti-CD19-PE (HIB19) (mouse monoclonal) | BD Biosciences | Cat# 555413, RRID: AB_395813 | FACS (1:20) |
| Antibody | Anti-CD16-PE-Cy7 (3 G8) (mouse monoclonal) | BD Biosciences | Cat# 557744, RRID: AB_396850 | FACS (1:20) |
| Antibody | Anti-CD56-PE (MY31) (mouse monoclonal) | BD Biosciences | Cat# 556647, RRID: AB_396511 | FACS (1:5) |
| Antibody | Anti-CD335-PE (9E2/NKp46) (mouse monoclonal) | BD Biosciences | Cat# 557991, RRID: AB_396974 | FACS (1:20) |
| Antibody | Anti-HLA-DR-FITC (TU36) (mouse monoclonal) | BD Biosciences | Cat# 555560, RRID: AB_395942 | FACS (1:5) |
| Recombinant DNA reagent | pCW57.1 mDux-CA | PMID:28459454 | RRID: Addgene_99284 | All-in-one lentivector with tet-inducible mouse Dux |
| Recombinant DNA reagent | pcDNA 3.1(-) mouse C/EBP delta | Johnson Laboratory | RRID: Addgene_12559 | Mouse C/EBPδ cDNA |
| Recombinant DNA reagent | TRE_3xFlag-C/EBPδ | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | TRE_ctrl | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiGuide-puro for GeCKO screen | PMID:25075903 | RRID: Addgene_1000000053 | Lentiviral pooled libraries for genome-wide CRISPR/Cas9 Knock-outs |
| Recombinant DNA reagent | lentiCRISPRv2 | PMID:25075903 | RRID: Addgene_52961 | Lentivector with CRISPR/Cas9 expression |
| Recombinant DNA reagent | lentiCRISPRv2-PHF8 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-CSRP1 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-HAND1 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-FBXW7 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-ATF3 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-STAT3 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-KLF5 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-IRF7 gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | lentiCRISPRv2-C/EBPβ gRNA | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | pLenti CMV GFP Blast (659-1) | PMID:19657394 | RRID: Addgene_17445 | Lentiviral eGFP expression vector |
| Recombinant DNA reagent | MSCV-PIG-empty | PMID:29316427 | RRID: Addgene_105594 | Empty control vector with GFP marker and puro resistance |
| Recombinant DNA reagent | MSCV S100a8 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a8 s1 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a8 s2 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a8 s3 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a8 s2+3 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a9 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a9 s1 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a9 s2 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a9 s3 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | MSCV S100a9 s4 prom_GFP | This paper | N/A | AG Roth (Institute of Immunology, WWU Münster) |
| Recombinant DNA reagent | psPAX2 | Trono Laboratory | RRID: Addgene_12260 | Lentiviral packaging plasmid |
| Recombinant DNA reagent | pCMV-VSV-G | Weinberg Laboratory | RRID: Addgene_8454 | Envelope protein for producing lentiviral particles |
| Sequence-based reagent | Cloning primers | This paper | Listed in Supplementary file 1a, c, d | |
| Sequence-based reagent | GeCKO library amplification and NGS primers | This paper | Listed in Supplementary file 1b | |
| Sequence-based reagent | Mutagenesis primers | This paper | Listed in Supplementary file 1e | |
| Sequence-based reagent | qPCR primers | This paper | Listed in Supplementary file 1f, g | |
| Peptide, recombinant protein | rm mouse GM-CSF | ImmunoTools | Cat# 12343137 | 40 ng/ml in vitro |
| Peptide, recombinant protein | LPS from Salmonella enteritidis | Sigma-Aldrich | Cat# L6011 | 0.5 mg/ml in vivo |
| Peptide, recombinant protein | LPS from Escherichia coli O55:B5 | Sigma-Aldrich | Cat# L2880 | 10 ng/ml in vitro |
| Peptide, recombinant protein | rm mouse IL-4 | Peprotech | Cat# 214-14 | 20 ng/ml in vitro |
| Peptide, recombinant protein | rm mouse IFN-y | ImmunoTools | Cat# 12343536 | 50 ng/ml in vitro |
| Commercial assay or kit | NucleoSpin RNA, Mini kit for RNA purification | Macherey Nagel | Cat# 740955.50 | |
| Commercial assay or kit | QuikChange II XL Site-Directed Mutagenesis Kit | Agilent Technologies | Cat# 200521 | |
| Commercial assay or kit | PureLink HiPure Plasmid Miniprep Kit | Thermo Fisher Scientific | Cat# K210002 | |
| Commercial assay or kit | NEBNext High-Fidelity 2× PCR Master Mix | NEB | Cat# M0541S | |
| Commercial assay or kit | QIAquick PCR Purification Kit | Qiagen | Cat# 28104 | |
| Commercial assay or kit | Bioanalyzer High Sensitivity DNA Analysis Agilent High Sensitivity DNA Kit | Agilent | Cat# 5067-4626 | |
| Commercial assay or kit | 20% PhiX spike in (Illumina PhiX control kit) | Illumina | Cat# FC-110-3001 | |
| Commercial assay or kit | NextSeq 500/550 High Output Kit v2.5 (75 Cycles) | Illumina | Cat# 20024906 | |
| Commercial assay or kit | NEBNext Ultra RNA Library Prep Kit for Illumina | NEB | Cat# E7530 | |
| Commercial assay or kit | LEGENDplex Mouse Inflammation Panel (13-plex) | BioLegend | Cat# 740446 | |
| Chemical compound, drug | Doxycycline | Sigma-Aldrich | Cat# D9891 | 2 µg/ml |
| Chemical compound, drug | Puromycin | InvivoGen | Cat# ant-pr-1 | 3–20 µg/ml |
| Chemical compound, drug | GSK-J4 HCl | SellekChem | Cat# S7070 | 5 µM |
| Chemical compound, drug | β-Estradiol | Sigma-Aldrich | Cat# E8875 | 1 µM |
| Chemical compound, drug | Polybrene | Sigma-Aldrich | Cat# 107689 | 8 µg/ml |
| Chemical compound, drug | Protease Inhibitor Cocktail | Sigma-Aldrich | Cat# P8340 | 1:1000 |
| Chemical compound, drug | Dynabead Protein G for Immunoprecipitation | Thermo Fisher Scientific | Cat# 10004D | ChIP (1:10) |
| Chemical compound, drug | M-PER Mammalian Protein Extraction Reagent | Thermo Fisher Scientific | Cat# 78501 | 1× |
| Chemical compound, drug | Dihydrorhodamine 123 | Sigma-Aldrich | Cat# D1054 | 15 µm |
| Chemical compound, drug | NEBNext Poly(A) mRNA Magnetic Isolation Module | NEB | Cat# E7490 | |
| Chemical compound, drug | DNA Polymerase I, Large (Klenow) Fragment | NEB | Cat# M0210 | |
| Chemical compound, drug | FluoSpheres Polystyrene Microspheres, 1.0 µm, blue-green fluorescent (430/465) | Thermo Fisher Scientific | Cat# F13080 | 5 × 106 per 0.5 × 106 cells |
| Software, algorithm | AliBaba2.1 | PMID:11808873 | http://gene-regulation.com/pub/programs/alibaba2/ | |
| Software, algorithm | FlowJo Software | BD Biosciences | https://www.flowjo.com/solutions/flowjo/ | |
| Software, algorithm | MaGeCK 0.5.9.3 | PMID:25476604 | https://sourceforge.net/p/mageck/wiki/Home/ |
Additional files
-
Supplementary file 1
List of oligonucleotides used for cloning, (qRT)PCR and mutagenesis.
(a) List of guides (stated in 5’–3’ orientation) for cloning into lentiCRISPR v2, related to Materials and methods. (b) List of primer (stated in 5’–3’ orientation) for amplifying GeCKO library and NGS, related to Materials and methods. (c) List of oligonucleotides (stated in 5’–3’ orientation, fw: forward, rv: reverse) for cloning steps to construct TRE_3xFlag-C/EBPδ vector, related to Materials and methods. (d) List of oligonucleotides (stated in 5’–3’ orientation, fw: forward, rv: reverse) for cloning steps to construct S100a8 and S100a9 reporter construct, related to Materials and methods. (e) List of oligonucleotides (stated in 5’–3’ orientation) for mutagenesis to disrupt specific sites within S100a8 and S100a9 reporter vectors, related to Materials and methods. (f) List of quantitative reverse transcription polymerase chain reaction (qRT-PCR) primer (in 5’–3’ orientation) used for qRT-PCR, related to Materials and methods. (g) List of chromatin immunoprecipitation (ChIP)-PCR primer (in 5’–3’ orientation) for S100a8 and S100a9 genomic locations, related to Materials and methods.
- https://cdn.elifesciences.org/articles/75594/elife-75594-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/75594/elife-75594-transrepform1-v2.docx





