Structure of the mitoribosomal small subunit with streptomycin reveals Fe-S clusters and physiological molecules
Figures
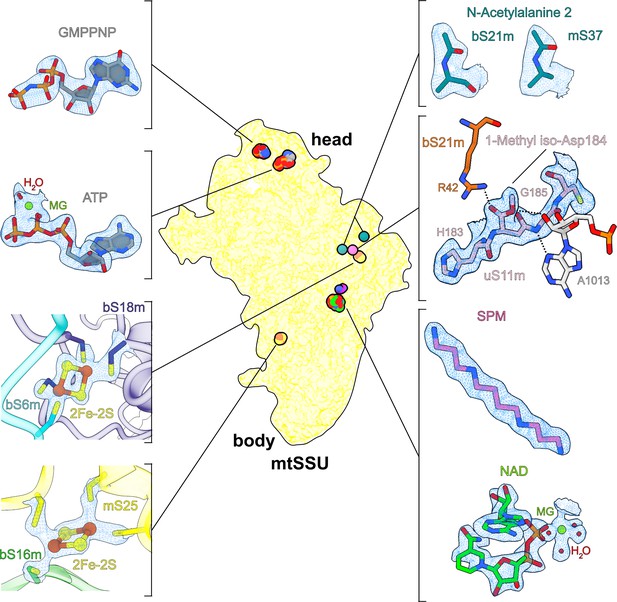
Overview of cofactors and modifications of the mitoribosomal small subunit.
Outlined view indicates the relative positions of identified cofactors and modifications. Cryo-EM densities, models, and surrounding environment are shown for GMPPNP, adenosine triphosphate (ATP), 2Fe–2S clusters, protein modifications, spermine, and nicotinamide adenine dinucleotide (NAD).
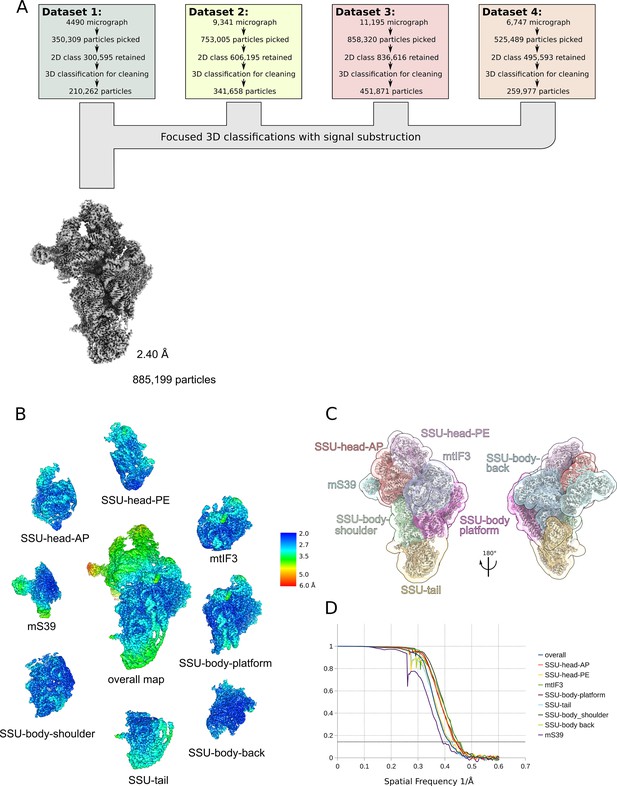
Cryo-EM data collection and processing.
(A) Cryo-EM data processing overview. (B) Binary masks used for local-masked refinements. (C) Density maps from overall refinements, filtered by their nominal resolution without sharpening, colored by local resolution. (D) Fourier shell correlation (FSC) curves of half maps from overall refinement and the local-masked refinements.
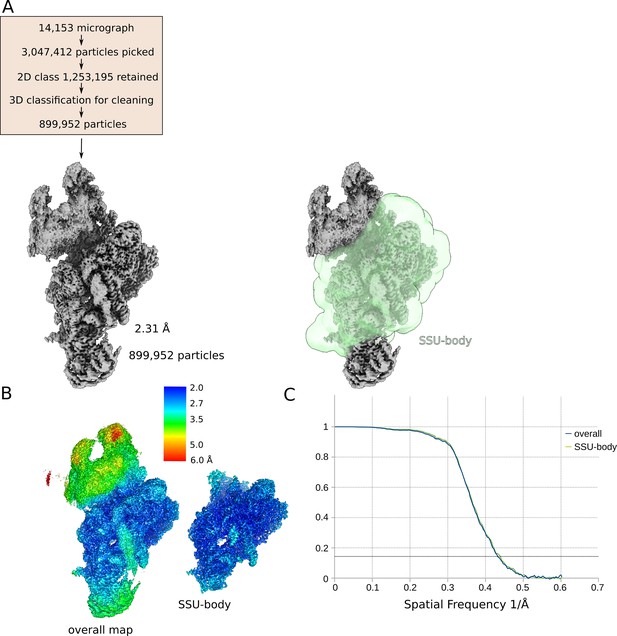
Cryo-EM data collection and processing for in vitro reconstituted small subunit (SSU):streptomycin complex.
(A) Cryo-EM data processing overview, final consensus map, and binary mask on SSU-body used for local-masked refinement. (B) Density maps from overall refinements, filtered by their nominal resolution without sharpening, colored by local resolution. (C) Fourier shell correlation (FSC) curves of half maps from overall refinement and the local-masked refinement.
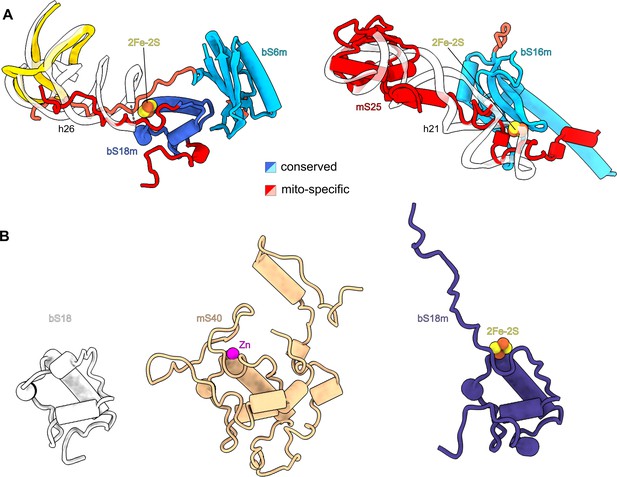
Iron–sulfur clusters bridge mitoribosomal proteins.
(A) Each of the two 2Fe–2S clusters is coordinated by mitochondria-specific components (red) from two different proteins. Superposition of 2Fe–2S cluster-binding sites in the human mitoribosome with corresponding sites in E. coli ribosome (PDB ID: 7K00, white) reveals that mitoribosomal protein elements participating in 2Fe–2S coordination compensate for missing rRNA helices. (B) The conserved overall fold of bS18 contributes to Zn2+ or 2Fe–2S binding in the mitoribosome.
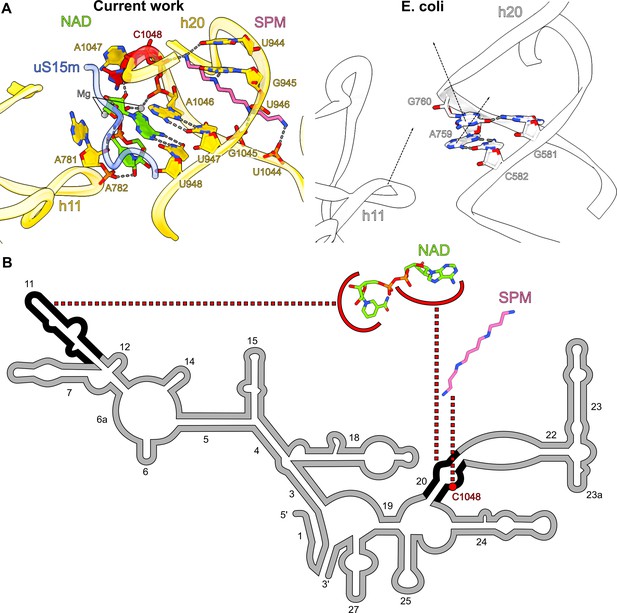
Spermine-NAD functionally compensates for rRNA and protein alterations.
(A) Comparison of the spermine-NAD binding site with E. coli ribosome (PDB ID: 7K00, white) shows that it provides structural stability that compensates for rRNA alterations. (B) 2D diagram of the SSU head rRNA (grey) with regions stabilized by NAD and spermine indicated (black); the associated insertion C1048 is highlighted in red.
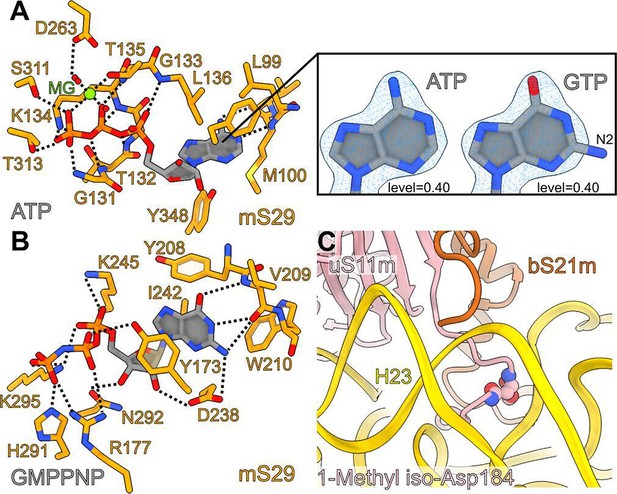
Interactions of adenosine triphosphate (ATP), GMPPNP in mS29, and 1-methyl iso-aspartate in uS11m.
(A) Interactions of the bound ATP with the surrounding residues of mS29. The right panel shows the fitting of adenine versus guanine to the density, featuring the amine group N2 on carbon 6 that does not fit when guanosine triphosphate (GTP) is modeled. (B) The newly identified binding pocket and the interactions of GMPPNP with the surrounding residues of mS29. (C) Bending of uS11m loop at the site of occurrence of 1-methyl iso-Asp 184 of uS11m to facilitate interactions with helix h23.
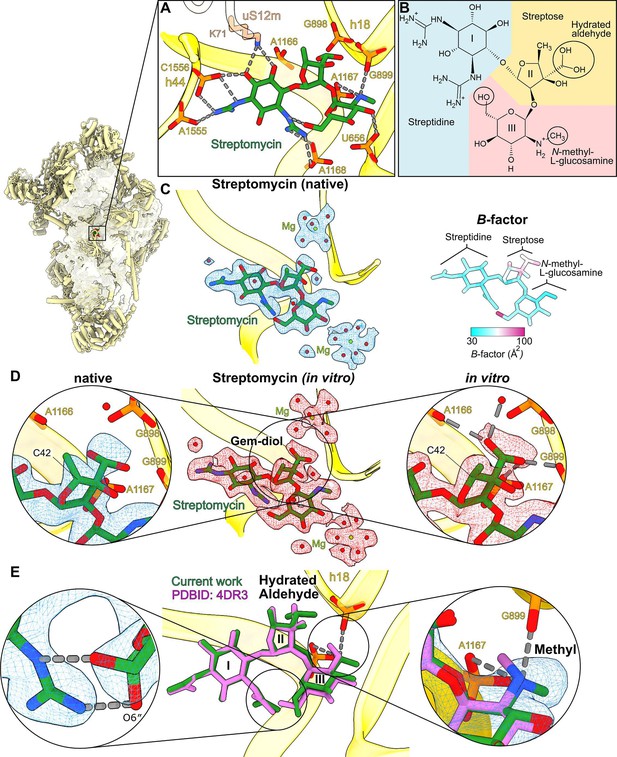
High-resolution features of streptomycin binding to the mitoribosome.
(A) Streptomycin interacts with uS12m and backbone phosphates of helices h18 and h44. (B) Chemical structure of the hydrated gem-diol form of streptomycin. (C) Left, density and model of the natively bound streptomycin along with surrounding water molecules and Mg2+ ions. Right, atomic B-factor distribution of the bound streptomycin shows higher relative flexibility of the streptose moiety. (D) Middle, density (red) and model of the in vitro formed complex. Left, zoom-in of the native state shows that the density of the gem-diol moiety is not well resolved. Right, zoom-in of the in vitro complex shows that the streptose moiety is better resolved with a defined orientation of the gem-diol moiety. A second notable difference is the inversion of chirality of C42, where the methyl group of streptose is located. (E) Comparison with the previously reported structure of T. thermophilus ribosome (PDB ID: 4DR3, pink). Close-ups of the chemical interactions of O6″ N-methyl-L-glucosamine in its alternative conformations (left zoom-in); and methyl moieties (right zoom-in) with their densities resolve discrepancies of the methyl-group orientation in the previous studies.
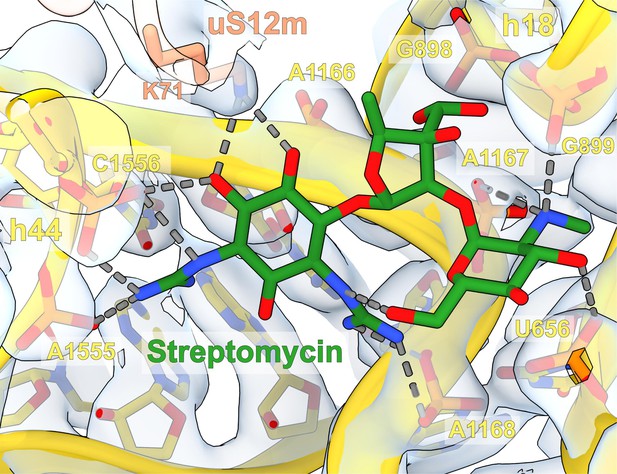
Densities of streptomycin interactions with the small subunit (SSU).
The bound streptomycin is shown as a model, and the interacting SSU region that forms the binding pocket is shown as a model with density (uS12m, h18 and h44 of rRNA). Specific interactions are indicated in dashed arrows.
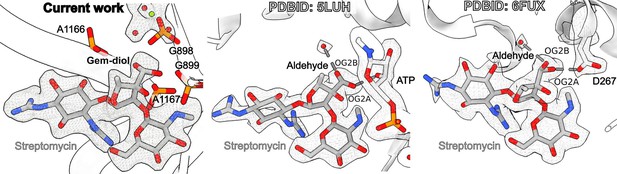
Comparison of the mitoribosome-bound streptomycin with the published X-ray electron density maps.
Left panel, current work with natively bound streptomycin. Middle panel, streptomycin modeled in two alternative conformations bound to adenylyltransferase AadA with streptose aldehyde (PDB: 5LUH, Stern et al., 2018). The density suggests a gem-diol moiety that can participate in the indicated hydrogen bonds. Right panel, streptomycin modeled in two alternative conformations bound to aminoglycoside phosphotransferase APH(3″)-Id (PDB: 6FUX, Alekseeva et al., 2019). The side chain carboxylate of D267 is likely to be a proton acceptor of a gem-diol that acts as a proton donor for the hydrogen bond.
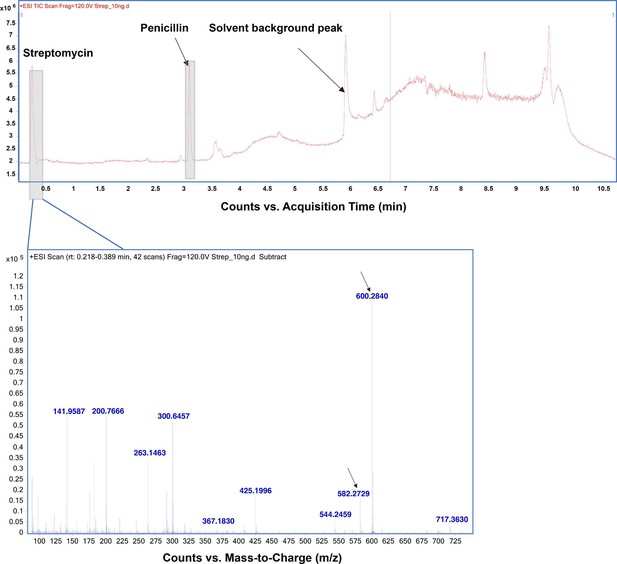
High-resolution mass spectrometry (HRMS) analysis for streptomycin.
Top, mass- spectra in positive ionization mode confirms the presence of streptomycin. Bottom, the arrows indicated corresponding m/z peaks for hydrated (600.28) and non-hydrated (582.27) forms of streptomycin.
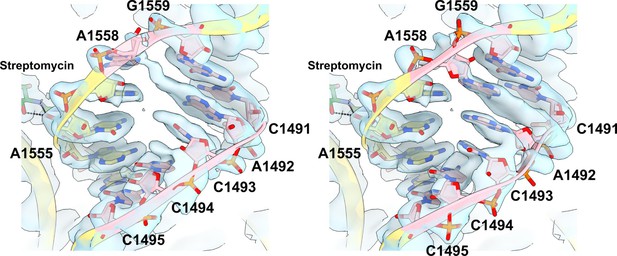
Alternative states of h44 rRNA.The density suggests that the residues colored pink in h44 adopt two alternative states (left and right).
The density shown as the surface represents a mixture of two states. A1558 is flipped out in one state (left), whereas its base is stacked between G1559 and A1492 in the other state, causing a shift of the residues in the other strand (right).
Tables
Cryo-EM collection, processing, model refinement, and validation statistics.
| Data collection and processing | Native SSU:streptomycin complex | In vitro formed SSU:streptomycin complex |
|---|---|---|
| Microscope | Titan Krios | Titan Krios |
| Detector | K2 Summit | K3 Summit |
| Magnification | 165,000 | 105,000 |
| Voltage [kV] | 300 | 300 |
| Total electron exposure [e−/Å2] | 30–32 | 40 |
| Defocus range [μm] | −0.2 to −3.6 | −0.2 to −3.7 |
| Pixel size [Å] | 0.83 | 0.846 |
| Symmetry imposed | C1 | C1 |
| Final particle | 885,199 | 899,952 |
| Resolution [Å] (overall/body/shoulder/platform/back/tail/head-AP/head-PE/mS39/mtIF3) | 2.40/–/2.23/2.28/2.29/2.41/2.26/2.39/2.57/2.20 | 2.31/2.28/–/–/–/–/–/–/– |
| Map-sharpening B-factor [Å2] (overall/shoulder/platform/back/tail/head-AP/head-PE/mS39/mtIF3) | −53/–/−47/−49/−56/−60/−52/−58/−68/−54 | −50/−49/–/–/–/–/–/–/– |
| Refinement | ||
| Model composition | ||
| Total atoms (non-hydrogen/hydrogen) | 71,880/59,406 | |
| Chains (RNA/protein) | 1/31 | |
| RNA residues (non-modified/m4C, m5C, m5U, m62A) | 950/1/1/1/2 | |
| Protein residues (non-modified/N-acetylAla/O1-methylisoAsp) | 5915/2/1 | |
| Metal ions (Mg2+/K+/Zn2+) | 62/21/1 | |
| Ligands (2Fe–2S/ATP/GMPPNP/NAD/spermine/streptomycin) | 2/1/1/1/1/1 | |
| Waters | 3087 | |
| Model to map CC (CCmask/CCbox/CCpeaks/CCvolume) | 0.90/0.79/0.79/0.88 | |
| Resolution [Å] by model-to-map FSC, threshold 0.50 (masked/unmasked) | 2.18/2.18 | |
| Average B-factor [Å2] (RNA/protein/metal ion and ligand/water) | 36/48/32/32 | |
| R.m.s. deviations, bond lengths [Å]/bond angles [°] | 0.002/0.408 | |
| Validation | ||
| Clash score | 1.39 | |
| Rotamer outliers [%] | 0 | |
| Ramachandran plot [%] (favored/ allowed/disallowed) | 98.01/1.95/0.03 | |
| CaBLAM outliers [%] | 0.71 | |
| Cβ outliers [%] | 0 | |
| MolProbity score | 0.87 | |
| EMRinger score | 6.16 | |
| PDB/EMDB accession code | 7P2E/EMD-13170 | –/EMD-15542 |





