Community diversity is associated with intra-species genetic diversity and gene loss in the human gut microbiome
Figures
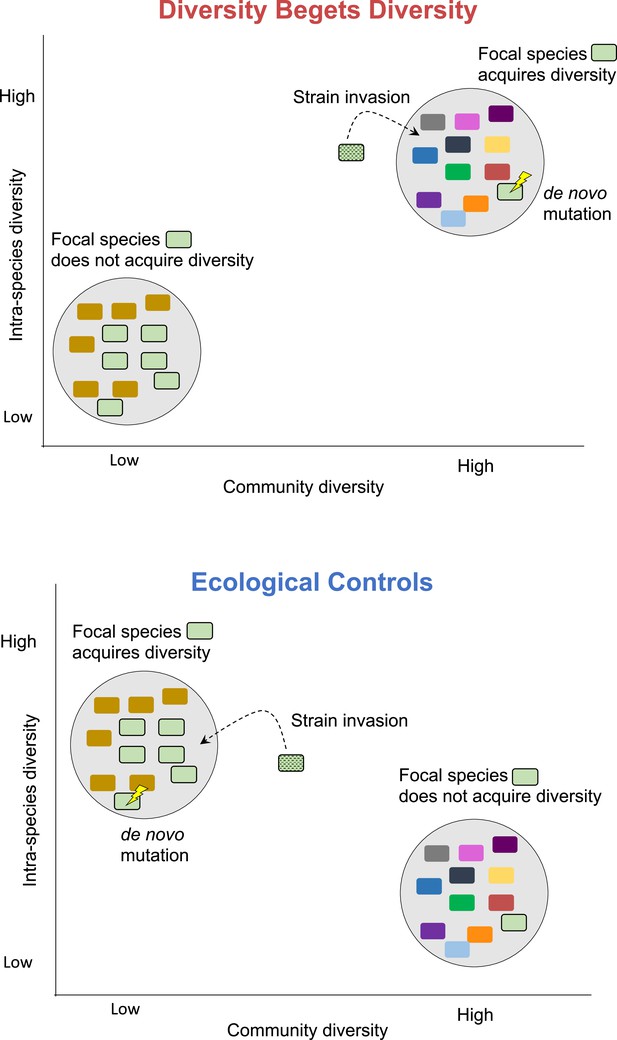
Diversity begets diversity (DBD) and ecological controls (EC) hypotheses illustrated.
Hypothetical microbial communities are illustrated as gray circles containing assemblages of microbial species, shown in different colors. 'DBD' means that the focal species is more likely to acquire diversity – through de novo mutation, invasion of a different strain of the same species, or a combination of both – in a community with high diversity. This is because new niches are created in a more diverse community. By contrast, 'EC' means that the focal species is more likely to acquire diversity through strain invasion or mutation in a community with low diversity. This is because niches remain unfilled in a low-diversity community, while niche space is saturated in a high-diversity community, impeding further diversification.
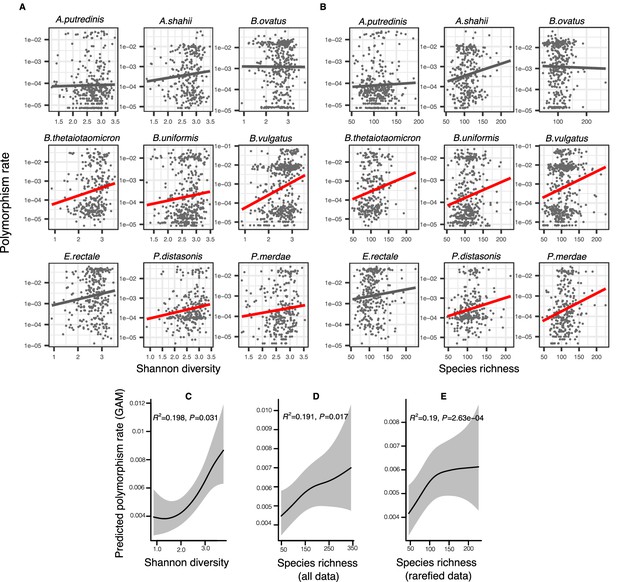
Positive association between community diversity and within-species polymorphism in cross-sectional Human Microbiome Project (HMP) samples.
(A) Scatter plots showing the relationship between community Shannon diversity and within-species polymorphism rate (estimated at synonymous sites) in the nine most prevalent species in HMP. (B) Scatter plots showing the relationship between species richness and within-species polymorphism rate in the nine most prevalent species in HMP. These are simple correlations to show the relationships in the raw data. Significant correlations are shown with red trendlines (Spearman correlation, p<0.05); non-significant trendlines are in gray. Results of generalized additive models (GAMs) predicting polymorphism rate in a focal species as a function of (C) Shannon diversity, (D) species richness estimated on all sequence data, and (E) species richness estimated on rarefied sequence data. GAMs are based on data from 69 bacterial species across 249 HMP stool donors. Adjusted R2 and Chi-square p-values corresponding to the predictor effect are displayed in each panel. Shaded areas show the 95% confidence interval of each model prediction. See Supplementary file 1a and Supplementary file 2 section 1 for detailed model outputs.
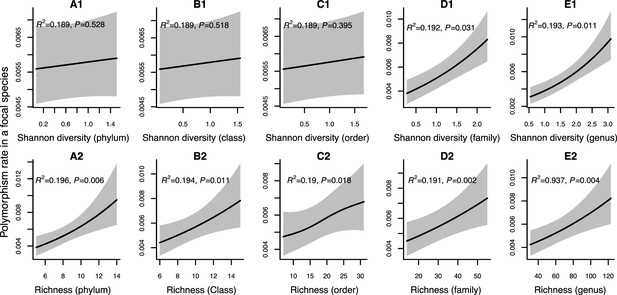
Results of generalized additive models predicting within-species polymorphism rate (at synonymous sites) as a function of community diversity at higher taxonomic levels (Human Microbiome Project [HMP] data).
(A1–E1) The predictor is Shannon diversity. (A2–E2) The predictor is richness. Adjusted R-squared (R2) and Chi-squared p-values corresponding to the predictor are displayed in each panel (gam.summary function from mgcv R package). Shaded areas show the 95% confidence interval of each model prediction. See Supplementary file 1c and Supplementary file 2 sections 2 and 3 for further details about model outputs.
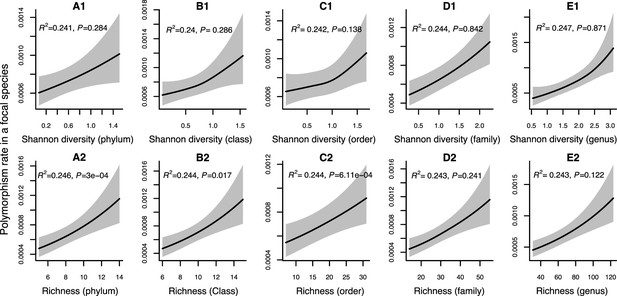
Results of generalized additive models predicting within-species polymorphism rate (at nonsynonymous sites) in a focal species as a function of community diversity at higher taxonomic levels (Human Microbiome Project [HMP] data).
(A1–E1) The predictor is Shannon diversity. (A2–E2) The predictor is richness. Adjusted R-squared (R2) and Chi-squared p-values corresponding to the predictor are displayed in each panel (gam.summary function from mgcv R package). Shaded areas show the 95% confidence interval of each model prediction. See Supplementary file 1d and Supplementary file 2 sections 5 and 6 for further details about model outputs.
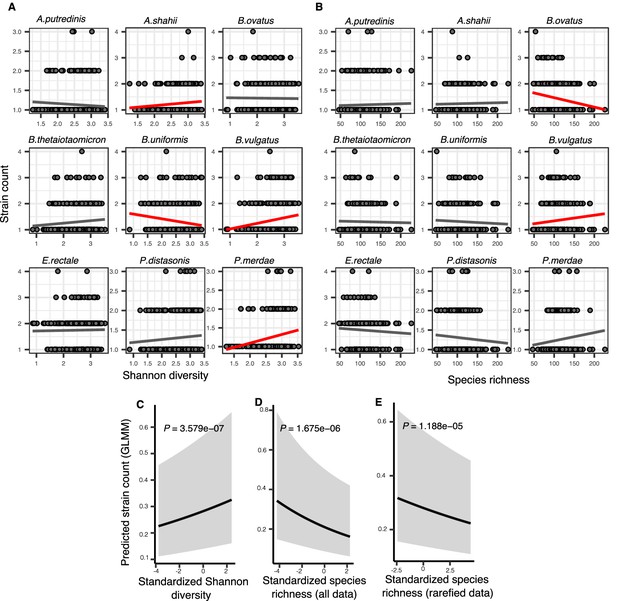
Associations between community diversity and strain number in cross-sectional Human Microbiome Project (HMP) samples.
(A) Scatter plots showing the relationship between Shannon diversity and the inferred number of strains within each of the nine most prevalent species in HMP. (B) Scatter plots showing the relationship between species richness and the inferred number of strains within each of the nine most prevalent species in HMP. Significant linear correlations are shown with red trendlines (Pearson correlation, p<0.05); non-significant trend lines are in gray. Results of generalized linear mixed models (GLMMs) predicting strain count in a focal species as a function of (C) Shannon diversity, (D) species richness estimated on all data, and (E) species richness estimated on rarefied sequence data. Diversity estimates (X-axis) are standardized to zero mean and unit variance in the models. The Y-axis shows the mean number of strains per focal species predicted by the GLMM. GLMMs are based on data from 184 bacterial species across 249 HMP stool donors. p-Values (likelihood ratio test) are displayed in each panel. Shaded areas show the 95% confidence interval of each model prediction. See Supplementary file 1e and Supplementary file 2 section 7 for detailed model outputs.
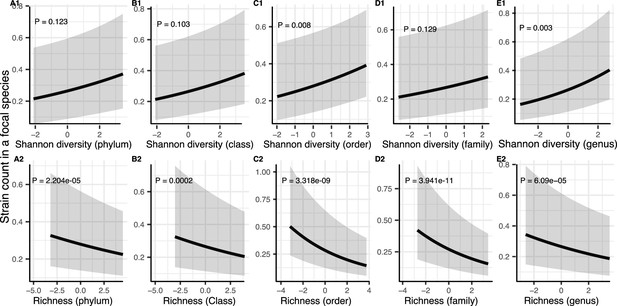
Results of generalized linear mixed models predicting strain count in a focal species as a function of community diversity at higher taxonomic levels (Human Microbiome Project [HMP] data).
Strain number in a focal species is positively correlated with Shannon (A1–E1) whereas its correlation with richness remains negative (A2–E2) through all taxonomic levels. The Y-axis is the predicted mean number of strains within a focal species. p-Values (drop1 function from R stats package, likelihood ratio test [LRT]). Shaded areas show the 95% confidence interval of each model prediction. See Supplementary file 1f and Supplementary file 2 section 9 for model details.
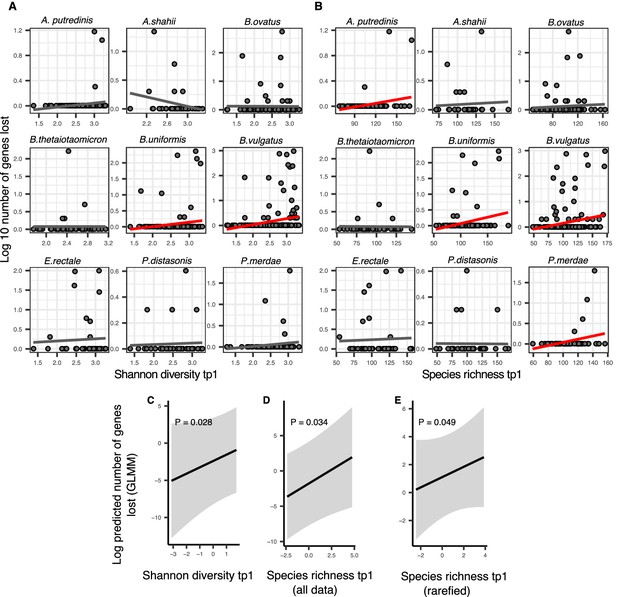
Positive association between community diversity and gene loss in Human Microbiome Project (HMP) time series.
(A) Scatter plots showing the relationship between Shannon diversity at time point 1 (tp1) and gene loss between tp1 and tp2 within each of the nine most prevalent species in HMP. (B) Scatter plots showing the relationship between species richness at tp1 and gene loss between tp1 and tp2 within each of the nine most prevalent species in HMP. Significant linear correlations are shown with red trendlines (Pearson correlation, p<0.05); non-significant trend lines are in gray. The Y-axis is plotted on a log10 scale for clarity. Results of generalized linear mixed models (GLMMs) predicting gene loss in a focal species as a function of (C) Shannon diversity, (D) species richness estimated on all data, and (E) species richness estimated on rarefied sequence data. p-Values (likelihood ratio test) are displayed in each panel. Shaded areas show the 95% confidence interval of each model prediction. The Y-axis is plotted on the link scale, which corresponds to log for negative binomial GLMMs with a count response. GLMMs are based on data from 54 bacterial species across 154 HMP stool donors sampled at more than one time point. See Supplementary file 1g and Supplementary file 2 section 10 for detailed model outputs.
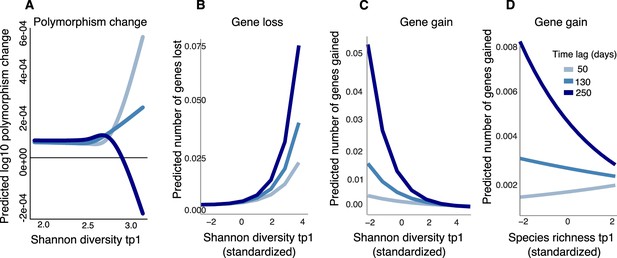
Community diversity is associated with increases in focal species polymorphism over short time lags and net gene loss in dense gut microbiome time series.
(A) Results of a generalized additive model (GAM) predicting polymorphism change in a focal species as a function of the interaction between Shannon diversity at the first time point and the time lag (days) between two time points in data from Poyet et al. The response (Y-axis) was log-transformed in the Gaussian GAM. Results of generalized linear mixed models (GLMMs) predicting (B) number of genes lost and (C) number of genes gained between two time points in a focal species as a function of the interaction between Shannon diversity at the first time point and the time lag between the two time points. (D) Results of the GLMM predicting the number of genes gained in a focal species as a function of the interaction between rarefied species richness at the first time point and the time lag between the two time points. The illustrated time lags correspond to the first quartile (50 days), the median (130 days), and the third quartile (250 days). See Supplementary file 1h and i and Supplementary file 2 section 11 for detailed model outputs. These analyses are based on data from 15 bacterial species across four stool donors from Poyet et al. Only statistically significant relationships are plotted. Non-significant relationships are not shown: the GAM predicting polymorphism change as a function of rarefied richness (p>0.05) and the GLMM predicting the number of genes lost as a function of rarefied richness (p>0.05).
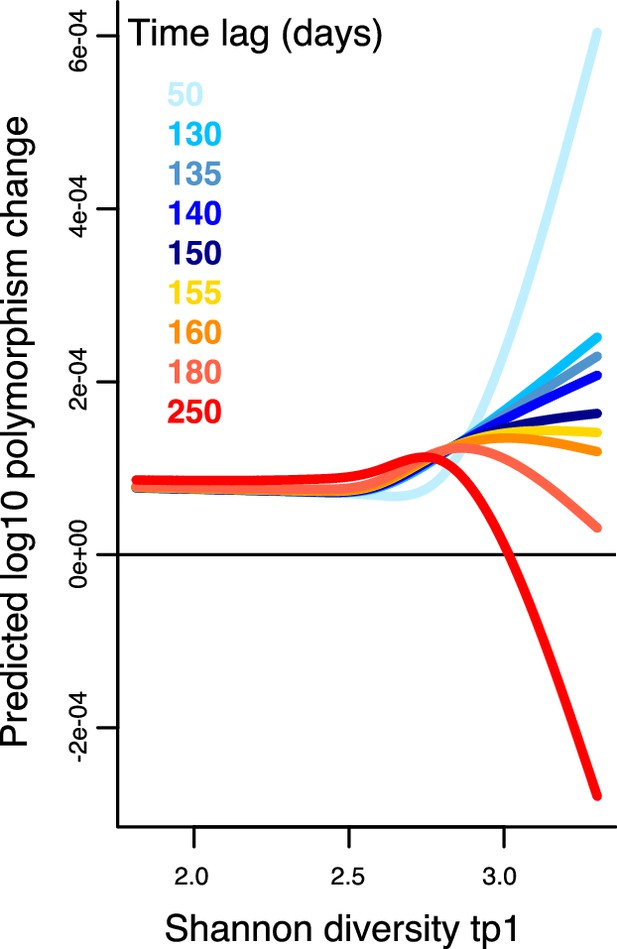
Results of a generalized additive model (GAM) predicting polymorphism change in a focal species as a function of the interaction between Shannon diversity at the first time point and the time lag (days) between two time points in the Poyet time series.
The response (Y-axis) was log-transformed in the Gaussian GAM. Several different time lags are shown to illustrate the inversion of the relationship around a lag time of 150 days. See Supplementary file 1h and Supplementary file 2 section 11 for further model details.
Additional files
-
Supplementary file 1
Supplementary tables a–i.
- https://cdn.elifesciences.org/articles/78530/elife-78530-supp1-v2.docx
-
Supplementary file 2
R syntax and statistics of all generalized models.
- https://cdn.elifesciences.org/articles/78530/elife-78530-supp2-v2.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/78530/elife-78530-mdarchecklist1-v2.docx





