Generating active T1 transitions through mechanochemical feedback
Figures
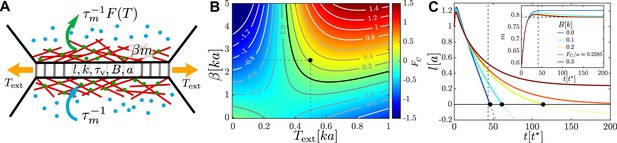
An active junction.
(A) An external pulling force of magnitude induces tension in a cell–cell junction of length , which consists of passive viscoelastic and active components. The passive component consists of a Maxwell element with stiffness and viscous relaxation time and a harmonic spring of stiffness and rest length connected to it in parallel. The active component is due to myosin motors (green and blue dots) with concentration that act to contract cortical actin filaments (red lines), exerting a force of magnitude . Myosin motors bind to the actin cortex with association rate and unbind with a tension-dependent dissociation rate . (B) Heatmap plot of the contraction force . For , the junction contraction rate is , where is the friction coefficient with the surrounding medium. The mechanochemical feedback loop is contractile in the top-right quadrant where , , and . Negative values of correspond to an extending junction. (C) Junction length vs. time for , , (black dot in B) for increasing values of the elastic barrier . An active corresponds to reaching . Increasing slows down contractions, until, for , the equilibrium length and no T1 is possible. Inset: myosin dynamics for the same set of junctions; the horizontal dashed line indicates , , , and . Length is measured in units of , time in units of , and force in units of .
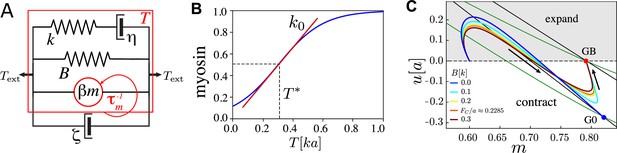
Key ingredients of the single active junction model.
(A) The junction is modelled as a Maxwell element with stiffness and viscosity (and the relaxation timescale ) connected in parallel with the active forcing, and an elastic barrier with stiffness and rest length . It is coupled in parallel to a dashpot with viscosity modelling dissipation with the surrounding medium. The red arrow indicates the feedback loop between tension and myosin kinetics. (B) Myosin–tension response curve, for with , , , . (C) Trajectories in the plane for several values of , where . G0 (GB) is the fixed point with no () elastic barrier. Fixed points are located at the intersection of their respective (black) and (green) nullclines. Arrows indicate the direction of motion. , , , and . Length is measured in units of , time in units , and force in units of .
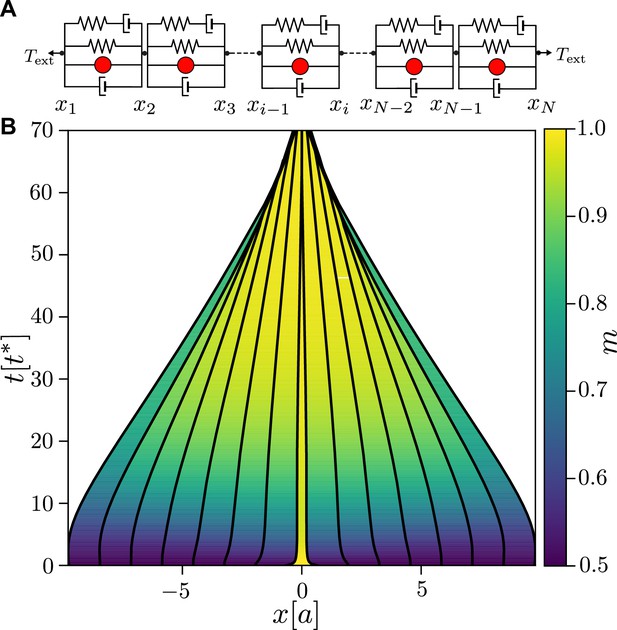
A linear chain of active junctions.
(A) Schematic of a linear chain for viscoelastic junctions with active tension feedback. The chain is formed by connecting in series the active viscoelastic elements shown in Figure 1—figure supplement 1A, with the pulling force of magnitude applied to both ends. (B) Time evolution of a linear chain for and with an initial pulse of myosin. The central contraction propagates rapidly outwards, resulting in a contraction and collapse of the chain. Activity and pulling force correspond to a point on the contraction–extension separatrix in Figure 1B, putting the system in a mechanically neutral state. The remaining parameters are the same as in Figure 1C.
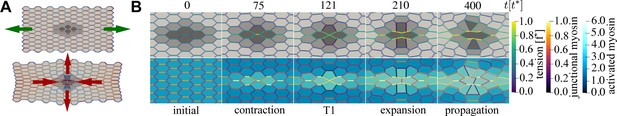
An active T1 transition event.
(A) Top panel: the mechanical anisotropy in the initial state is produced by applying pulling forces (green arrows) in the horizontal direction to the left and right boundaries. Bottom panel: the final state after the active T1 shows a clear convergence–extension deformation (red arrows). Cells are coloured by type: passive (light grey), buffer (medium grey), and active (dark grey). Junctions are coloured by junctional myosin. (B) Time sequence of the active T1 transition measured from the moment activity and viscoelasticity were switched on. Cells in the top row are coloured by type and junctions are coloured by tension. Cells in the bottom row are coloured by activated myosin , and junctions are coloured by myosin. Parameters: , , (active), (buffer), (passive), , , , , , , , , with () cells in the horizontal (vertical) direction. Units: length (), time (), force ().
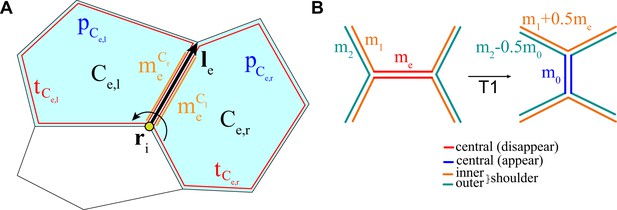
Schematic representation of the key ingredients in the vertex model with active junctions.
(A) The total force on vertex i can be expressed as a sum of forces due to junctions. For the junction , and are the cells to the left and the right, respectively, when looking in the direction of the vector is the myosin attached to junction e due to the cell to the left (right) of it. The area () and perimeter () terms are defined in the text above. (B) Example of the redistribution of myosin after a T1 transition for one side of the junction, and one of its inner and outer shoulder junctions.
Simulation of a passive system with applied horizontal pulling for .
Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). In this video activity is turned off, and there is no difference between those three cell types.
Low activity system with no T1 transitions.
Activity is set to for the four central cells and the patch is pulled horizontally with , , and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition.
Activity is set to for the four central cells and the patch is pulled horizontally with , , and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
System with low pulling force.
Activity is set to for the four central cells and the patch is pulled horizontally with , , and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
System with too high pulling force.
Activity is set to for the four central cells and the patch is pulled horizontally with , , and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
System with too high activity.
Activity is set to for the four central cells and the patch is pulled horizontally with , , and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition with low and .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition with low .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition with low .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition at intermediate values of and .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition at high .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition at high .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
Active T1 transition at high and .
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, cells are coloured by type, active (dark grey), buffer (intermediate grey), passive (light grey). Activity of buffer cells is set to .
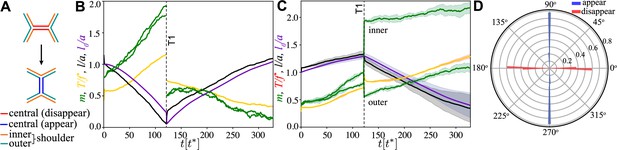
Junction dynamics during the active T1 transition shown in Figure 2.
(A) Definition of central (red/blue – junction that disappears/appears), inner (orange), and outer (blue-green) shoulder junctions through the T1 transition. (B) Central junction: myosin, (green; two curves for myosin on two sides of the junction), tension, (yellow), junction length, (black), and rest length l0 (purple) vs. time. The vertical line indicates the T1 transition, at which point junctional myosin is redistributed according to the rules outlined in Figure 2—figure supplement 1B. (C) Same as in panel (A) but averaged over four shoulder junctions, with variance indicated as shade. (D) Polar histogram of the orientation of the first T1 event measured with respect to the pulling direction, from simulations. Blue (red) indicates appearing (disappearing) junctions. Parameters are and and as in Figure 2.
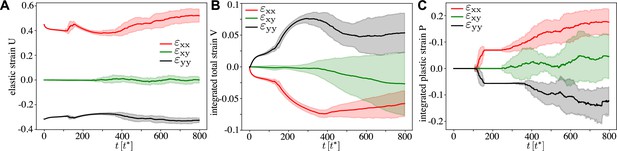
Continuous strain tensors through the active T1 transition, for , , and the other parameters corresponding to Figure 2 and Figure 3.
(A) Statistical strain tensor defined in Equation 20, with the initial undeformed state used as reference configuration. (B) Total strain, that is, integrated tensor defined in Equation 24. (C) Integrated plastic strain tensor defined in Equation 22. One can see that the active T1 proceeds at near constant elastic strain with cells elongated in the pulling direction, that is, and . Conversely, there is clear convergence–extension since and in the total strain. Here, the traces are an average over simulations, and the shading corresponds to the error of the mean.
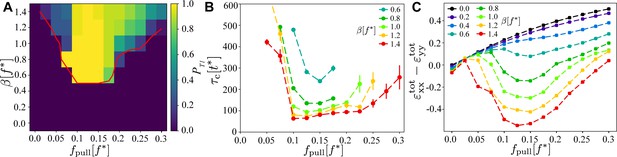
Existence and timescales of T1 transitions in the vertex model with active junctions as a function of and , averaged over simulations with different realisations of the myosin noise.
(A) Probability of a central T1 transition. The red line is the 50% probability contour of any T1 occurring in the simulation. (B) Typical timescale for the T1 transition to occur, measured as the length of the contraction phase. The other parameters are the same as in Figure 2. (C) Magnitude of the convergence–extension deformation as a function of and characterised by measuring induced by the T1 transition.
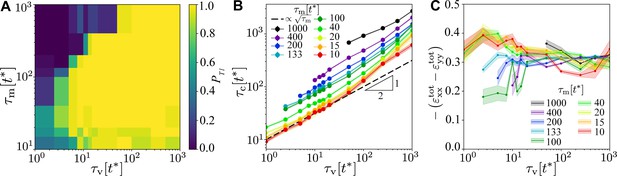
Robustness of the T1 mechanism as a function of and for and .
(A) Probability of a central T1 event, averaged over simulations with different realisations of the myosin noise. The probability of any T1 event is 1 throughout. (B) Contraction time to collapse for the central T1 as a function of , for different values of , for points where a central T1 event occurred in at least 25% of simulations. (C) Peak of the total convergence extension strain , showing very weak dependence on viscoelastic and myosin timescales. Shading in panels (B) and (C) indicates the standard error of the mean.
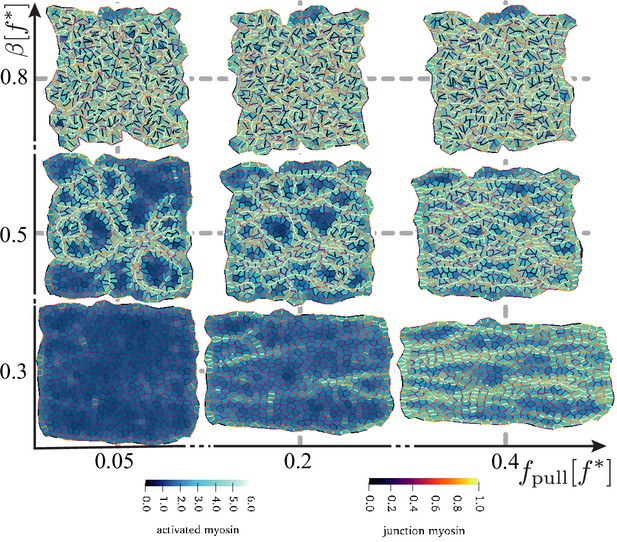
Disordered active tissue at time as a function of the magnitude of the pulling force and activity .
The region of convergence–extension is at the centre of the diagram, around and . The remaining parameters are the same as in Figure 2.
Video of simulations of active contractions in disordered tissues for a range of activities, β, and magnitudes of the pulling force, .
The system is first stretched for with activity switched off. During the initial stretching period, myosin is allowed to build anisotropy, but it does not feed back onto tension. This results in the tissue extending in the direction of external force and contracting in the direction perpendicular to it. After the tissue had reached mechanical equilibrium, the activity was turned on, with myosin being allowed to exert active contractions. All systems were simulated for an additional . Snapshots used to make the video were recorded once every . Simulations parameters are given in the caption of Figure 6.
Active T1 transitons in a random patch.
Activity is set to and the patch is pulled horizontally with with and . Left panel - junction and cell myosin; right panel - junction tension, and all cells are active.
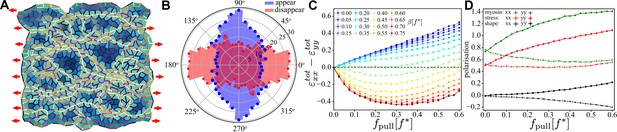
Characterisation of convergence-extension in a random tissue patch.
(A) Snapshot of a random tissue patch for and at ,that is during the convergence–extension flow. Red arrow indicates that a constant pulling force is applied throughout the entire simulation. (B) Angular histogram of T1 events for same values of and . Cells and junctions are coloured as in Figure 6. (C) Magnitude of the convergence–extension deformation as a function of characterised by measuring induced by the T1 transition. (D) Anisotropy along (, solid line) and perpendicular to (, dashed line) the direction of the external pulling force for myosin (green), mechanical stress (red), and shape tensor (black) as functions of for at . In (C) and (D), each point was averaged over independent samples and the error bar is smaller than the symbol size.
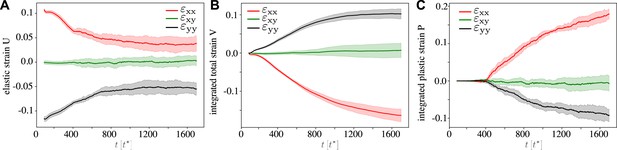
Continuous strain tensors for the fully active tissue at , , and the other parameters corresponding to Figure 2 and Figure 3.
(A) Statistical strain tensor defined in Equation 20, with the initial undeformed state used as reference configuration. (B) Total strain, that is, integrated tensor defined in Equation 24. (C) Integrated plastic strain tensor defined in Equation 22. Convergence–extension flow proceeds with cells elongated in the pulling direction, that is, and while and in the total strain. Here, the traces are an average over simulations, and the shading corresponds to the standard deviation.

Measuring T1 transitions in the fully active random patch.
A representative example taken from a sample of simulations for , . (A) Appearing and disappearing junctions as a function of simulation frame number and angle, before filtering. Inset: raw T1 histogram. (B) Same dataset after filtering procedure.
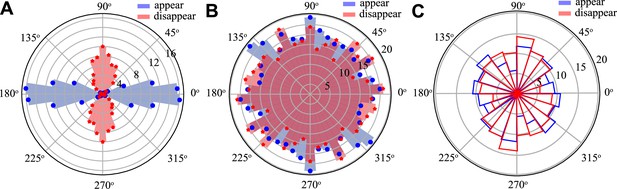
T1 histograms in the parameter range where no convergence–extension occurs.
(A) Passive tissue at pulled with , T1s are passive and aligned along pulling direction. (B) Active tissue at with no applied force , here T1s are distributed isotropically. (C) Experimental T1 distribution in the anterior region of the streak, strongly resembling the active isotropic case.
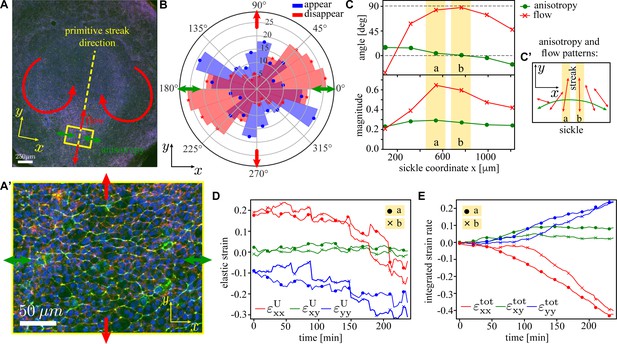
Analysis of the tissue flows in the early-stage chick embryo.
(A) Image of a typical early-stage chick embryo prior to the gastrulation (i.e. primitive streak formation). The primitive streak will form along the yellow dashed line. The direction of myosin anisotropy is shown by the green double-headed arrows, and the direction of the tissue flow is indicated by the red arrows. x-axis is chosen to coincide with the long direction of the sickle-shaped active region in the embryo’s posterior (Rozbicki et al., 2015). (A’) Zoom-in of the rectangular region on the posterior side of the embryo; myosin II (green), actin (red), and nuclei (blue). (B) Measured distribution of the orientation of T1 events in a circular patch of diameter ≈ 190 tracked over the period of ≈ 6 hr (cf. model distribution in Figure 7B). Blue (red) denotes junctions that appear (disappear); arrows have the same meaning as in (A). (C) Angle (red) and magnitude (green) of tissue shape anisotropy (dots) and tissue flow (crosses) for rectangular patches along the sickle with corresponding anisotropy and flow patterns shown in (C’). Components of the elastic strain tensor (D) and the total integrated strain tensor (E; definition in Equation 24) as a function of time during first 4 hr of the streak formation for two central regions of the sickle (yellow stripes in C). Details of the analysis are given in ‘Materials and methods’.
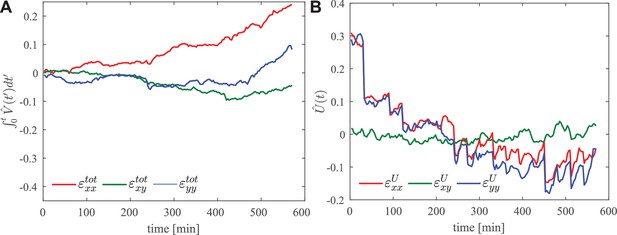
Region of interest in the anterior of the embryo.
(A) Integrated total strain and (B) elastic strain .
Image sequence taken at the base of the forming streak showing cell intercalations.
Centres of some cells are labelled with coloured dots for easier identification. Crossed blue and red lines indicate individual intercalation events. Blue lines indicate the direction of junction contraction and red lines the direction of extension of newly formed junctions. White scale bar 25 μm.
Image sequence taken in the region of epiblast in front of forming streak.
Centres of some cells are labelled with coloured dots for easier identification. Crossed blue and red lines indicate individual intercalation events. Blue lines indicate the direction of junction contraction and red lines the direction of extension of newly formed junctions. Images were taken at 3 min intervals. White scale bar 50 μm.
Tables
Values of the parameters in the single-junction model.
Units: length (), time (), force ().
| Base | ||
|---|---|---|
| Parameter | Description | |
| Spring constant | ||
| Barrier rest length | ||
| Friction with substrate | ||
| Model | ||
| Parameter | Description | Value range |
| Barrier spring constant | ||
| Applied external tension | ||
| Myosin activity | ||
| Viscoelastic time | ||
| Myosin time | ||
| Myosin reference level | 0.5 | |
| Threshold tension | ||
| Slope of vs. at | ||
| Tension-independent myosin dissociation | 1 | |
Values of the parameters used in the vertex model with active junctions.
Units: length (), time (), force ().
| Base | ||
|---|---|---|
| Parameter | Description | |
| Hexagonal cell edge length | ||
| Perimeter modulus | ||
| Spring constant | ||
| Friction with substrate | ||
| Model | ||
| Parameter | Description | Value range |
| Area modulus | ||
| Target cell area | ||
| Target cell perimeter | ||
| Pulling force | ||
| Myosin activity | ||
| Viscoelastic time | ||
| Myosin time | ||
| Threshold tension | ||
| Slope of vs. at | ||
| Myosin reference level | 0.5 | |
| Total cell myosin | 6 | |
| Variance of myosin fluctuations | 1 | |
| Tension-independent myosin dissociation | 0.1 | |





