Characterization of full-length CNBP expanded alleles in myotonic dystrophy type 2 patients by Cas9-mediated enrichment and nanopore sequencing
Figures
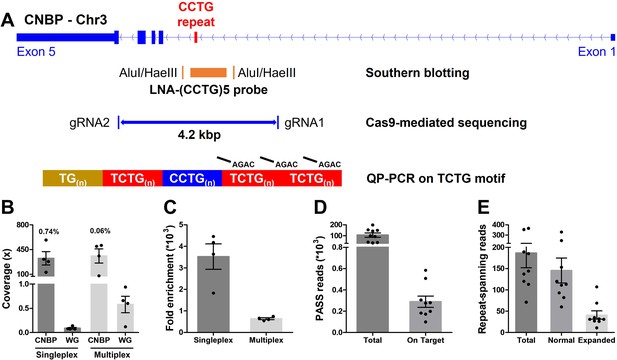
Cas9-mediated sequencing of the CNBP microsatellite.
(A) Experimental methods applied retrospectively to study the CNBP microsatellite in nine confirmed dystrophy type 2 (DM2) patients. The positions of AluI and HaeIII restriction sites (142 bp upstream and 108 bp downstream of the CCTG array, respectively) and the (CCTG)5 probe (orange rectangle) for Southern blot hybridization are shown, along with the gRNAs cleavage site for Cas9-mediated enrichment (boundaries of blue line = 4.2 kb) and the annealing position of P4TCTG primers for quadruplet-repeat primed PCR (QP-PCR). (B) Average coverage and (C) fold enrichment obtained by Cas9-mediated enrichment in DM2 patients following singleplex (n = 4) or multiplex (n = 4) runs. Numbers above bars represent the percentage of on-target reads. (D) Total and on-target number of PASS reads generated for each DM2 patient and (E) number of reads fully spanning the CNBP microsatellite and attributable to either the normal or expanded alleles. Data are plotted as means ± standard error of the mean (SEM). WG, whole genome. Sequencing statistics of singleplex and multiplex experiments are reported in detail in Supplementary file 1.
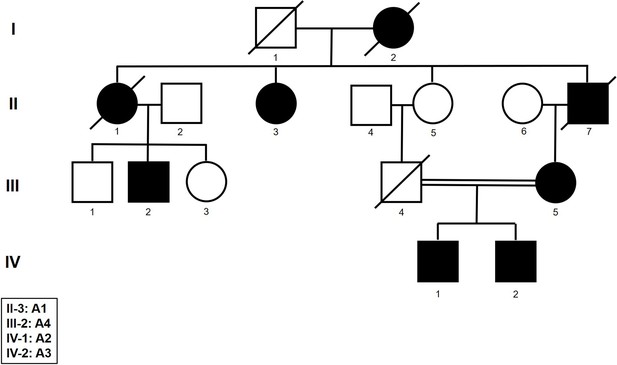
Pedigree of the dystrophy type 2 (DM2) family analyzed.
Genetic pedigree showing the relationship among A1–A4 familiar cases analyzed in this study. DNA sample from III-5 was not available for ONT sequencing.
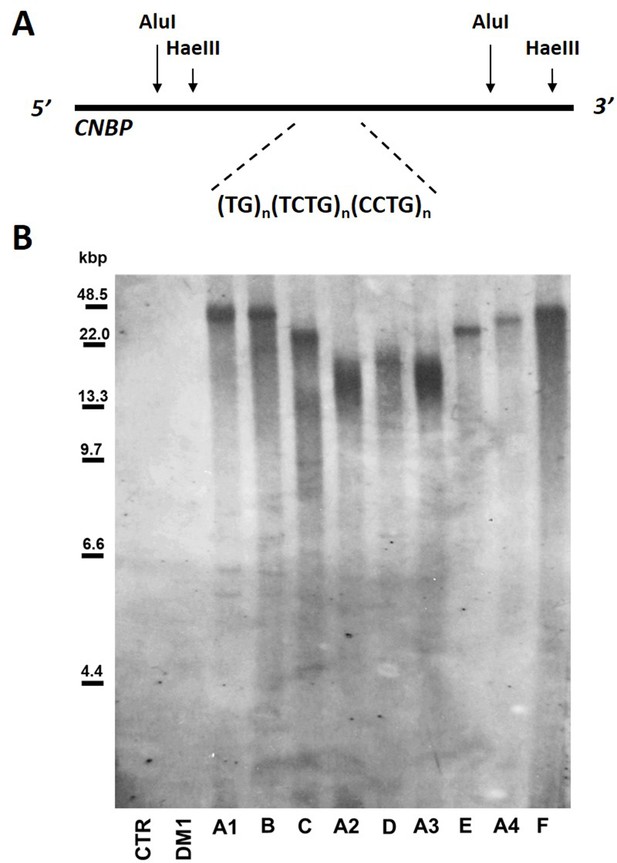
Southern blot analysis of expanded alleles in dystrophy type 2 (DM2) patients.
(A) Restriction map of the DM2 locus indicating the AluI and HaeIII restriction sites used for the digestion of genomic DNA. (B) Southern blot analysis of genomic DNA double digested with AluI and HaeIII and probed with a digoxigenin (DIG)-labeled (CCTG)5 locked nucleic acid (LNA) probe. Lane 1, CTR, healthy control sample; lane 2, DM1 sample; lanes 3–11, DM2 samples. Molecular markers are indicated on the left. The original scan of the Southern blot analysis reported in panel B and the whole figure incorporating the original uncropped scan can be found in Figure 1—figure supplement 2—source data 1 and 2, respectively.
-
Figure 1—figure supplement 2—source data 1
Original scan of the Southern blot analysis reported in Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/80229/elife-80229-fig1-figsupp2-data1-v2.zip
-
Figure 1—figure supplement 2—source data 2
Figure 1—figure supplement 2 including the original uncropped scan of the Southern blot analysis.
(A) Restriction map of the dystrophy type 2 (DM2) locus indicating the AluI and HaeIII restriction sites used for the digestion of genomic DNA. (B) Southern blot analysis of genomic DNA double digested with AluI and HaeIII and probed with a digoxigenin (DIG)-labeled (CCTG)5 locked nucleic acid (LNA) probe. Lane 1, CTR, healthy control sample; lane 2, DM1 sample; lanes 3–11, DM2 samples. Molecular markers are indicated on the left. *indicates aspecific bands, as reported by Nakamori et al., 2009.
- https://cdn.elifesciences.org/articles/80229/elife-80229-fig1-figsupp2-data2-v2.zip
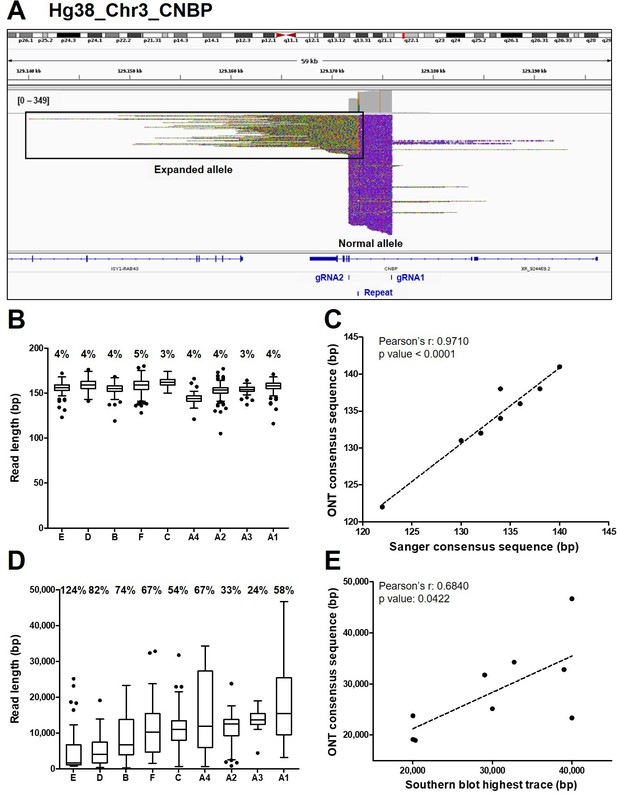
Analysis of ONT sequencing data from normal and expanded CNBP alleles.
(A) Integrative Genomics Viewer (IGV) visualization of ONT sequencing data at the CNBP locus of a representative dystrophy type 2 (DM2) patient following Cas9-mediated enrichment. Reads generated from the normal allele feature clear cuts on both sides of the CNBP repeat, whereas those derived from the expanded allele are longer, soft-clipped and do not match the reference genome, as expected. Length distributions of reads derived from the normal alleles (B) and expanded alleles (D) of each patient. Boxes represent the interquartile range (IQR) of lengths, the horizontal line is the median, whiskers and outliers are plotted according to Tukey’s method. (C) Correlation between the length of ONT and Sanger consensus sequences for the normal allele (n = 9). (E) Correlations between the maximum length of ONT sequences (longest complete read) and the upper edge of the Southern blot trace for the expanded allele (n = 9). Numbers on top of panels (B) and (D) indicate the coefficient of variation of normal and expanded alleles, respectively.
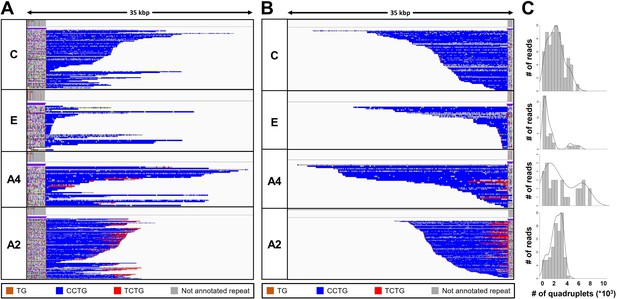
Analysis of the expanded-repeat CNBP alleles in dystrophy type 2 (DM2) patients.
Integrative Genomics Viewer (IGV) visualization (35-kbp windows) of ONT-targeted sequencing data from the expanded alleles of four representative DM2 patients. Complete reads were aligned at the 5′ end (A) and then at the 3′ end (B) in order to identify the repeat pattern that characterizes the expanded microsatellite locus. Each motif in the expanded alleles was visualized using a different color, as indicated in the key. Samples C and E contained a ‘pure’ CCTG expansion (blue) whereas samples A4 and A2 also contained the unexpected TCTG motif (red) downstream of CCTG. (C) Abundance of quadruplets identified in each patient. The y-axis shows the number of ONT reads with a certain number of repeats, whereas the x-axis shows the number of quadruplet repeats identified. ONT reads were grouped into 500 bp bins. The gray line represents the estimated kernel density of the underlying solid gray distribution of ONT reads.
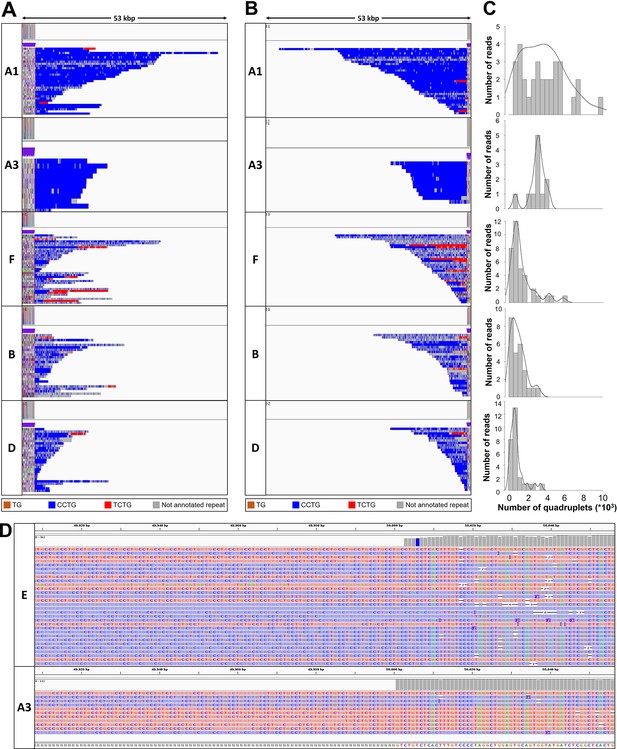
Analysis of the CNBP repeat motif for the expanded alleles in dystrophy type 2 (DM2) patients.
Integrative Genomics viewer (IGV) visualization (53-kbp window) of ONT-targeted sequencing data from the expanded alleles of the five remaining DM2 samples. Complete reads were aligned at the 5′ end (A) and subsequently at the 3′ end (B) in order to identify the repeat pattern characterizing the expanded microsatellite locus. Each motif in the expanded alleles was visualized using a different color, as indicated in the key. All samples feature the unexpected TCTG motif of variable length downstream of the CCTG motif. (C) Abundance of quadruplets identified in each patient. The y-axis shows the number of ONT reads with a certain number of repeats, whereas the x-axis shows the number of quadruplet repeats identified. ONT reads were grouped into 500 bp bins. The gray line represents the estimated kernel density of the underlying solid gray distribution of ONT reads. (D) IGV visualization of ONT-targeted sequencing data from the expanded alleles of two representative patients (E and A3) carrying a pure CCTG expansion and a repeat with the TCTG motif, respectively.
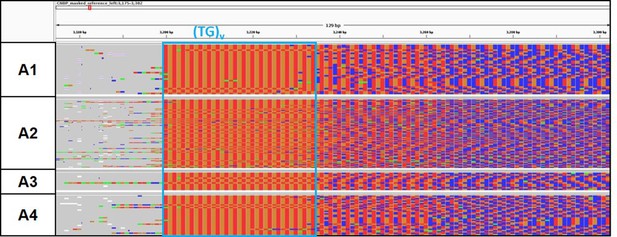
Analysis of the CNBP 5′-end (TG)v repeat motif of the CNBP expanded alleles of A1–A4 dystrophy type 2 (DM2) family members.
Integrative Genomics viewer (IGV) visualization (~130 bp window) of ONT-targeted sequencing data from the expanded alleles of the DM2 family members. Complete reads were aligned at the 5′ end in order to identify the repeat pattern characterizing the expanded microsatellite locus. Nucleotides of the expanded repeat have been colored with A: green, C: blue, G: orange, and T: red.
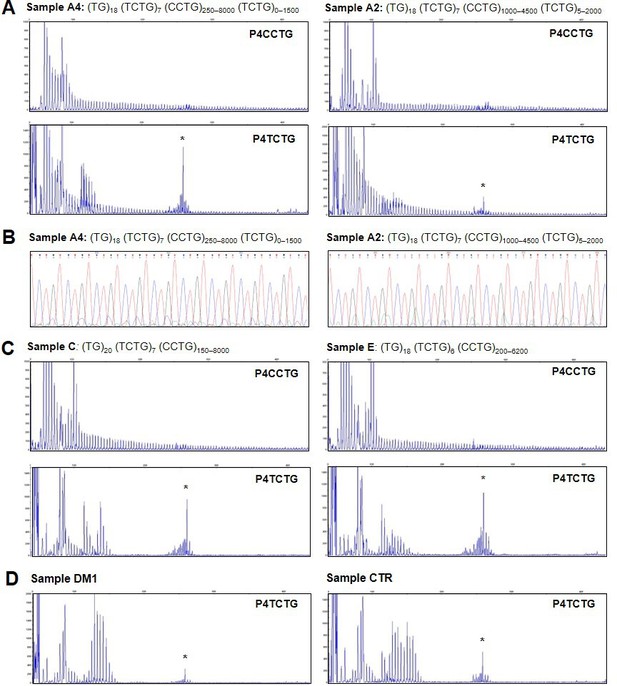
Analysis of the TCTG motif by quadruplet-repeat primed PCR (QP-PCR) and Sanger sequencing.
(A) Representative QP-PCR profiles of genomic DNA samples from patients A2 and A4 and showing the presence of the TCTG block. Upper panels show QP-PCR results using the conventional P4CCTG primer. Lower panels show QP-PCR results using primer P4TCTG. (B) Sanger sequencing of the QP-PCR products from P4TCTG reaction confirming the presence of the TCTG sequence. (C) QP-PCR profiles of genomic DNA samples from patients C and E showing only the traditional dystrophy type 2 (DM2) motif CCTG. For each patient, the composition of CNBP expanded alleles above the QP-PCR tracks reflects the ONT sequencing data. Asterisks (*) indicate nonspecific signals, also visible in the QP-PCR profiles of DM1 and CTR samples (D).
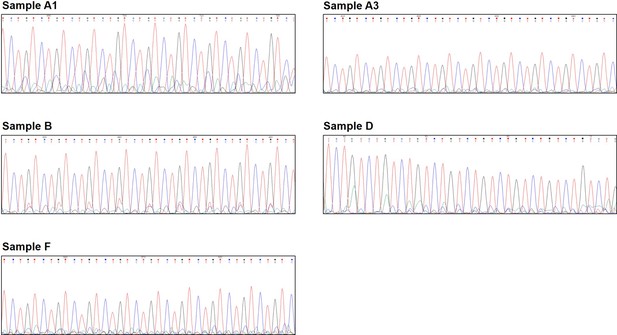
Sanger sequencing of quadruplet-repeat primed PCR (QP-PCR) products showing the presence of the (TCTG)n motif.
QP-PCR was carried out using primer P4TCTG and genomic DNA from dystrophy type 2 (DM2) patients A1, A3, B, D, and F. Sanger sequencing of the QP-PCR products confirmed the presence of the (TCTG)n array at the 3′ end of the (CCTG) expansions.
Tables
Demographic and molecular features of the dystrophy type 2 (DM2) patients.
For each patient, the table shows the ID, sex, age, age at onset, repeat length, and structure of the CNBP normal allele characterized by Sanger sequencing, and the estimated repeat length of the expanded allele determined by Southern blotting (–, not available).
| Normal allele | Expanded allele | |||||
|---|---|---|---|---|---|---|
| Sample ID | Sex | Age | Age at onset | Repeat length (bp) | Repeat structure | Repeat length (bp) |
| A1 | F | 75 | 70 | 136 | (TG)24 (TCTG)7 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 40,000 |
| A2 | M | 27 | 25 | 130 | (TG)17 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 20,000 |
| A3 | M | 21 | – | 132 | (TG)20 (TCTG)8 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 20,323 |
| A4 | M | 65 | 61 | 122 | (TG)19 (TCTG)9 (CCTG)12 | 32,745 |
| B | F | 49 | 44 | 134 | (TG)21 (TCTG)7 (CCTG)6 GCTG CCTG TCTG (CCTG)7 | 40,000 |
| C | M | 20 | – | 140 | (TG)24 (TCTG)6 (CCTG)7 GCTG CCTG TCTG (CCTG)7 | 29,027 |
| D | M | 44 | 39 | 134 | (TG)19 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 20,000 |
| E | F | 61 | 43 | 134 | (TG)19 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 30,000 |
| F | M | 56 | 50 | 138 | (TG)21 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 39,000 |
CNBP repeat analysis based on Cas9-mediated sequencing of the normal alleles.
For each patient, the table shows the characteristics of normal CNBP alleles based on the analysis of ONT sequencing data, in terms of length and structure. The table reports the percentage identity of consensus sequences reconstructed from ONT or Sanger sequencing data and the incongruences in ONT-derived sequences are highlighted in bold.
| Normal allele | Sample ID | Repeat length (bp) | Repeat structure | Identity with Sanger sequence |
|---|---|---|---|---|
| A1 | 136 | (TG)24 (TCTG)7 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 100.0% | |
| A2 | 131 | (TG)17 TGCTG (TCTG)8 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 99.2% | |
| A3 | 132 | (TG)20 (TCTG)8 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 100.0% | |
| A4 | 122 | (TG)19 (TCTG)9 (CCTG)12 | 100.0% | |
| B | 134 | (TG)21 (TCTG)7 (CCTG)6 GCTG CCTG TCTG (CCTG)7 | 100.0% | |
| C | 141 | (TG)24 TGCTG (TCTG)5 (CCTG)7 GCTG CCTG TCTG (CCTG)7 | 99.3% | |
| D | 138 | (TG)21 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 97.1% | |
| E | 134 | (TG)19 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 100.0% | |
| F | 138 | (TG)21 (TCTG)9 (CCTG)5 GCTG CCTG TCTG (CCTG)7 | 100.0% |
CNBP repeat analysis based on Cas9-mediated sequencing of the expanded alleles.
For each patient, the table shows the characteristics of expanded CNBP alleles based on the analysis of ONT sequencing data, in terms of length and structure. The expanded (CCTG)x are colored in blue and (TCTG)y in red. Potential incongruences in ONT-derived sequences are highlighted in bold. The table indicates also the fraction of reads carrying the unexpected TCTG motif at the 3′ end of the CNBP microsatellite expansion.
| Expanded allele | Sample ID | Repeat length (bp) (min–max) | Repeat structure | Number of reads carrying the TCTG motif |
|---|---|---|---|---|
| A1 | 3241–46,685 | (TG)20 (TCTG)7 ( | 14 (45%) | |
| A2 | 864–23,779 | (TG)18 (TCTG)7 | 92 (86%) | |
| A3 | 4429–18,983 | (TG)19 (TCTG)7 | 8 (73%) | |
| A4 | 660–34,284 | (TG)18 (TCTG)7 | 9 (29%) | |
| B | 344–23,358 | (TG)18 (TCTG)7 | 6 (23%) | |
| C | 700–31,753 | (TG)20 (TCTG)7 | 0 (0%) | |
| D | 383–19,143 | (TG)18 (TCTG)7 | 3 (11%) | |
| E | 848–25,162 | (TG)18 (TCTG)6 | 0 (0%) | |
| F | 1533–32,824 | (TG)15 (TCTG)10 | 15 (43%) |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Homo sapiens) | CNBP | Ensembl | HGNC:13164 | Hg38 |
| Biological sample (Homo sapiens) | Anti-coagulated peripheral blood | Policlinico Tor Vergata, Rome, Italy | Patient A1, A2, A3, A4, B, C, D, E, F | |
| Sequence-based reagent | Digoxigenin (DIG)-labeled locked nucleic acid (LNA) probe | Nakamori et al., 2009 | DIG-LNA probe | (CCTG)5 |
| Sequence-based reagent | P4TCTG | This paper | PCR primers | agc gga taa caa ttt cac aca gga TCT GTC TGT CTG TCT GTC TGT |
| Sequence-based reagent | CL3N58_DR-[FAM] | This paper | PCR primers | GCC TAG GGG ACA AAG TGA GA |
| Sequence-based reagent | P3 | This paper | PCR primers | AGC GGA TAA CAA TTT CAC ACA GGA |
| Sequence-based reagent | crRNA1_CNBP | This paper | CRISPR RNA | CCA CCT GAT TCA CTG CGA TA |
| Sequence-based reagent | crRNA2_CNBP | This paper | CRISPR RNA | GGC TTC TCA TTC CAC GAC CA |
| Sequence-based reagent | Native barcodes | Oxford Nanopore Technologies (ONT) | EXP-NBD104 | |
| Commercial assay or kit | DIG-High Prime DNA Labeling and Detection Starter Kit II | Roche | Cat. No. 11585614910 | |
| Commercial assay or kit | Flexigene DNA Kit | Qiagen | Cat. No. 51206 | |
| Commercial assay or kit | BigDye Terminator v3.1 Cycle Sequencing Kit | Thermo Fisher | Cat. No. 4337458 | |
| Commercial assay or kit | Nanobind CBB Big DNA HMW Kit | Circulomics | SKU 102-301-900 | |
| Commercial assay or kit | NucleoSpin Blood L Kit | Macherey-Nagel | Item number: 740954.20 | |
| Commercial assay or kit | Qubit dsDNA BR Assay Kit | Thermo Fisher Scientific | Cat. No. Q32853 | |
| Commercial assay or kit | TapeStation DNA ScreenTape & Reagents | Agilent Technologies | Cat. No. 5067–5365 5067–5366 | |
| Software, algorithm | GeneMapper Software 6 | Applied Biosystems | Cat. No. 4475074 | |
| Software, algorithm | CHOPCHOP | Labun et al., 2019 | chopchop.cbu.uib.no/ | |
| Software, algorithm | Guppy v3.4.5 | Computational Biology Research Center – AIST | ||
| Software, algorithm | NanoFilt v2.7.1 | De Coster et al., 2018 | ||
| Software, algorithm | BBMap suite v38.87 | https://sourceforge.net/projects/bbmap/ | ||
| Software, algorithm | Tandem Repeat Finder v4.09 | Benson, 1999 | ||
| Software, algorithm | Minimap2 v2.17-r941 | Li, 2018 | ||
| Software, algorithm | Integrative Genomics Viewer (IGV) v2.8.3 | Robinson, 2011 | ||
| Software, algorithm | Scripts for the generation of consensus sequences and repeat annotations for the normal allele | https://github.com/MaestSi/CharONT2 | Script Name: CharONT2 | |
| Software, algorithm | Scripts for the annotation of repeats and the generation of simplified reads for the expanded allele | https://github.com/MaestSi/MosaicViewer_CNBP | Script Name: MosaicViewer_CNBP v1.0.0 | |
| Software, algorithm | MinKNOW V20.06.5 | Oxford Nanopore Technologies | ||
| Commercial assay or kit | Alt-R S.p. HiFi Cas9 Nuclease v3 | IDT | Cat. No. 1081060 | Recombinant Cas9 nuclease for target excision (see M&M) |
| Commercial assay or kit | Alt-R CRISPR-Cas9 tracrRNA | IDT | Cat. No. 1072532 | Structural RNA for gRNA formation (see M&M) |
| Commercial assay or kit | AMPure XP Beads | Beckman-Coulter | Product No. A63881 | Magnetic beads for nucleic acid purification (see M&M) |
| Commercial assay or kit | CutSmart buffer 10× | New England BioLabs | Cat. No. B7204 | Buffer for gDNA dephosphorylation, RNP formation, and target excision (see M&M) |
| Commercial assay or kit | Blunt/TA Ligase Master Mix | New England BioLabs | Cat. No.: M0367S | T4 DNA ligase for native barcode ligation to dA-tailed ends (see M&M) |
| Commercial assay or kit | FLO-MIN106D (R9.4.1) flow cell | Oxford Nanopore Technologies | FLO-MIN106D | Flowcell for ONT sequencing (see M&M) |
Additional files
-
Supplementary file 1
Sequencing statistics of singleplex and multiplex experiments.
The table reports the feature of each Cas9-mediated sequencing experiment performed and the sequencing statistics of each ONT run. Average values are also provided for singleplex and multiplex runs separately.
- https://cdn.elifesciences.org/articles/80229/elife-80229-supp1-v2.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/80229/elife-80229-mdarchecklist1-v2.pdf





