Phage tRNAs evade tRNA-targeting host defenses through anticodon loop mutations
Figures
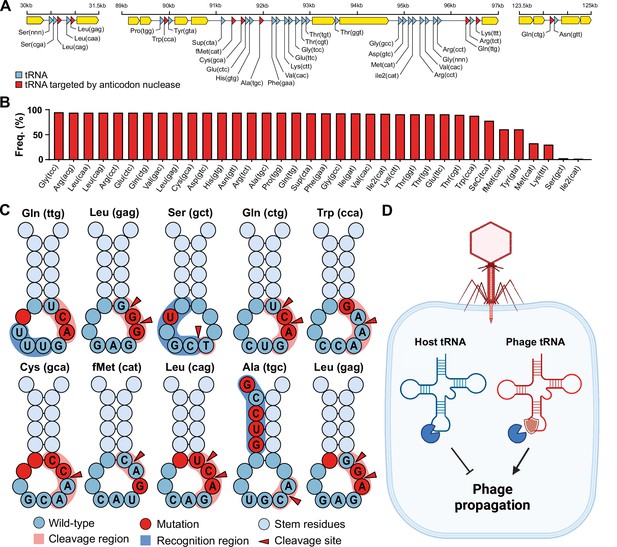
Phage transfer RNAs (tRNAs) are predicted to be anticodon nuclease resistant.
(A) The genomic context of the tRNA clusters containing 36 tRNAs present in C1 mycobacteriophage Rizal (Russell and Hatfull, 2017). (B) Prevalence of individual phage-encoded tRNAs in the C1 mycobacteriophage cluster, composed of 161 phages. (C) Mutations in the anticodon loop of phage tRNAs in comparison to host tRNAs, located in the cleavage site of anticodon nucleases. (D) Proposed mechanism of action of phage tRNAs. During phage infection, tRNAses are activated and deplete the host tRNA pool via tRNA cleavage to prevent phage propagation. Phage tRNAs are insensitive to cleavage, allowing the phage to propagate.
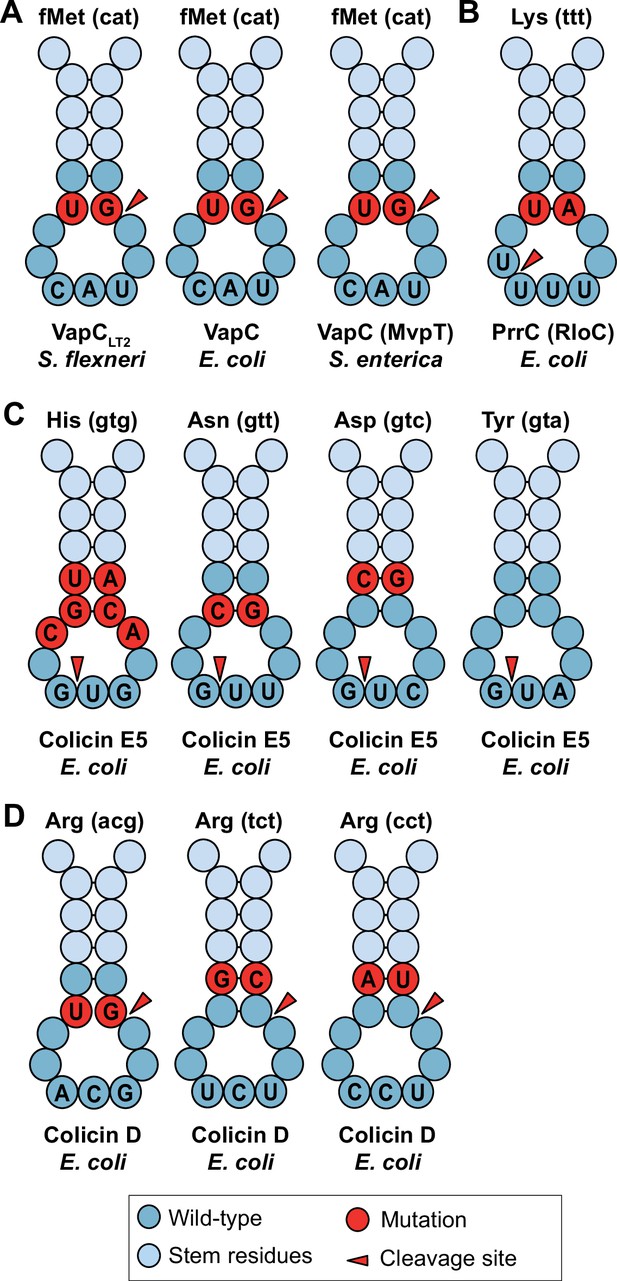
Phage transfer RNAs (tRNAs) from enterobacteria.
(A) Mutations in the anticodon loop of phage tRNAs in comparison to host tRNAs, located in the cleavage site of anticodon nucleases of tRNA fMet (cat) that are targeted by VapCs. (B) Mutations in the anticodon loop of phage tRNAs in comparison to host tRNAs, located in the anticodon loop of tRNA Lys (ttt) that is targeted by PrrC (Rloc), which is repaired by the tRNA-Lys ligase. (C) Mutations in the anticodon loop of phage tRNAs in comparison to host tRNAs, located in the anticodon loops of tRNAs that are targeted by Colicin E5 in the anticodon itself. (D) Mutations in the anticodon loop of phage tRNAs in comparison to host tRNAs, located in the anticodon loops of tRNAs that are targeted by Colicin D from E. coli.
Additional files
-
Supplementary file 1
Phage versus host-encoded transfer RNA (tRNA) comparisons.
(a) Overview of all C1 mycobacteriophage encoded tRNAs. (b) Overview of anticodons encoded by C1 mycobacteriophages compared to their hosts'. (c) Analysis of anticodon loop mutations and anticodon avoidance in other species.
- https://cdn.elifesciences.org/articles/85183/elife-85183-supp1-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/85183/elife-85183-mdarchecklist1-v2.docx





