Genetic validation of PfFKBP35 as an antimalarial drug target
Figures
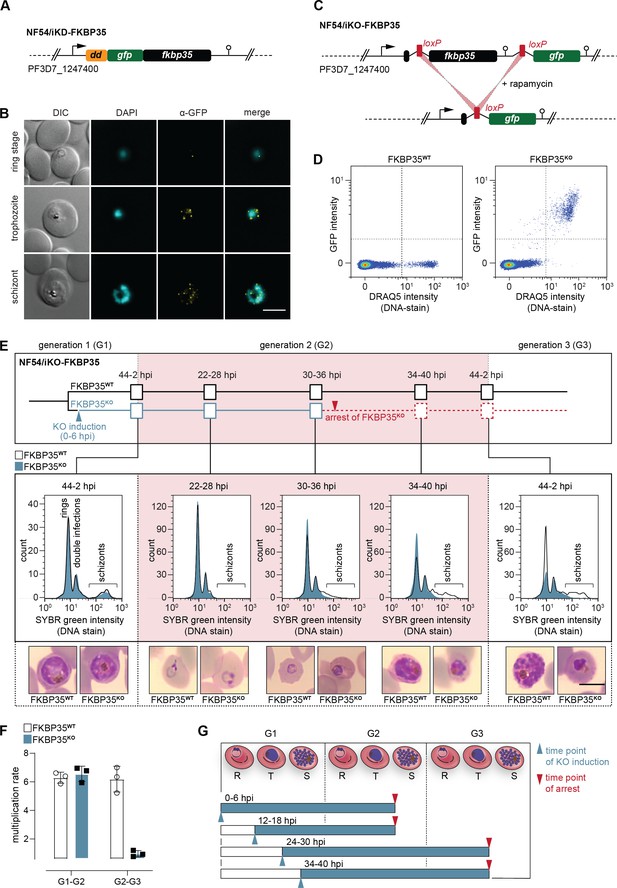
PfFKBP35 depletion causes delayed parasite death.
(A) Schematic of the N-terminally modified fkbp35 locus in NF54/iKD-FKBP35 parasites. dd, destabilization domain; gfp, green fluorescent protein. (B) Localization of PfFKBP35 in NF54/iKD-FKBP35 parasites at the ring, trophozoite, and schizont stage assessed by immunofluorescence. DNA was stained with DAPI. Representative images are shown. Scale bar: 5 µm. DIC, differential interference contrast. (C) Schematic of the modified fkbp35 locus in NF54/iKO-FKBP35 parasites before and after loxP recombination. gfp, green fluorescent protein. (D) Flow cytometry plot showing GFP expression in G1 schizonts of FKBP35KO and FKBP35WT parasites. DNA was stained with DRAQ5. (E) Time course showing the development of NF54/iKO-FKBP35 parasites. Upper panel: schematic of the experimental setup. Middle panel: DNA content of the populations assessed by flow cytometry based on SYBR Green intensity. Lower panel: Giemsa-stained blood smears. The knock-out was induced at 0–6 hpi. Representative data of three biological replicates are shown. Scale bar: 5 µm. (F) Multiplication rates of FKBP35KO and FKBP35WT parasites from generation 1 to generation 2 (G1–G2) and G2–G3. The knock-out was induced at 0–6 hpi. Multiplication rates were determined using flow cytometry. Data points represent the multiplication rates from G1–G2 and G2–G3. n = 3, error bars represent the standard deviation. (G) Schematic showing time point of growth arrest (red arrow) following PfFKBP35 knock-out at different hpi (blue arrow). R, ring stage; T, trophozoite stage; S, schizont stage.
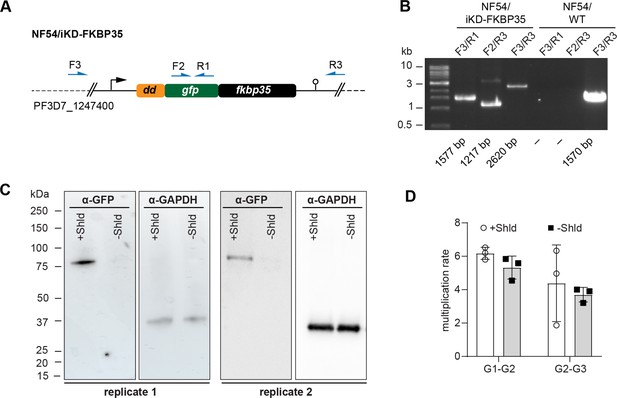
Characterization of NF54/iKD-FKBP35 parasites.
(A) Schematic of the modified fkbp35 locus (PF3D7_1247400) in NF54/iKD-FKBP35 parasites. The 5′-end of fkbp35 was tagged with the sequences of a destabilization domain (dd) and a green fluorescent protein (gfp). The DD tag allows regulating protein levels post-translationally using the stabilizing compound Shield-1 (Shld) (Armstrong and Goldberg, 2007; Banaszynski et al., 2006). Names and binding sites of the primers used for diagnostic PCRs are indicated. (B) Diagnostic PCRs performed on gDNA of NF54/FKBP35 iKD parasites to confirm correct editing of the locus. PCRs performed on NF54/WT gDNA served as control. Numbers at the bottom indicate expected fragment sizes. (C) Expression levels of the DD-GFP-FKBP35 fusion protein were assessed by Western blot. Parasite cultures were split at 0–6 hpi and treated with Shld or the vehicle control EtOH. Protein lysates were collected 40 hr later. α-GAPDH served as loading control. The expected size of the DD-GFP-FKBP35 fusion protein is 74.2 kDa. (D) Multiplication rates of NF54/iKD-FKBP35 parasites from generation 1 to generation 2 (G1–G2) and G2–G3. Parasites were split at 0–6 hpi and treated with Shld or the vehicle control EtOH. Multiplication rates were determined using flow cytometry. Data points represent the multiplication rates from G1–G2 and G2–G3, respectively. n = 3; error bars represent the standard deviation.
-
Figure 1—figure supplement 1—source data 1
Agarose gel electrophoresis of peqGREEN-stained DNA and SDS-polyacrylamide gel electrophoresis (SDS-PAGE) of protein samples.
- https://cdn.elifesciences.org/articles/86975/elife-86975-fig1-figsupp1-data1-v1.zip
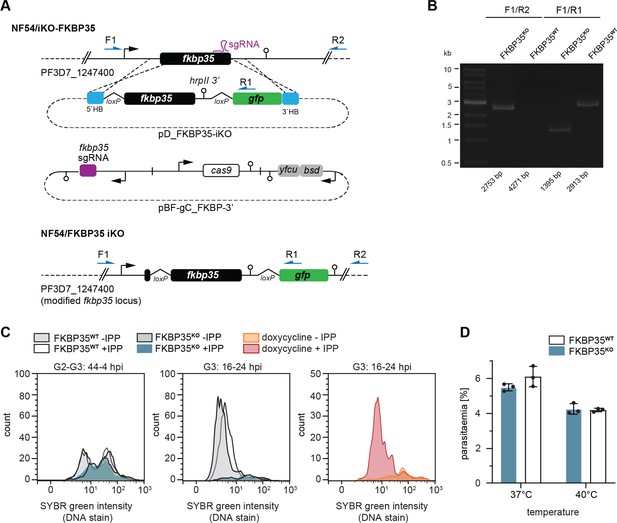
CRISPR/Cas9-based generation of transgenic parasite lines.
(A) Schematic of the wild-type fkbp35 locus (PF3D7_1247400) and the two plasmids used for CRISPR/Cas9 gene editing (pD_FKBP35-iKO and pBF-gC_FKBP-3′) to generate the modified fkbp35 locus of NF54/iKO-FKBP35 parasites. The modification consists of an intron containing a loxP site that was introduced after the first 38 bp of the coding sequence and an hrpII terminator inserted after the stop codon followed by another loxP site and a gfp sequence. The sequence flanked by the loxP sites was recodonized to prevent homologous recombination during gene editing. Names and binding sites of the primers used for diagnostic PCRs are indicated. (B) Diagnostic PCRs on gDNA of NF54/iKO-FKBP35 parasites confirm correct locus editing and rapamycin-induced loxP recombination. Numbers at the bottom indicate the expected band sizes. (C) DNA content analysis of FKBP35KO and FKBP35WT parasites in the presence or absence of isopentenyl pyrophosphate (IPP)-supplementation. Disruption of isoprenoid precursor biosynthesis in the apicoplast – the only essential function of this organelle – is a known cause for delayed death in P. falciparum (Wiley et al., 2015; Kennedy et al., 2019). Inhibition of isoprenoid synthesis by doxycycline can thus be restored by supplementing the parasite culture medium with the isoprenoid precursor IPP (Yeh and DeRisi, 2011). However, IPP supplementation did not alter the fate of FKBP35KO cells, demonstrating that their death in G2 is independent of apicoplast-dependent processes as shown by DNA content analysis. Doxycycline-treated wild-type parasites rescued by IPP-supplementation served as positive control. The knock-out was induced at 0–6 hpi. Representative data of three biological replicates are shown. (D) Parasite survival after exposure to 40°C for 6 hr. Given the reported interactions of PfFKBP35 with heat shock protein 70 (HSP70) and HSP90 (Leneghan and Bell, 2015; Kumar et al., 2005), we tested the parasite’s response to heat stress. PfHSP70 is required to ensure parasite survival under heat shock conditions by stabilizing the digestive vacuole membrane trough binding to phosphatidylinositol 3-phosphate (Lu et al., 2020). Despite this association, we did not detect altered heat-susceptibility between FKBP35KO and FKBP35WT cells. The knock-out was induced at 0–6 hpi. n = 3, error bars represent the standard error of the mean.
-
Figure 1—figure supplement 2—source data 1
Agarose gel electrophoresis of peqGREEN-stained DNA.
- https://cdn.elifesciences.org/articles/86975/elife-86975-fig1-figsupp2-data1-v1.zip
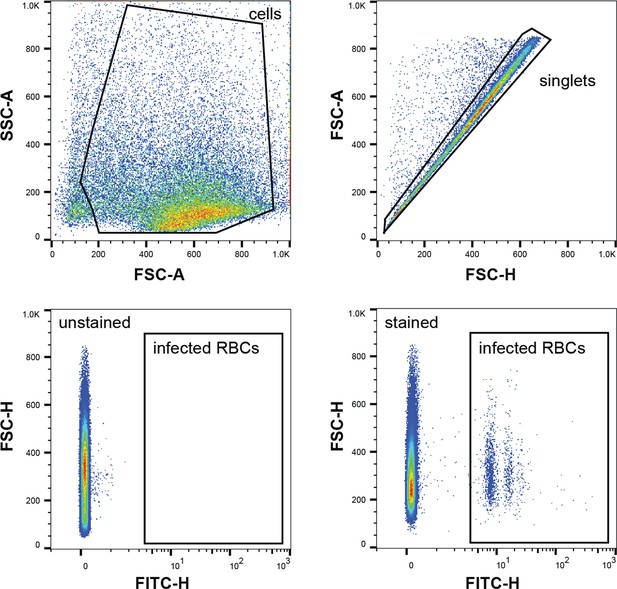
Gating strategy of flow cytometry data.
Representative flow cytometry plots of formaldehyde/glutaraldehyde-fixed parasite cultures are shown. The first plot shows the gate to distinguish cells from small debris (SSC-A vs. FSC-A). The ‘singlets’ gate (FSC-A vs. FSC-H) removes events with more than one cell. SYBR Green fluorescence intensity is used to identify parasite-infected RBCs (FSC-H vs. FITC-H). SSC, side scatter. FSC, forward scatter. FITC, fluorescein-5-isothiocyanate. A, area. H, height.
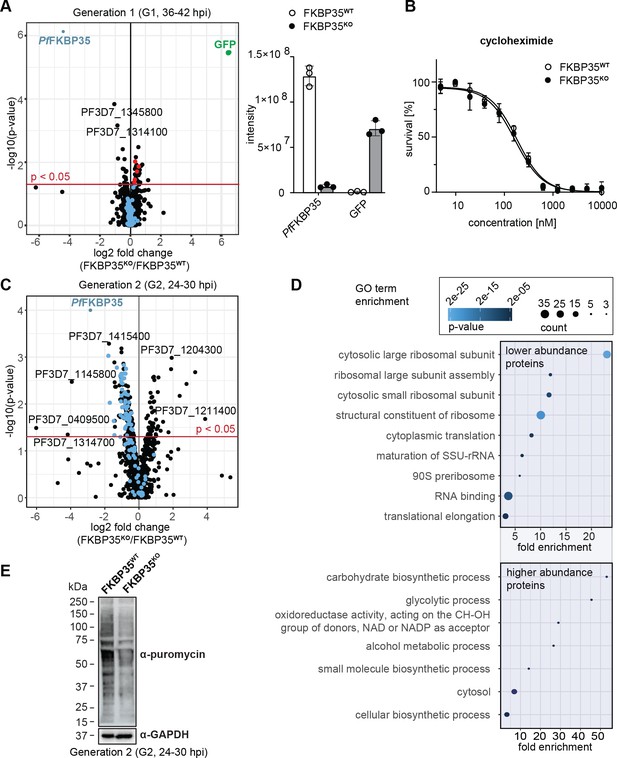
PfFKBP35 is crucial for ribosome homeostasis and protein synthesis.
(A) Left panel: Proteome analysis of schizonts (36–42 hpi) in G1. The volcano plot shows change in protein abundance in FKBP35KO compared to FKBP35WT. Ribosomal proteins are highlighted in blue, proteins associated with the GO terms ‘pre-ribosome’ or ‘nucleolus’ in red. P-values were calculated using empirical Bayes moderated t-tests and adjusted for multiple testing using the Benjamini-Hochberg method. n = 3. Right panel: relative difference in PfFKBP35 and GFP abundance expressed as signal intensity measured by mass spectrometry. n = 3, error bars represent the standard deviation. (B) Dose–response effect of the translation inhibitor cycloheximide on FKBP35KO and FKBP35WT parasites. The knock-out was induced at 0–6 hpi and parasites were subsequently exposed to cycloheximide for 48 hr. Parasite survival was quantified by flow cytometry. n = 3, error bars represent the standard error of the mean. (C) Proteome analysis of trophozoites (24–30 hpi) in G2. The volcano plot shows change in protein abundance in FKBP35KO compared to FKBP35WT. Ribosomal proteins are highlighted in blue. P-values were calculated using empirical Bayes moderated t-tests and adjusted for multiple testing using the Benjamini-Hochberg method. n = 3. (D) GO enrichment analysis of significantly deregulated proteins in FKBP35KO trophozoites in G2. GO enrichment analysis was performed using the PANTHER ‘Overrepresentation Test’ with the test type ‘Fisher’s exact’ and the correction method ‘false discovery rate.’ (E) Assessment of translation activity using SUnSET. FKBP35KO and FKBP35WT trophozoites of G2 were incubated with puromycin for 1 hr. Incorporated puromycin was detected using an α-puromycin antibody. α-GAPDH served as loading control. n = 3, a representative Western blot is shown.
-
Figure 2—source data 1
SDS-polyacrylamide gel electrophoresis (SDS-PAGE) of protein samples.
- https://cdn.elifesciences.org/articles/86975/elife-86975-fig2-data1-v1.zip
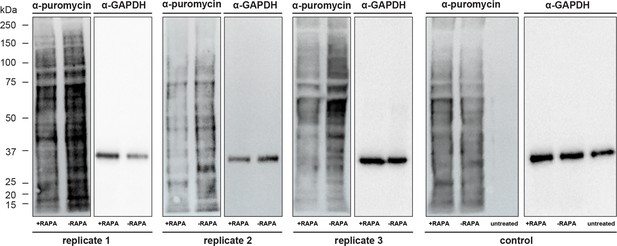
Assessment of translation rates using SUnSET.
FKBP35KO and FKBP35WT trophozoites of G1 were incubated with puromycin for 1 hr. Incorporated puromycin was detected using an α-puromycin antibody; α-GAPDH served as loading control. Replicate 3 is also shown in main Figure 2 (mirrored). The control was performed using an NF54/DiCre parasite line. Untreated control parasites were cultured in the absence of puromycin, confirming the specificity of the α-puromycin antibody.
-
Figure 2—figure supplement 1—source data 1
SDS-polyacrylamide gel electrophoresis (SDS-PAGE) of protein samples.
- https://cdn.elifesciences.org/articles/86975/elife-86975-fig2-figsupp1-data1-v1.zip
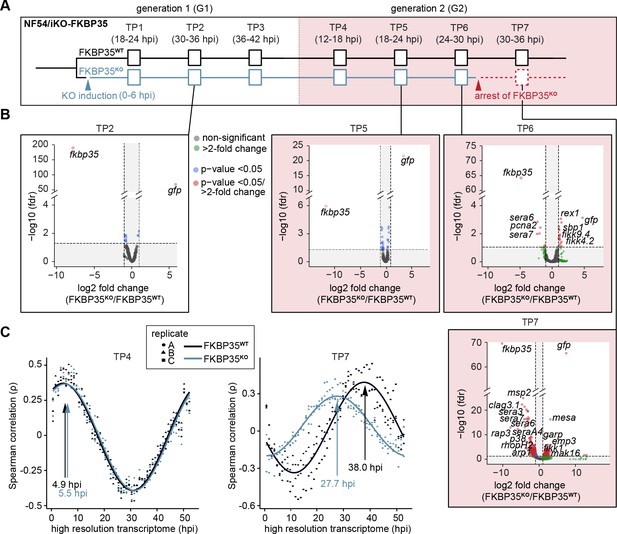
The knock-out of PfFKBP35 remains silent at the transcriptomic level.
(A) Setup of the RNA-seq experiment. RNA was collected at seven indicated time points (TP). (B) Volcano plots show differential gene expression between FKBP35KO and FKBP35WT parasites. Genes are color-coded according to the fold change and false discovery rate (FDR). Paired differential expression tests were performed using the DESeq2 likelihood ratio test. n = 3. (C) Mapping of FKBP35KO and FKBP35WT transcriptomes to a high-resolution reference (Bozdech et al., 2003). Spearman rank coefficients (ρ) show correlation between the sampled transcriptomes and the reference dataset (Bozdech et al., 2003). The TPs with the highest correlation coefficient are indicated.
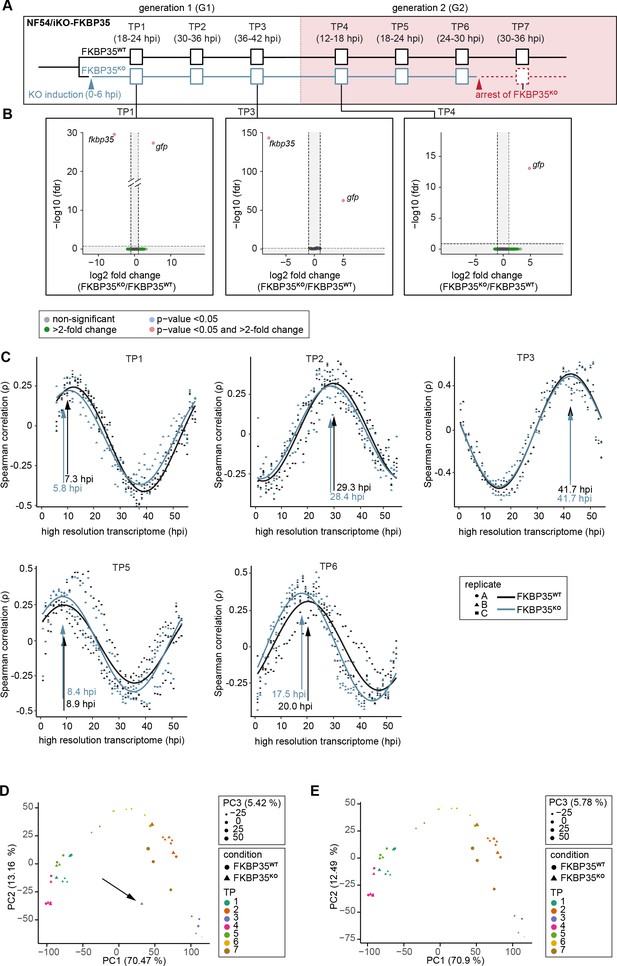
Transcriptomics reveals a cell cycle arrest of FKBP35KO parasites.
(A) Schematic of the experimental setup. The knock-out was induced at 0–6 hpi. RNA was collected at indicated time points (TPs). (B) Volcano plots show differential gene expression between FKBP35KO and FKBP35WT parasites. Genes are color-coded according to the fold change and false discovery rate (FDR). Paired differential expression tests were performed using the DESeq2 likelihood ratio test. n = 3. (C) Mapping of FKBP35KO and FKBP35WT transcriptomes to a high-resolution reference dataset (Bozdech et al., 2003). Spearman rank coefficients (ρ) describe the correlation between the sampled transcriptomes and the reference dataset. The TPs with the highest correlation coefficient are indicated. (D) Principal component analysis (PCA) of RNA-seq data by condition and TP. The outlier at TP3 (highlighted by an arrow) was removed from the analysis. (E) PCA of RNA-Seq data by condition and TP without the TP3 outlier mentioned in (D). Exclusion of this sample did not affect the overall pattern.
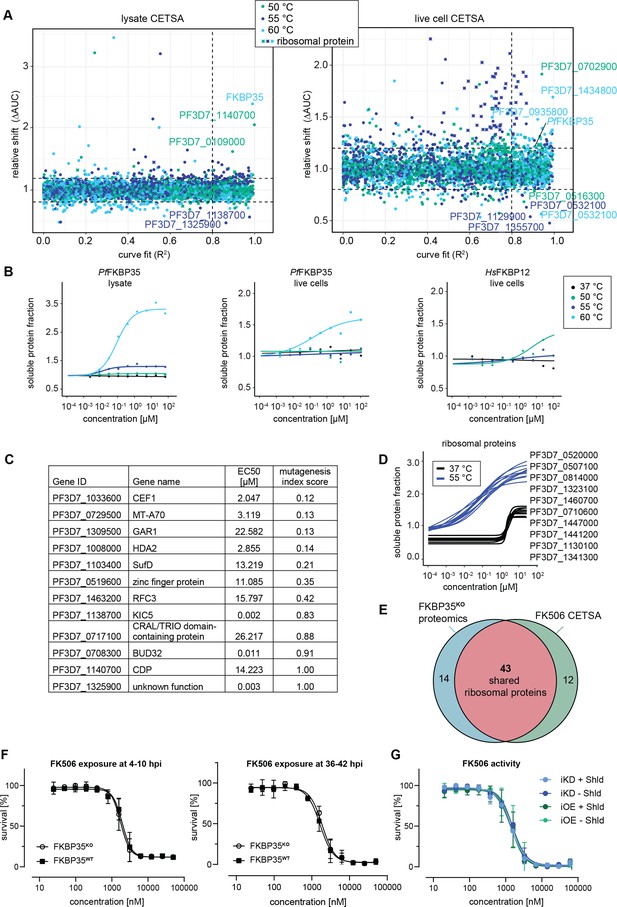
Cellular thermal shift assays (CETSA) identify FK506 interaction partners.
(A) CETSA results. The relative shift in protein abundance (area under the curve of heat-challenged samples normalized against the non-denaturing control, ΔAUC) as a function of R2 (goodness of fit as a measure for the dose–response effect). The dashed lines indicate an arbitrary cutoff of three times the median absolute deviation (MAD) of each dataset and R2 = 0.8. Three different temperatures were used for protein denaturation (indicated by colors). (B) Dose-dependent stabilization of P. falciparum and human FKBPs detected using lysate and live cell CETSA. Bullets represent the abundance of non-denatured proteins in response to increasing FK506 concentrations under thermal challenge relative to the DMSO control. Colors indicate the temperatures used for heat challenge; the black curve represents the non-denaturing (37°C) control. (C) Proteins with the most pronounced stability shift under thermal challenge in the lysate CETSA experiment are listed. The EC50 is the concentration at which the half-maximal effect of the FK506-induced thermal stabilization is reached. The mutagenesis index score determined by Zhang et al., 2018 is a measure to predict if a protein is essential for asexual growth: 0 = essential, 1 = non-essential. (D) Fitted live cell CETSA curves of 10 representative ribosomal proteins at 37°C and 55°C. Ribosomal proteins show an FK506 dose-dependent increase in the soluble protein fraction. (E) Venn diagram showing the overlap between ribosomal proteins deregulated in the PfFKBP35 knock-out cell line (less abundant in G2 FKBPKO trophozoites determined by proteomics) and the ribosomal proteins stabilized by FK506 in live cell CETSA. The number of factors identified in either or both assays is indicated. (F) Dose–response effect of FK506 on FKBP35KO and FKBP35WT parasites. The knock-out was induced at 0–6 hpi and parasites were exposed to FK506 at 4–10 hpi (left panel) and 36–42 hpi in G1 (right panel). Parasite survival was measured after reinvasion (10–16 hpi) in G2. n = 3, error bars represent the standard error of the mean. (G) Dose–response effect of FK506 on NF54/iKD-FKBP35 and NF54/iOE-FKBP35 parasites in the presence and absence of Shield-1 (Shld). Parasites were split at 0–6 hpi and subsequently treated with Shld or the vehicle control EtOH. Data points represent the mean of three and two independent biological replicates for the NF54/iOE-FKBP35 and NF54/iKD-FKBP35 cell line, respectively. Error bars represent the standard error of the mean.
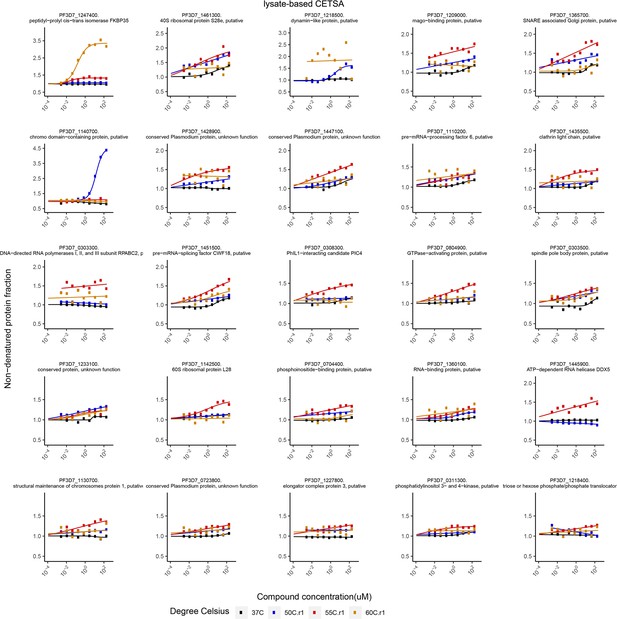
Putative FK506 interaction partners identified by protein lysate-based cellular thermal shift assays (CETSA).
Proteins with an area under the curve (∆AUC in protein lysate-based CETSA) greater than two median absolute deviations and an R2 greater than 0.8 are listed. Bullets represent the abundance of non-denatured proteins in response to increasing FK506 concentrations under thermal challenge relative to a DMSO control. Colors indicate the temperatures used for protein denaturation; the black curve represents the non-denaturing control.
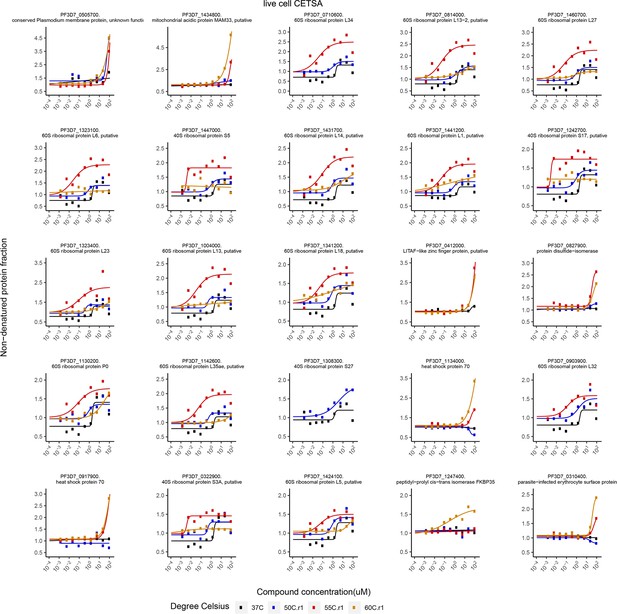
Putative FK506 interaction partners identified by live cell cellular thermal shift assays (CETSA).
Proteins with an area under the curve (∆AUC in live cell CETSA) greater than two median absolute deviations and an R2 greater than 0.8 are listed. Bullets represent the abundance of non-denatured proteins in response to increasing FK506 concentrations under thermal challenge relative to a DMSO control. Colors indicate the temperatures used for heat challenge; the black curve represents the non-denaturing control.
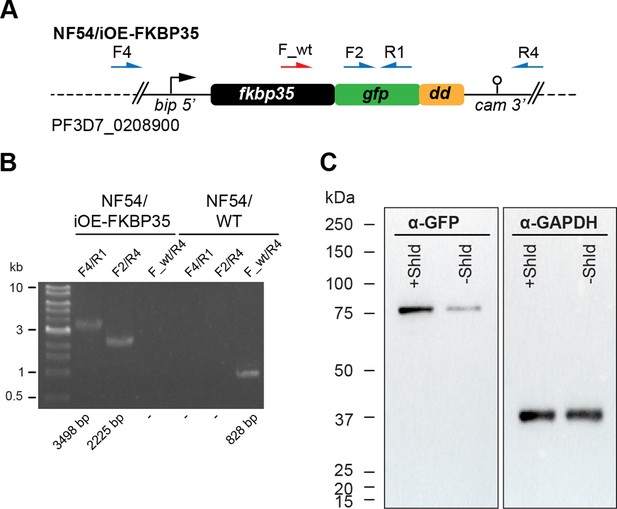
Characterization of NF54/iOE-FKBP35 parasites.
(A) Schematic of the modified p230p locus (PF3D7_0208900) in NF54/iOE-FKBP35 parasites, in which the overexpression cassette was introduced. This cassette consists of a bip promoter upstream of coding sequences of fkbp35, green fluorescent protein (gfp) and destabilization domain (dd), followed by the cam 3′ region. Names and binding sites of the primers used for diagnostic PCRs are indicated. The primer F_wt only binds the unmodified p230p sequence. (B) Diagnostic PCRs performed on gDNA of NF54/iOE-FKBP35 parasites to confirm correct editing of the locus. PCRs performed on NF54/WT gDNA served as control. Numbers at the bottom indicate the expected band sizes. (C) Expression levels of the FKBP35-GFP-DD fusion protein were assessed by western blot. Parasite cultures were split at 0–6 hpi and treated with Shield or the vehicle control EtOH. Protein lysates were collected 40 hr later. α-GAPDH served as loading control. The expected size of the FKBP35-GFP-DD fusion protein is 74.2 kDa.
-
Figure 4—figure supplement 3—source data 1
Agarose gel electrophoresis of peqGREEN-stained DNA and SDS-polyacrylamide gel electrophoresis (SDS-PAGE) of protein samples.
- https://cdn.elifesciences.org/articles/86975/elife-86975-fig4-figsupp3-data1-v1.zip
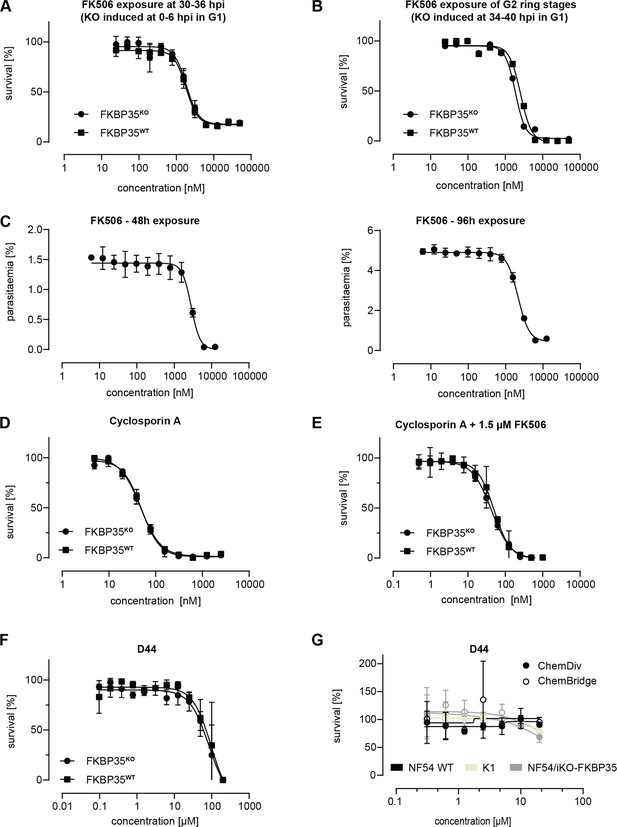
FKBP-targeting drugs fail at inhibiting parasite PfFKBP35.
(A) Dose–response effect of FK506 on FKBP35KO and FKBP35WT parasites. The knock-out was induced at 0–6 hpi and parasites were exposed to FK506 at 30–36 hpi. Survival rates were measured by flow cytometry in G2. n = 3, error bars represent the standard error of the mean. (B) Dose–response effect of FK506 on FKBP35KO and FKBP35WT parasites. The knock-out was induced at 34–40 hpi in G1. Ring stage parasites of G2 were exposed to FK506 for 48 hr and survival rates were measured by flow cytometry in G3. Data points represent survival rates measured in a single experiment performed in technical duplicates. (C) Dose–response effect of FK506 on FKBP35WT parasites over two IDCs. Parasites at 0–6 hpi were exposed to FK506 and incubated for 96 hr. Parasitemia was measured by flow cytometry after 48 hr and 96 hr. Medium containing the appropriate drug concentrations was changed after 48 hr. n = 3, error bars represent the standard error of the mean. (D) Dose–response effect of cyclosporin A on FKBP35KO and FKBP35WT parasites. Cylocsporin A, a known inhibitor of calcineurin when complexed with FKBP in other organisms (Dumont, 2000), did not alter drug susceptibility between FKBP35KO and FKBP35WT parasites, suggesting that PfFKBP35 is not involved in regulating calcineurin activity. The knock-out was induced at 0–6 hpi. Parasites were exposed to cyclosporin A for 48 hr. n = 3, error bars represent the standard error of the mean. (E) Dose–response effect of cyclosporin A in combination with a constant FK506 concentration on FKBP35KO and FKBP35WT parasites. The knock-out was induced at 0–6 hpi. Parasites were subsequently exposed to cyclosporin A and 1.5 µM FK506 for 48 hr. n = 3, error bars represent the standard error of the mean. (F) Dose–response effect of D44 on FKBP35KO and FKBP35WT. The knock-out was induced at 0–6 hpi and parasites were exposed to D44 for 48 hr thereafter. Survival rates were determined using flow cytometry. n = 3, error bars represent the standard error of the mean. (G) Dose–response effect of D44 on NF54 WT, K1, and NF54/iKO-FKBP35 parasites after 72 hr exposure. Survival rates were determined using a hypoxanthine incorporation assay. Data points represent the mean of two technical replicates. Error bars represent the standard error of the mean.
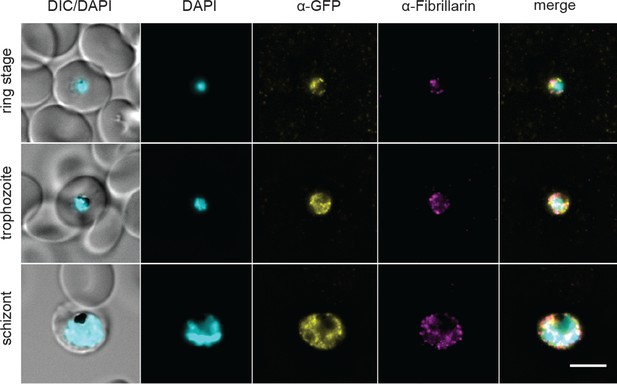
PfFKBP35 localizes near fibrillarin-dense nuclear regions.
Localization of PfFKBP35 (GFP) and fibrillarin in NF54/iKD-FKBP35 parasites at the ring, trophozoite and schizont stage assessed by immunofluorescence. DNA was stained with DAPI. Representative images are shown. Scale bar: 5 µm. DIC, differential interference contrast.
Additional files
-
Supplementary file 1
Mass spectrometry data comparing the proteomes of FKBP35KO and FKBP35WT parasites.
- https://cdn.elifesciences.org/articles/86975/elife-86975-supp1-v1.xlsx
-
Supplementary file 2
RNA-seq data comparing the transcriptomes of FKBP35KO and FKBP35WT parasites.
- https://cdn.elifesciences.org/articles/86975/elife-86975-supp2-v1.xlsx
-
Supplementary file 3
CETSA data showing the relative abundance of soluble proteins under non-denaturing and denaturing conditions in response to increasing FK506 concentrations.
- https://cdn.elifesciences.org/articles/86975/elife-86975-supp3-v1.xlsx
-
Supplementary file 4
Oligonucleotides used in this study.
- https://cdn.elifesciences.org/articles/86975/elife-86975-supp4-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/86975/elife-86975-mdarchecklist1-v1.docx





