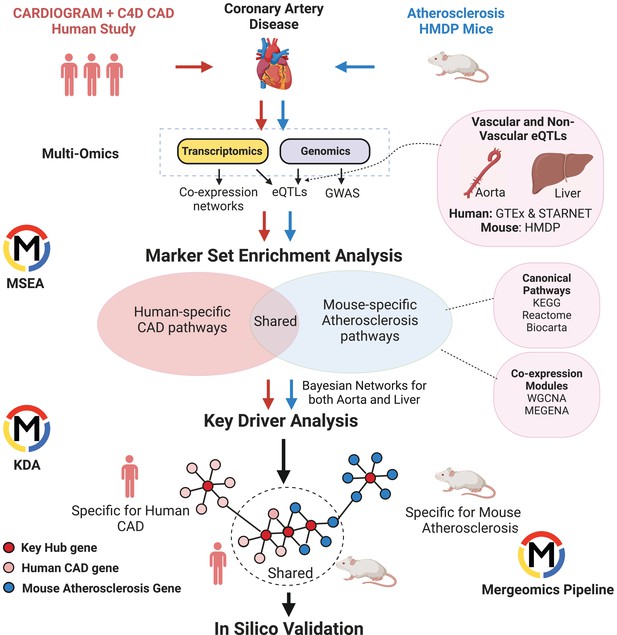
Shared and distinct pathways and networks genetically linked to coronary artery disease between human and mouse
Figures
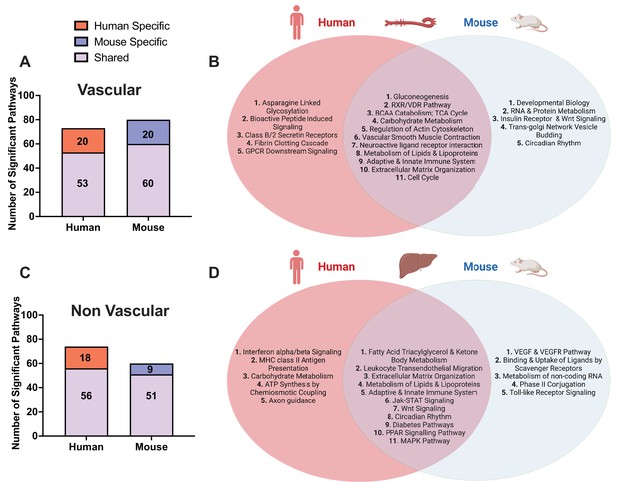
Shared and species-specific biological pathways.
(A) Bar plot highlighting the number of significant pathways shared and unique between mouse and human in vascular tissue (FDR<0.05). (B) Venn diagram highlighting the top shared and unique significant pathways between mouse and human in vascular tissue. (C) Bar plot highlighting the number of significant pathways shared and unique between mouse and human in non-vascular tissue (liver) (FDR<0.05). (D) Venn diagram highlighting the top shared and unique significant pathways between mouse and human in non-vascular tissue (liver).
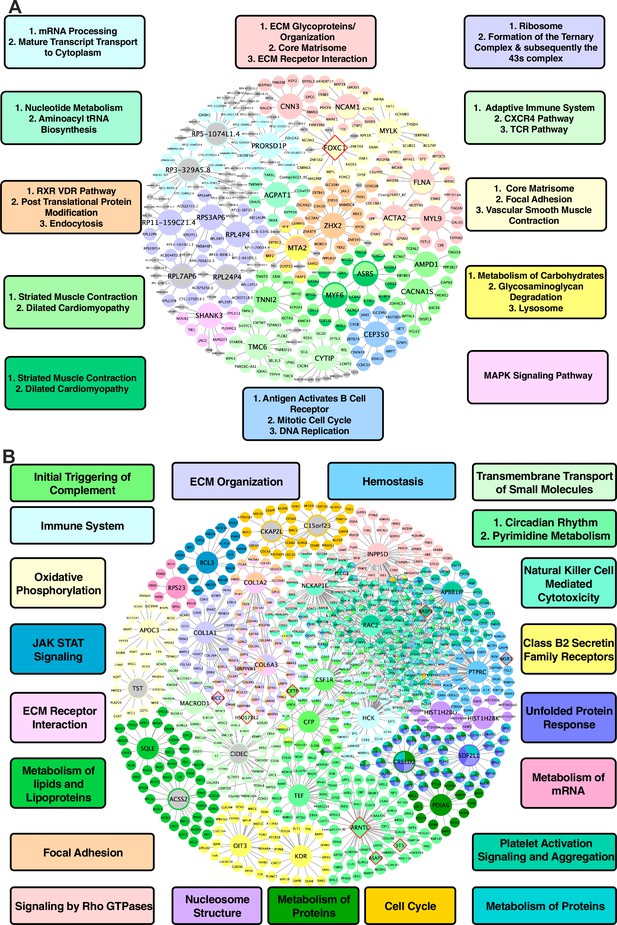
Shared networks between mice and humans.
(A) Vascular tissue gene regulatory network shared between mice and humans (B) Non-vascular gene regulatory network shared between mice and humans. Each node is color coded based on the pathway/module that the genes are derived from with larger nodes signifying key driver genes. Red border diamonds represent CAD GWAS hits uncovered after the CARDIOGRAM+C4D GWAS (2016 onwards) and pink border diamonds represent CAD GWAS hits prior to the CARDIOGRAM+C4D GWAS.
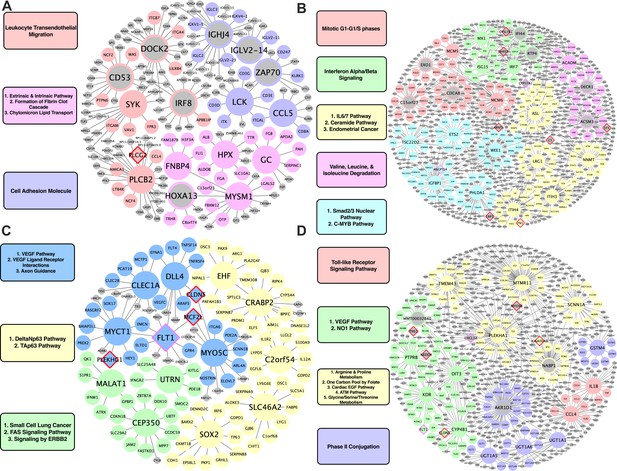
Species-specific networks.
(A) Human vascular gene regulatory network, (B) Human non-vascular gene regulatory network, (C) Mouse vascular gene regulatory network, (D) Mouse non-vascular gene regulatory network. Each node is color coded based on the pathway/module that the genes are derived from with larger nodes signifying key driver genes. Red border diamonds represent CAD GWAS hits uncovered after the CARDIOGRAM+C4D GWAS (2016 onwards) and pink border diamonds represent CAD GWAS hits prior to the CARDIOGRAM+C4D GWAS.
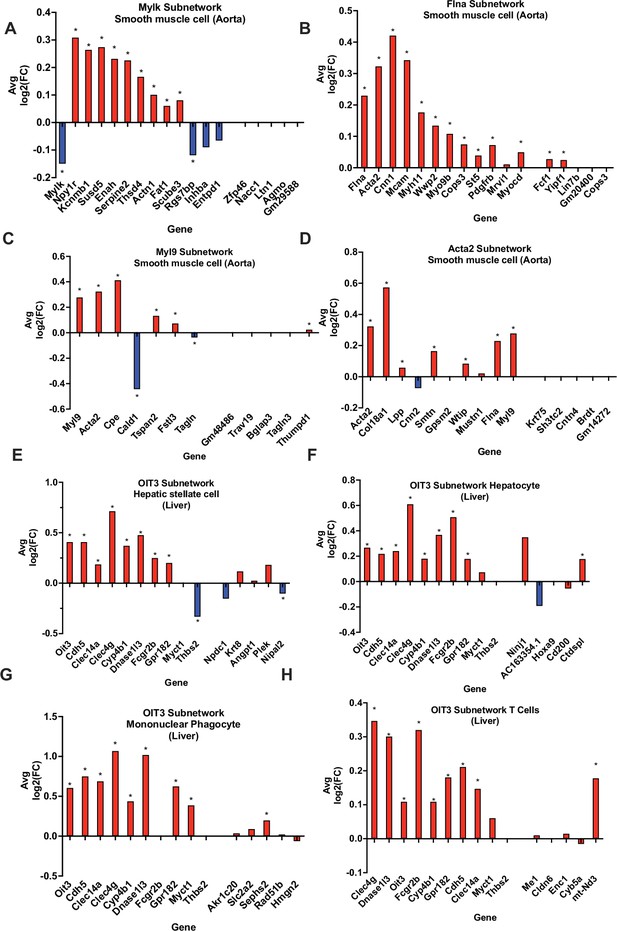
In silico validation of select KDs using single cell RNA-sequencing data.
(A-D) Aorta KDs and subnetwork genes, where AvgLog2(FC) is representing gene expression change between atherogenic diet and chow diet groups. (E-H) Liver OIT3 KD and subnetwork genes, where AvgLog2(FC) is representing gene expression changes between NASH and control. The last 5 genes in each plot are randomly selected negative control genes. We utilized a Wilcoxon rank sum test with bonferroni correction to derive significance for each gene. * represents a p<0.05.
Additional files
-
Supplementary file 1
Co-expression module numbers, MSEA pathway results and KD correlations with CAD clinical traits.
(A) The number of modules derived from WGCNA and MEGENA from vascular and non-vascular tissues. (B) Marker Set Enrichment Analysis (MSEA) results with an FDR <5% for Human CAD GWAS and Mouse atherosclerosis GWAS in vascular and non-vascular tissue. (C) Second round of MSEA results for the independent supersets with an FDR <5% and cross-species comparison. (D) Key driver correlations in vascular tissue with clinical traits relevant to CAD. (E) Key driver correlations in non-vascular tissue with clinical traits relevant to CAD.
- https://cdn.elifesciences.org/articles/88266/elife-88266-supp1-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/88266/elife-88266-mdarchecklist1-v1.docx