Septin 7 interacts with Numb to preserve sarcomere structural organization and muscle contractile function
Figures

Variants of Numb.
(A) The PCR strategy used to detect Numb splice variants is shown. Red arrows depict the locations of the forward and reverse primers used. Exons (black boxes) are numbered 1–10. (B) cDNA was prepared using total RNA isolated from denervated (Dn) or control (C) gastrocnemius muscle and from undifferentiated (U) and differentiated (D) C2C12 myoblasts and amplified using primers spanning exons 1–6 (upper panel) or 8–10 (lower panel). PCR products were analyzed by agarose gel electrophoresis. Gastrocnemius muscles were removed at 7 d after left sciatic nerve transection. Differentiated C2C12 cells were harvested at 5 d after transferring cells to differentiation media. PTBL and PTBs refer to phosphotyrosine binding domain with or without exon 3; PRRs: proline-rich region, s refers to the PRR without exon 9, which is the only form present in muscle and myogenic cells.
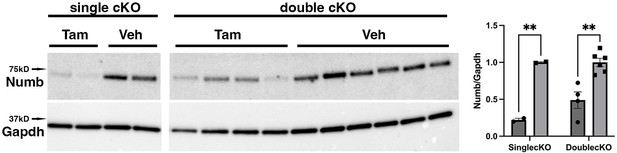
Representative Western blot showing downregulation of Numb levels in extensor digitorum longus (EDL) muscle from animals used for ex vivo physiology experiments.
Data were analyzed by unpaired t-test. **p<0.01. Data as mean +/- STD. The lanes were cut from the same blot.
-
Figure 2—source data 1
Original western blot analysis of data shown Figure 2 with anti-Numb (top image) and anti-GAPDH (bottom image).
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig2-data1-v1.pdf
-
Figure 2—source data 2
Figure 2 original western blot analysis (anti-Numb, top, and anti-GAPDH, bottom) with the bands shown in Figure 2 highlighted and sample labels.
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig2-data2-v1.pdf
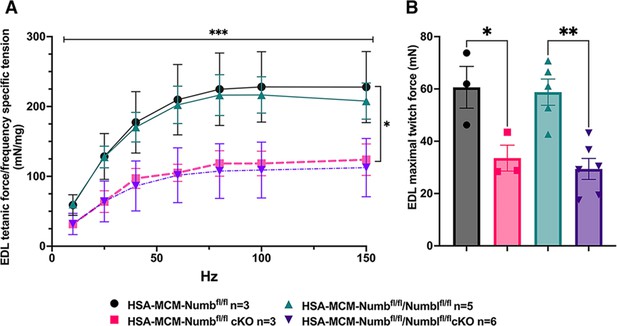
Effect of Numb or Numb/Numbl conditional knockout (cKO) on contractile function of extensor digitorum longus (EDL) muscle during ex vivo physiological testing.
(A) Specific tension generated during tetanic contraction at the indicated frequencies is shown for weight normalized EDL muscle harvested from HSA-MCM/Numb(f/f) or HSA-MCM/Numb(f/f)/Numbl(f/f) mice at 14 d after starting injections of tamoxifen or vehicle. Statistical analysis was performed with repeated-measure ANOVA followed by Sidak’s multiple-comparison test. F = 28.29, DFn = 6, DFd = 22; ***p<0.001, force × frequency interaction ***p<0.001; (B) Maximum force generated during a single twitch is shown. Statistical analysis was performed with one-way ANOVA with Tukey’s post hoc test (F = 10.07, DFn = 3, DFd = 13). *p<0.05; **p<0.01. N = 3–6. Data expressed as mean +/- STD.
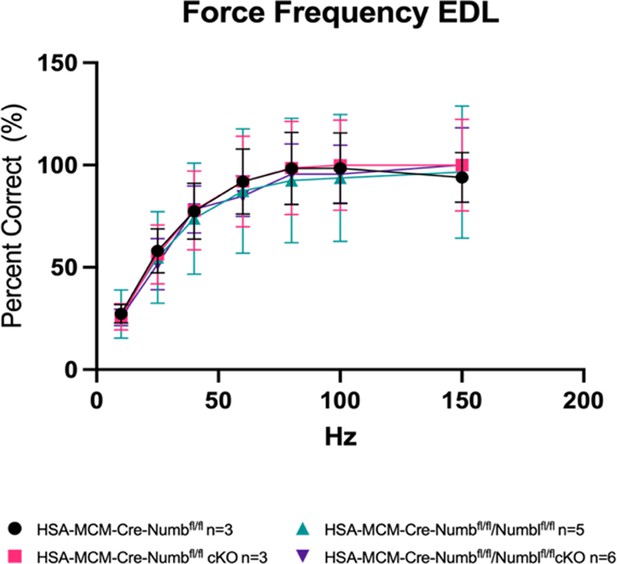
Force–frequency data from Figure 3 were replotted for each group as percent maximal force produced for that group.
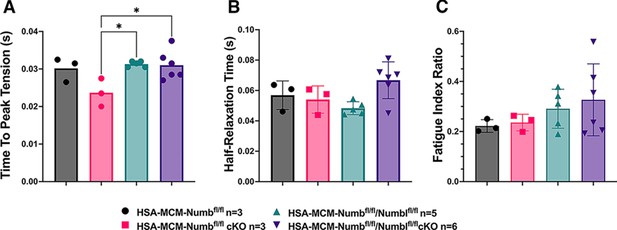
Time to peak tension (A), half-relaxation time (B), and fatigue index (C) are shown for HSA-MCM/Numb(f/f) and double (HSA-MCM/Numb(f/f)/Numbl(f/f) mice) at 14 d after starting injections of tamoxifen or vehicle.
Data were analyzed by one-way ANOVA with Tukey’s post hoc test. (A) F = 4.561, DFn = 3, DFd = 13; *p<0.05; (B) F = 3.679, DFn = 3, DFd = 13; (C) F = 0.9772, DFn = 3, DFd = 13. N = 3–6. Datas expressed as mean +/- STD.
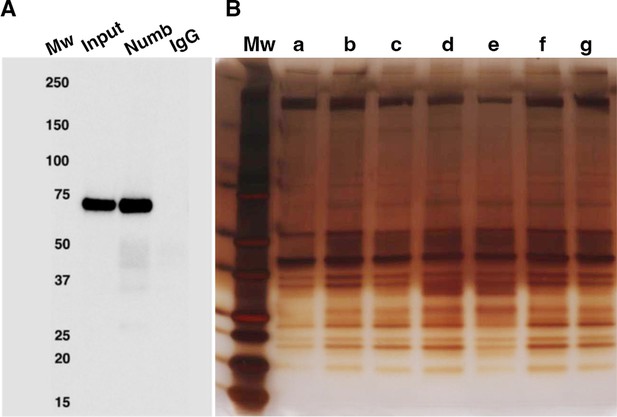
Numb immunoprecipitation.
(A) Representative western blot of Numb immunoprecipitated from differentiated C2C12 myotubes. Input: original C2C12 lysate; Numb: goat anti-Numb IgG, control IgG: normal goat IgG. The blot was probed with a rabbit monoclonal anti-Numb antibody (B). Proteins recovered in each of the seven immunoprecipitations used for liquid chromatography coupled with mass spectrometry (LC/MS/MS) analysis were resolved by SDS-PAGE and visualized by silver staining.
-
Figure 5—source data 1
Original images of western blot analysis shown in Figure 5A (anti-Numb, top, and anti-GAPDH, bottom).
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig5-data1-v1.pdf
-
Figure 5—source data 2
Original images of western blot analysis (anti-Numb, top, and anti-GAPDH, bottom) with the bands shown in Figure 5A highlighted and sample labels.
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig5-data2-v1.pdf
-
Figure 5—source data 3
Original image of silver-stained IP gel shown in Figure 5B.
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig5-data3-v1.pdf
-
Figure 5—source data 4
Original image of silver-stained IP gel with the bands shown in Figure 5B highlighted and sample labels.
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig5-data4-v1.pdf
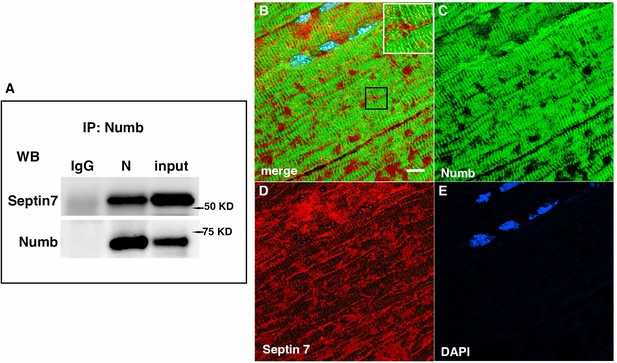
Western blot showing a representative co-immunoprecipitation of Septin 7 by anti-Numb antibody.
(A) IgG: immunoprecipitation with control Ig; N: immunoprecipitation with anti-Numb; input: original C2C12 lysate. The upper panel was probed with anti-Septin 7, the lower panel with anti-Numb. (B–D) Immunolocalization of Numb and Septin 7 in muscle. Longitudinal cryosections of TA muscle from C57Bl6 mice were immunostained for Numb and Septin 7: (B) merged image; (C) anti-Numb antibody (green); (D) anti-Septin 7 antibody (red); (E) DAPI staining of nuclei (blue). Scale bar, 10 μm. The inset in (A) shows higher magnification (×2.3) of the area in the black square.
-
Figure 6—source data 1
Original images of western blot analysis shown in Figure 6A (anti-Septin, left, and anti-Numb, right).
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig6-data1-v1.jpg
-
Figure 6—source data 2
Figure 6A images of western blot analysis (anti-Septin 7, left, and anti-Numb, right) with the bands shown in Figure 6A highlighted and sample labels.
- https://cdn.elifesciences.org/articles/89424/elife-89424-fig6-data2-v1.pdf
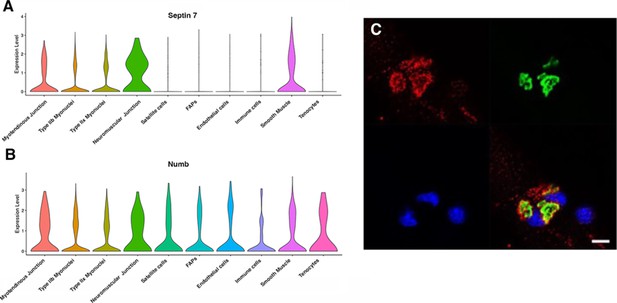
Septin 7 is enriched at neuromuscular junctions.
(A, B) Violin plots generated by Myoatlas showing abundance of Septin 7 (A) and Numb (B) mRNA in nuclei of TA muscle from 5-month-old mice (Dho et al., 1999). (C) Representative confocal microscopy image of the neuromuscular junction of an isolated myofiber immunostained with anti-Septin 7 (red) and with a fluorescently tagged α-bungarotoxin (green); nuclei were stained with DAPI (blue). Scale bar, 10 μm.

Representative confocal images of isolated mouse hindlimb fibers immunostained with anti-Numb (green) (B, E) and anti-Septin 7 antibodies (red) (C–F); merge: merged images (A, D).
Nuclei were labeled with DAPI (blue). (A–C) Myofiber from a control mouse (vehicle treated); (D–F) myofiber from a Numb cKO mouse (tamoxifen treated). Scale bar, 10 μm.
Tables
Numb-interacting proteins identified by mass spectrometry.
| UniProt ID | Protein | Abundance ratio (Numb/control) | p-Value (control vs Numb) | Control count(out of 7) | Numb count (out of 7) |
|---|---|---|---|---|---|
| APCA4_MOUSE | Anaphase-promoting subunit 4 | ∞ | 0 | 0 | 7 |
| NIPA_MOUSE | Nuclear-interacting partner of Alk | ∞ | 0 | 0 | 7 |
| PDE4D_MOUSE | cAMP specific 3–5-cyclic phosphodiesterase 4D | ∞ | 0 | 0 | 6 |
| WNK1_MOUSE | Serine-Threonine kinase Wnk1 | 289 | 5.4E–4 | 4 | 7 |
| SEPT7_MOUSE | Septin 7 | 82.9 | 3.6E–3 | 4 | 7 |
| APC 10_MOUSE | Anaphase-promoting complex subunit 10 | ∞ | 0 | 0 | 5 |
| SEPT10_MOUSE | Septin 10 | ∞ | 0 | 0 | 4 |
| HOME3_MOUSE | Homer protein homologue | ∞ | 0 | 0 | 4 |
| TRAF7_MOUSE | E3 ubiquitin ligase Traf7 | ∞ | 0 | 0 | 4 |
| RUVB1_MOUSE | RuvB-like 1 | ∞ | 0 | 0 | 5 |
| SEPT2_MOUSE | Septin 2 | 379 | 0.01 | 5 | 7 |
| SEPT9_MOUSE | Septin 9 | 148 | 0.01 | 4 | 7 |
-
Table 1—source data 1
MS/MS data for Septin 7 from representative IP sample (67) showing peptides abundance and example mass spectra.
- https://cdn.elifesciences.org/articles/89424/elife-89424-table1-data1-v1.pdf
-
Table 1—source data 2
MS/MS data for cAMP-specific 3,5 cyclic phosphodiesterase 4D (PDE4D) from representative IP sample (520) showing peptides abundance and example mass spectra.
- https://cdn.elifesciences.org/articles/89424/elife-89424-table1-data2-v1.pdf
-
Table 1—source data 3
MS/MS data for nuclear -interacting protein of ALK (NIPA) from representative IP sample (1617) showing peptides abundance and example mass spectra.
- https://cdn.elifesciences.org/articles/89424/elife-89424-table1-data3-v1.pdf
-
Table 1—source data 4
MS/MS data for anaphase-promoting complex subunit 2 (ANC2) from representative IP sample (67) showing peptides abundance and example mass spectra.
- https://cdn.elifesciences.org/articles/89424/elife-89424-table1-data4-v1.pdf
-
Table 1—source data 5
MS/MS data for anaphase-promoting complex subunit 4 (APC4) from representative IP sample (1617) showing peptides abundance and example mass spectra.
- https://cdn.elifesciences.org/articles/89424/elife-89424-table1-data5-v1.pdf
-
Table 1—source data 6
MS/MS data for anaphase-promoting complex subunit 10 (APC10) from representative IP sample (520) showing peptides abundance and example mass spectra.
- https://cdn.elifesciences.org/articles/89424/elife-89424-table1-data6-v1.pdf
Additional files
-
Supplementary file 1
List of MS/MS data for all the proteins for which peptides fragments were identified in the IP samples.
- https://cdn.elifesciences.org/articles/89424/elife-89424-supp1-v1.xlsx
-
Supplementary file 2
Manually curated annotation for the Numb-binding proteins identified by IP-MS/MS showing domains, function, and associated diseases including linkage to disorders of the skeleton.
- https://cdn.elifesciences.org/articles/89424/elife-89424-supp2-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/89424/elife-89424-mdarchecklist1-v1.pdf





