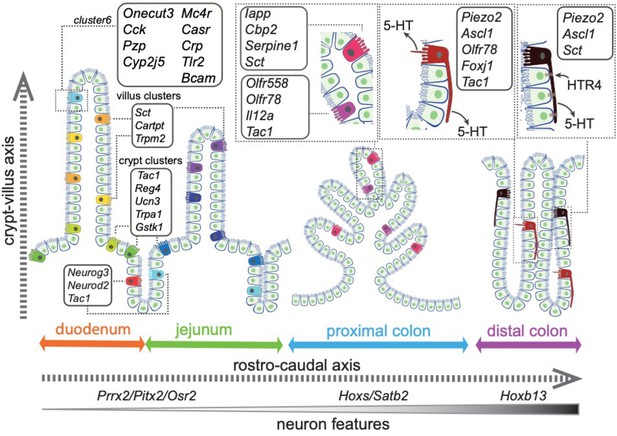
Stratification of enterochromaffin cells by single-cell expression analysis
Figures
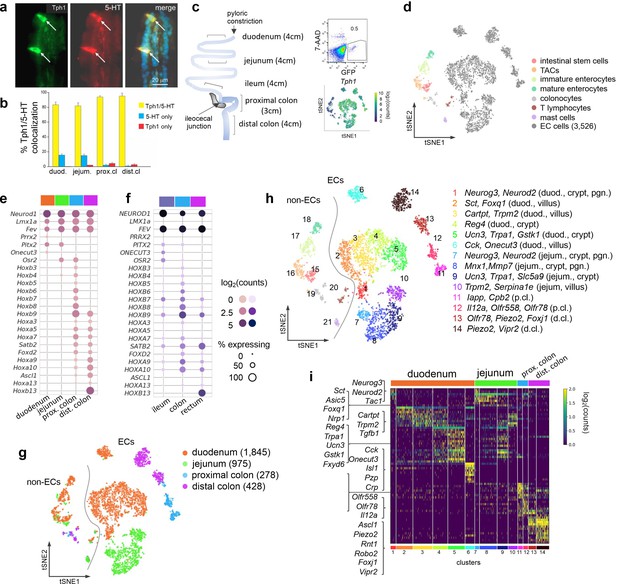
scRNA-seq identifies distinct intestinal EC cell clusters.
Dual IF staining of 5-hydroxytryptamine (5-HT) and GFP (representing Tph1) (a) and their quantification/colocalization in the indicated regions of Tph1-bacTRAP mice (b). Schematic showing the isolation and enrichment of GFP+ cells from the indicated regions of the GI tract for scRNA-seq (c). Upper right: Fluorescence-activated cell sorting (FACS) plot of dissociated gut epithelial cells from Tph1-bacTRAP mice. Gate shows GFP+ cells, which account for ~0.5% of total viable gut epithelial cells (7-AAD− cells). Lower right: t-SNE projection of all GFP+ cells superimposed on an expression heatmap of Tph1 in GFP+ cells. t-SNE projection of all GFP+ cells isolated from Tph1-bacTRAP mice in the second cell profiling experiment, from which 3526 EC cells were identified and subjected to further analysis (d). TACs: transit amplifying cells; EC cells: enterochromaffin cells. (e, f) Transcription factors (TFs) that are differentially expressed in the mouse EC cells isolated from Tph1-bacTRAP mice along the rostro-caudal axis presented by regions as indicated in the color bar (e). Expression data of the human orthologues of the same TFs were extracted from human gut mucosa dataset (GSE125970). Enteroendocrine (EEC) cells were selected and presented by regions (f). Size of the circles represents percentage of expression and the intensity of the circles represents aggregated expression of indicated TFs in cells partitioned by regions. (g) t-SNE projection of all GFP+ cells color-coded by their regions in the GI tract. Numbers of cells retained from each region are indicated in parentheses. Dashed line demarcates the separation of non-EC cells (including stem cells, TACs, immature enterocytes, mature enterocytes, colonocytes, T lymphocytes, and mast cells) versus EC cells. (h) tSNE projection of all GFP+ cells color-coded by clusters that were identified via the Louvain method. Fourteen clusters of EC cells (clusters 1–14) and 7 clusters of non-EC cells were identified (clusters 15–21, as described in (d)). Key marker genes are listed for each cluster. duod.: duodenum, jejum.: jejunum, pgn: progenitor. (i) Heatmap of the top 5–10 signature genes for each cluster presented as normalized log2(counts) in all EC cells (in columns). Color-coded bar at the bottom represents the clusters identified in (h).
-
Figure 1—source data 1
Expression data for specific detected genes.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig1-data1-v1.xlsx
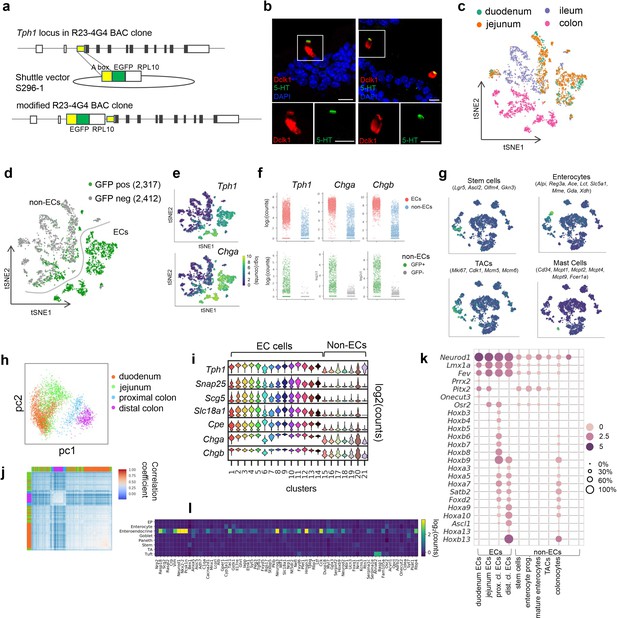
scRNA-seq identifies distinct intestinal enterochromaffin (EC) cell clusters.
(a) Schematic of Tph1-bacTRAP allele generation. A R23-4G4 BAC clone containing the Tph1 locus was used. Yellow box: ‘A box’, a 420-bp fragment immediately upstream of start codon of Tph1 that was used for homologous recombination. A sequence encoding a fusion protein of EGFP and RPL10 was cloned immediately downstream of the A box. (b) Dual IF staining of 5-hydroxytryptamine (5-HT) and Dclk1 (tuft cell marker) in the duodenum of wild-type mice. Scale bar: 10 µm. (c, d) t-SNE projections of all cells (4729) from single-cell profiling capturing both GFP+ (2317) and GFP− (2412) cells from the indicated regions of Tph1-bacTRAP mice. Single cells are color-coded by regions in (c) or by GFP positivity in (d). (e) t-SNE projections of cells shown in (c, d) superimposed on heatmaps representing normalized counts for the indicated genes. (f) Comparison of Tph1, Chga, and Chgb expression in EC cells versus non-EC cells (upper) or in the GFP+ versus GFP− cells among non-EC cells (lower). (g) t-SNE projection of cells from the second profiling capturing GFP+ cells from the duodenum, jejunum, and proximal and distal colon as presented in Figure 1d, g, h. Superimposed heatmaps represent the aggregated transcript counts of the indicated marker genes for each identified cell type. TACs: transit amplifying cells. (h) Principal component analysis (PCA) plot of EC cells (as shown in Figure 1d, g, h) from the second profiling and color-coded by regions. (i) Violin plots showing expression of well-established marker genes of EC cells in all clusters. (j) Unsupervised hierarchical clustering of EC cells based on transcriptome similarity (correlation coefficient) computed from the top 800 genes extracted from PC1 and PC2 (as shown in (h)). Color bars indicate the regions of EC cells. (k) Transcription factors (TFs) differentially detected along the rostro-caudal axis are presented in all cells (as shown in Figure 1d, g, h) partitioned by cell types or by regions (for ECs). Size of the circles represents percentage of expression and intensity of the circles represents aggregated expression of indicated TFs. (l) Signature genes of the EC clusters (as shown in Figure 1i) showing enrichment in EEC cells but not in the other epithelial cell types of the intestine. Heatmap illustrates median expression of the indicated genes extrapolated from GEO dataset GSE92332. EP: early progenitor, TA: transit amplifying cell.
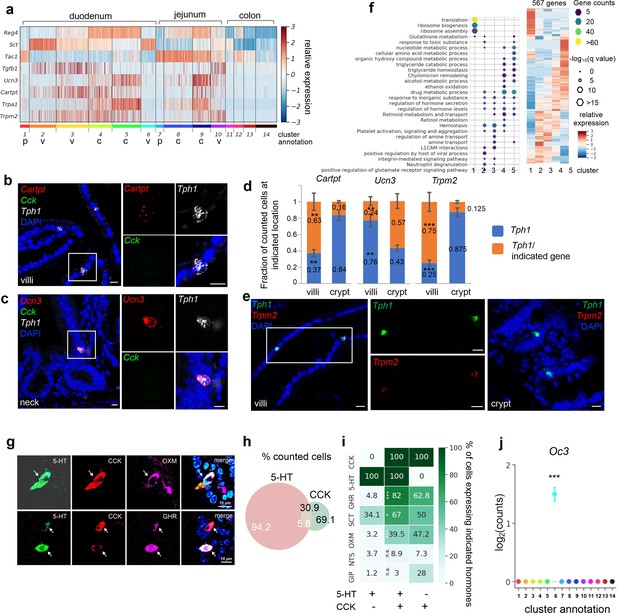
SI enterochromaffin (EC) cells are predicted to switch sensors and hormone compositions along the crypt–villus axis.
(a) Heatmap of representative genes with differential expression patterns between clusters annotated as crypt or villus. Relative gene expression (z-score) is shown across all single EC cells. Color-coded bar at the bottom represents the clusters. p: progenitor, v: villus, c: crypt. smRNA-FISH of Cartpt, Tph1, and Cck (b); Ucn3, Tph1, and Cck (c); or Trmp2 and Tph1 (e). The boxed regions are enlarged on the right and split into individual channels. Data shown are representative examples from three independent mice. Scale bars: 10 µm. (d) Quantitation of Cartpt, Ucn3, and Trmp2 positivity in the Tph1+ cells in the crypt versus villus. For each quantitation, 130–160 Tph1+ cells were counted from the crypt or villus of the duodenum in three independent mice. **p < 0.01, ***p < 0.001; unpaired two-tailed Student’s t-test comparing the fractions in the villi against those in the crypt. (f) Gene ontology analysis of the 567DE genes identified in the duodenal clusters. DE genes were determined by false discovery rate (FDR) <10–10 against every other cluster (among clusters 1–5) based on Wilcoxon rank sum test and corrected by the Benjamini–Hochberg procedure. Relative expression (z-score) of DE genes is shown on the right, GO analysis of DE genes on the left. Size of the hexagons represents the q-value of enrichment after −log10 transformation, and the density represents the number of genes per GO term. Accumulative hypergeometric testing was conducted for enrichment analysis and Sidak–Bonferroni correction was applied to correct for multiple testing. Cells co-expressing 5-HT, CCK, and a third hormone product shown by IF staining, such as a Gcg product (GHR), oxyntomodulin (OXM, upper panel) and the Ghrl product, ghrelin (GHR, lower panel). Arrows point to the triple-positive cells. Note that the relative levels of hormones vary considerably among cells, as shown in the 5-HT, CCK, and ghrelin triple stain. Venn diagram showing co-expression of 5-HT and CCK based on IF staining with indicated antibodies. Numbers represent percentages out of all 5-HT+ cells (white) or CCK+ cells (black). Summary of 5-HT or CCK single-positive cells and 5-HT/CCK double-positive cells (by columns) producing a third hormone (by rows) as identified by IF staining. Heatmap and annotated numbers represent the percentage out of all cells in individual columns. ***p < 10–10, **p < 10–2 by hypergeometric tests against the 5-HT single-positive cells. (g–i) are based on IF staining for indicated peptides/hormones in three different animals. Point plot depicting the median expression of Oc3 in each cluster showing enrichment of Oc3 expression in cluster 6 (Cck+/Tph1+ cells) from the Tph1-bacTRAP dataset. Error bars represent upper and lower quantiles. ***p < 10–50; two-tailed Kolmogorov–Smirnov statistic between the observed Oc3 distribution versus the bootstrap-facilitated randomization control distribution (median of n = 500 randomizations shown in gray boxplots).
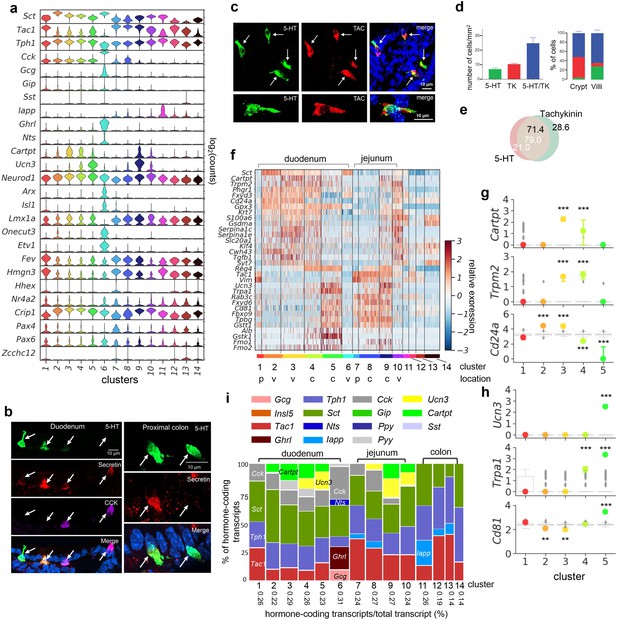
SI enterochromaffin (EC) cells are predicted to switch sensors and hormone compositions along the crypt–villus axis.
(a) Expression (log2(counts)) of peptide hormones along with Tph1 and transcription factors in the EC clusters (as shown in Figure 2a). (b) Co-expression of secretin, 5-hydroxytryptamine (5-HT) and CCK examined by IF staining. Co-expression of tachykinin and 5-HT examined by IF staining (c) and quantitated (d) in the jejunum. (e) Venn diagram showing co-expression of tachykinin and 5-HT in the jejunum based on IF staining. Numbers present the percentage out of all counted 5-HT+ cells (white) or tachykinin+ cells (black). (f) Heatmap of genes exhibiting differential enrichment along the crypt–villus axis. Relative expression (z-score) of indicated genes are shown across all single EC cells. Color-coded bar at the bottom represents the clusters. p: progenitor, v: villus, c: crypt. Representative gene enrichment in villus clusters (g) or crypt clusters (h). Note that cluster 4 shares some features in common with crypt and villus clusters and may represent cells at an intermediate stage of development. Point plots depict the median expression of indicated genes in each cluster and the error bars represent upper and lower quantiles. Clusters are represented by colors. The gray box plots depict the distributions of the median expression based on size-matched randomization control experiments (median of n = 500 randomizations). The boxes present the quantiles and whiskers show the rest of the distribution. ***p < 10–50, **p < 10–30; two-tailed Kolmogorov–Smirnov statistic between the observed gene distribution and randomized control distribution. (i) Marimekko chart showing the cluster-wise distribution of hormone-coding transcripts (color-coded) in the 14 clusters of EC cells. The width of each column represents the percentage, stated at the bottom, of all hormone-coding transcripts (plus Tph1) out of all transcripts. Height of each bar represents the percentage of indicated hormone-coding transcripts out of all hormone-coding transcripts.
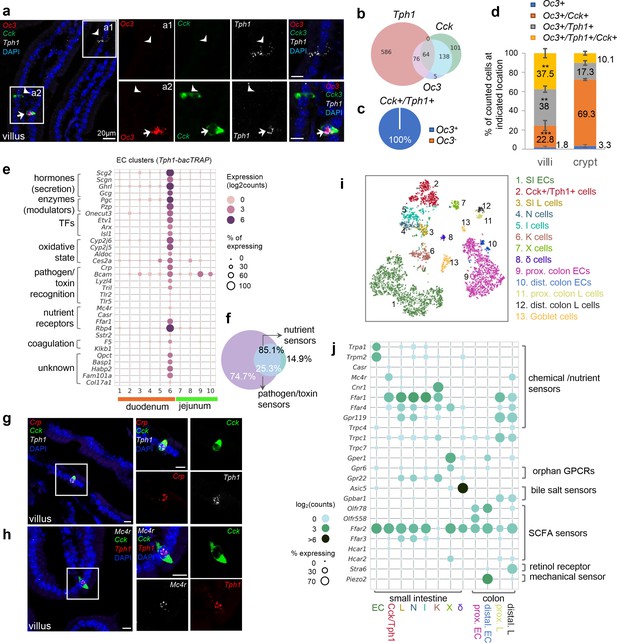
Cck, Oc3, and Tph1 specify an enterochromaffin (EC) subpopulation with a dual sensory signature.
(a) smRNA-FISH of Cck/Oc3/Tph1. Two boxed regions (a1, a2) are enlarged on the right and split into individual channels. An arrow points to a Cck+/Oc3+/Tph1+ cell in the villi and arrowheads point to Tph1 or Cck single-positive cells with no Oc3 expression in the villi. Images are representative from four different animals. Scale bars: 20 µm. (b) Venn diagram showing overlaps of Tph1-, Cck-, and Oc3-expressing cells based on smRNA-FISH performed on duodenal sections. Numbers of counted cells are annotated based on four sections per mouse in four different animals. (c) Pie chart showing all counted Cck+/Tph1+ cells partitioned by Oc3 positivity based on smRNA-FISH. (d) Quantitation of Oc3 expressing cells by smRNA-FISH in the villi versus in the crypts. **p < 0.01, ***p < 0.001, unpaired two-tailed Student’s t-test comparing the percentages in the villi against those in the crypts. (e) Signature genes identified in Cck+/Oc3+/Tph1+ cells by scRNA-seq in the Tph1-bacTRAP dataset. Size of the circles represents percentage of expression and intensity of the circles represents aggregated expression of indicated genes. (f) Venn diagram showing co-expression of two sets of genes summaried by the aggregated expression of genes associated with pathogen/toxin recognition (Crp, Lyz4, Tril, Tlr2, and Tlr5) versus genes associated with nutrient sensing and homeostasis (Mc4r and Casr) in the Tph1-bacTRAP dataset. smRNA-FISH of Crp, Cck, and Tph1 (g) and Mc4r, Cck, and Tph1 (h). The boxed regions are enlarged on the right and split into individual channels. Data are representative from three different animals. Scale bars: 10 µm. tSNE representation of single Neurod1-tdTomato+ cells isolated from the gut. Cells are color-coded by clusters that were identified via the Louvain method. Dot plot of genes coding G-protein-coupled receptors (GPCRs), TRP channels, SLC transporters, purinergic receptors, and prostaglandin receptors, identified in clusters as shown in (i). (j) Gene expression in specific endocrine cell types. Genes were selected if detected in >10% of cells in at least one of the indicated clusters and excluded if detected in >20% of enterocytes or colonocytes. Size of the circles represents percentage of expression and intensity of the circles represents aggregated expression of indicated genes partitioned by clusters.
-
Figure 3—source data 1
Gene expression data .
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig3-data1-v1.xlsx
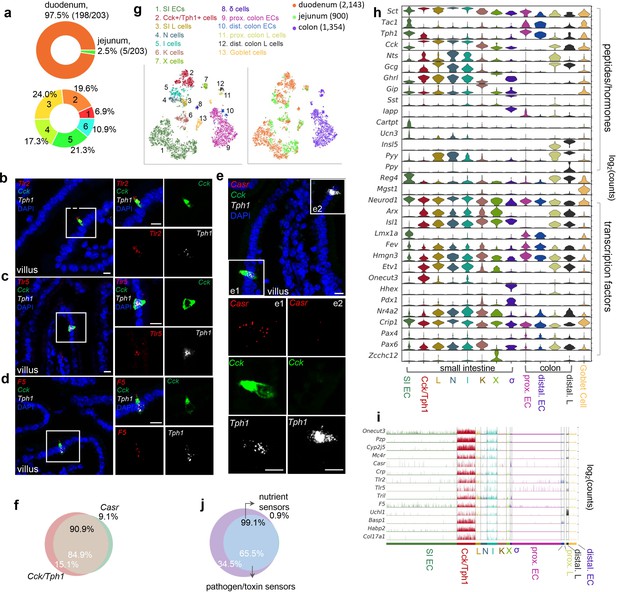
Cck, Oc3, and Tph1 specify an enterochromaffin (EC) subpopulation with a dual sensory signature.
(a) Upper: Regional composition of cluster 6 cells. Regions are color-coded. 203 cells were identified in cluster 6 in the Tph1-bacTRAP dataset. Lower: EC cells in duodenum are constituted by six clusters. Numbers and colors indicate clusters. smRNA-FISH of Tlr2, Cck, and Tph1 (b); Tlr5, Cck, and Tph1 (c); F5, Cck, and Tph1 (d); and Casr, Cck, and Tph1 (e). Two dashed boxes (e1, e2) show a representative Cck+/Tph1+ double-positive cell stained positive for Casr (e1) and a Tph1+ single-positive cell without Casr (e2). Scale bars: 10 µm. (f) Quantitation of Casr (black text) and double Cck+/Tph1+-positive cells (white text) in the duodenum based on smRNA-FISH. 64 cells were quantitated from four mice. (g) tSNE representation of single Neurod1-tdTomato+ cells isolated from the gut. Cells are color-coded by clusters identified via Louvain method (left) or by regions (right). (h) Expression (log2(counts)) of peptide hormones along with Tph1 and transcription factors in the enteroendocrine (EEC) clusters. (i) Expression of signature genes identified in the Cck+/Tph1+ cluster from the Neurod1-tdTomato dataset. Cells are shown in the columns and color-coded by clusters, and genes are shown in the rows. (j) Venn diagram showing the co-expression of two sets of genes summarized by the aggregated expression of genes associated with pathogen/toxin recognition (Crp, Lyz4, Tril, Tlr2, and Tlr5) versus genes associated with nutrient sensing and homeostasis (Mc4r and Casr) in the Neurod1-tdTomato dataset.
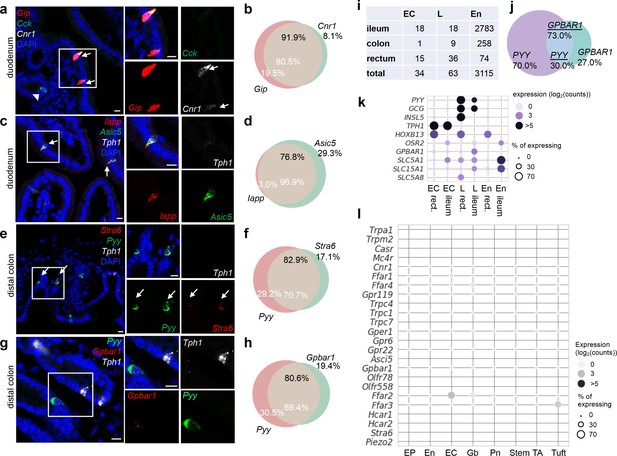
Distinct molecular sensors identified in EC cells versus other enteroendocrine (EEC) cells.
smRNA-FISH of Cnr, Gip, and Cck (a); Asic5, Iapp, and Tph1; (c) Stra6, Pyy, andTph1 (e); or Gpbar1, Pyy, and Tph1 (g) in wild-type mice. Venn diagrams of the corresponding quantitation based on smRNA-FISH experiment (b, d, f, h). For each quantitation, 105–156 cells were counted from three mice. Scale bars: 10 µm. (i) Breakdown of single human mucosal cells (GEO dataset GSE125970) in indicated categories. En: enterocytes. EC: enterochromaffin cells. L: Gcg- and/or Pyy-positive L cells. Since only 10 EEC cells were identified in the colon, they were not assessed in the subsequent analysis. (j) Venn diagram of GPBAR1 and PYY co-expression in human gut mucosal cells. Data were extracted from GEO dataset GSE125970. (k) Dot plot of key sensors differentially expressed in human ileal versus colonic L cells. Data were extracted from GSE125970. Single cells that are positive for TPH1, or positive for PYY or/and GCG are annotated as EC or L cells, respectively. En: enterocytes. Enterocytes and regions are annotated as in the provided metadata. Note, HOXB13 or OSR2 expression mark rectum or ileum origin, respectively. (l) Dot plot showing the expression level and percentage of sensors identified in EECs (Figure 3j). Data were extrapolated from GSE92332. Note, many of sensors (10/24) identified in our study were not detected in GSE92332, possibly due to the small number of EEC cells (258) in this dataset. EP: early progenitor, En: enterocyte, EC: enterochromaffin cell, Gb: goblet cell, Pn: Paneth cell, TA: transit amplifying cell.
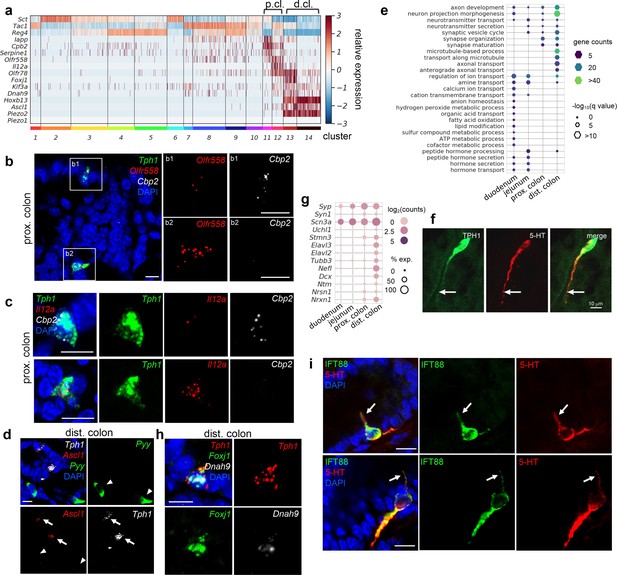
Distinct subpopulations of enterochromaffin (EC) cells are resolved in the colon.
(a) Heatmap showing the key signature genes identified in the colonic EC cells. Relative expression (z-score) of indicated genes are shown across all single EC cells based on the Tph1-bacTRAP dataset. Color bars at the bottom represent clusters and the ones on the top represent regions. p.cl.: proximal colon; d.cl.: distal colon. (b) smRNA-FISH of Tph1, Olfr558, and Cpb2 in the proximal colon. Two dashed boxes (b1, b2) are enlarged on the right to show two representative Tph1+ cells with preferential Cbp2 expression (b1) or Olfr558 expression (b2). (c) smRNA-FISH of Tph1, Il12a, and Cpb2 in the proximal colon. Two representative Tph1+ cells with preferential Cbp2 expression (upper) or Il12a expression (lower) are shown. (d) smRNA-FISH of Ascl1, Tph1, and Pyy in the distal colon. Arrows point to Ascl1 staining in Tph1+ cells. Arrowheads point to the absence of Ascl1 staining in Pyy+ cells. (e) GO analysis based on DE genes identified in regional EC cells. Hypothalamic neuron enriched genes were identified based on data from GEO dataset GSE74672, of which 60.7% genes were detected in gut EC cells. DE genes among the regional EC cells were determined by false discovery rate (FDR) <10–10 against every other region based on Wilcoxon rank sum test and corrected by Benjamini–Hochberg procedure. Size of the hexagons represents the q-value of enrichment after −log10 transformation, and the density represents the number of genes per GO term. Accumulative hypergeometric testing was conducted for enrichment analysis and Sidak–Bonferroni correction was applied to correct for multiple testing. (f) IF staining against 5-hydroxytryptamine (5-HT) and GFP (representing Tph1) in the distal colon. Note the prominent axon-like extension in the distal colon EC cell. (g) Dot plot showing representative neuron-related genes enriched in distal colon. (h) smRNA-FISH of Tph1, Foxj1, and Dnah9 in the distal colon. (i) IF staining against IFT88 and 5-HT in the distal colon. Data shown are two representative examples from four mice. Scale bars in panels b–d, f, i: 10 µm. Images in panels b–d, f are representative from three mice.
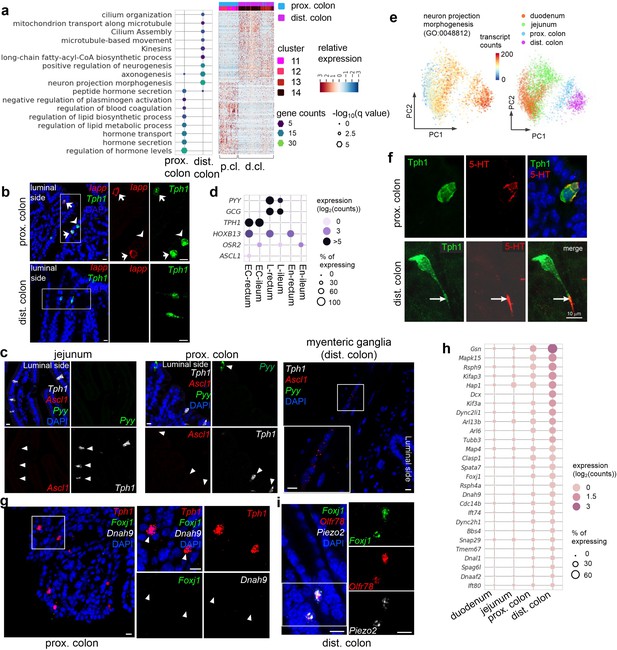
Distinct subpopulations of enterochromaffin (EC) cells are resolved in the colon.
(a) GO analysis of DE genes identified between the proximal and distal colonic EC cells (left). Data are based on the Tph1-bacTRAP dataset. DE genes: false discovery rate (FDR) <10–10, Wilcoxon rank sum test corrected with the Benjamini–Hochberg procedure for multiple hypothesis testing. Unsupervised hierarchical clustering of single cells (in columns) based on identified DE genes is shown on the right. (b) smRNA-FISH of Iapp and Tph1 in the proximal colon (upper) and distal colon (lower). Arrows point to Iapp/Tph1 double-positive cells. Arrowheads point to the absence of Iapp in some Tph1+ cells. (c) smRNA-FISH of Ascl1, Tph1, and Pyy in the jejunum, proximal colon and myenteric ganglia from distal colon. Arrowheads point to the absence of Ascl1 in either Tph1+ or Pyy+ cells. Associated with Figure 4d. (d) Dot plot showing ASCL1 enrichment in human rectal EC cells. Data were extracted from GEO dataset GSE125970. Single cells that are positive for TPH1, or positive for PYY and/or GCG are annotated as EC or L cells, respectively. En: enterocytes. Enterocytes and regions are annotated according to the provided metadata. HOXB13 and OSR2 expression mark the origins of rectum and ileum, respectively. Size of the circles represents percentage of expression and intensity of the circles represents aggregated expression of indicated genes. (e) Principal component analysis (PCA) plots color-coded by the aggregated transcript counts of the genes (based on the Tph1-bacTRAP dataset) in the indicated GO term (right) and by regions (left). (f) IF staining against 5-hydroxytryptamine (5-HT) and GFP (representing Tph1) in the proximal colon (upper) and the distal colon (lower). Note the axon-like extension is not observed in the proximal colonic EC cells. (g) smRNA-FISH of Tph1, Foxj1, and Dnah9 in the proximal colon. The boxed region is enlarged on the right. Arrowheads point to the absence of Foxj1 and Dnah9 in Tph1+ cells in the proximal colon. Associated with Figure 4h. (h) Dot plot showing expression levels and frequency of genes associated with ‘cilium assembly’ (R-MMU-5617833) and ‘cilium organization’ (GO:0044782) in EC cells along the rostro-caudal axis of the gut. (i) smRNA-FISH of Foxj1, Olfr78, and Piezo2 in the distal colon. The boxed region is enlarged on the right and split into individual channels. Scale bars in panels b, c, f, g, i: 10 µm. Images are representative from three mice.
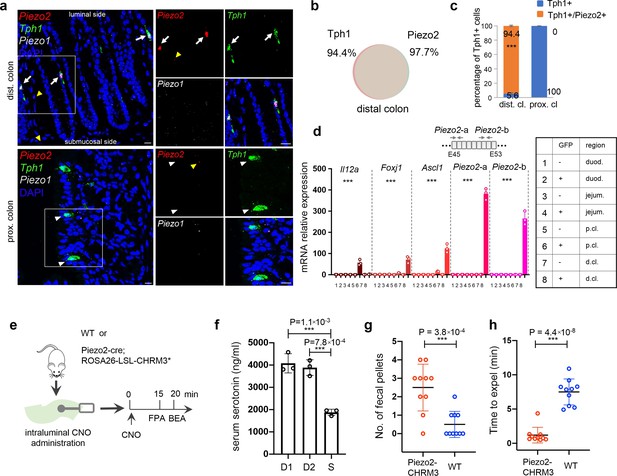
Piezo2 is highly enriched in distal colonic enterochromaffin (EC) cells that mediate colon motility.
(a) smRNA-FISH of Piezo2, Piezo1, and Tph1 in distal (upper) and proximal colon (lower). Arrows point to Piezo2/Tph1 double-positive cells in the distal colon. White arrowheads point to absence of Piezo2 in Tph1+ cells in the proximal colon. Yellow arrowheads point to sparse staining of Piezo2 within the lamina propria. Images are representative from four mice. Scale bars: 10 µm. (b) Venn diagram showing the co-expression of Tph1 and Piezo2 transcripts in the distal colon based on smRNA-FISH. Percentages are of double-positive cells out of respective single-positive cells. A total of 274 cells were quantitated from four mice. (c) Quantitation of Piezo2 and/or Tph1 expressing cells in distal and proximal colon based on smRNA-FISH. A total of 386 cells were quantitated from four wild-type mice. ***p < 0.001; unpaired two-tailed Student’s t-test of the double-positive fractions in distal colon versus proximal colon. (d) qPCR validation of regional enriched genes in EC cells. GFP− and GFP+ cells were sorted from the duodenum, jejunum, and proximal and distal colon in the Tph1-bacTRAP mice. Relative gene expression was computed relative to the values in the GFP− cells isolated from the duodenum (indicated as 1), after normalization by the aggregates of three house-keeping genes (B2m, Gapdh, and Rpl13a). Two sets of primers were used to detect Piezo2 expression. E45: exon 45 of the Piezo2 gene (NCBI reference sequence NM_001039485.4). ***p < 0.0001; two-way ANOVA test for both variables of GFP positivity and regions of gut, as well as the interaction of the two variables. Data are representative from three independent experiments. (e) Schematic of intraluminal administration of clozapine-N-oxide (CNO) followed by fecal pellet assay (FPA) and bead expulsion assay (BEA). FPA and BEA were conducted in the same cohorts at 15 and 20 min after CNO administration. (f) Serum serotonin level determined by ELISA from blood samples collected retro-orbitally 15 min after CNO administration (n = 3 mice per group). S: saline control, D1 (dose 1): 120 ng/kg, D2 (dose 2): 60 ng/kg. Representative data from two independent experiments are shown. Separate animal cohorts were used for serum serotonin assay versus FPA and BEA assays. (g) FPA from fecal pellets collected for 15 min after CNO administration. n = 10 in both WT and Piezo2-CHRM3* cohorts. Either WT or Piezo2-cre;ROSA26-LSL-CHRM3* (referred to as Piezo2-CHRM3*) mice were treated with CNO at 60 ng/kg once intraluminally. Representative data from three independent experiments. (h) BEA performed 20 min after CNO administration. A glass bead was inserted 2 cm into the distal colon, and the time to expel it was monitored in each animal. n = 10 in both cohorts. Representative data from three independent experiments.
-
Figure 5—source data 1
Quantitative PCR data, fecal pellet data, bead expulsion, serotonin levels.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig5-data1-v1.xlsx
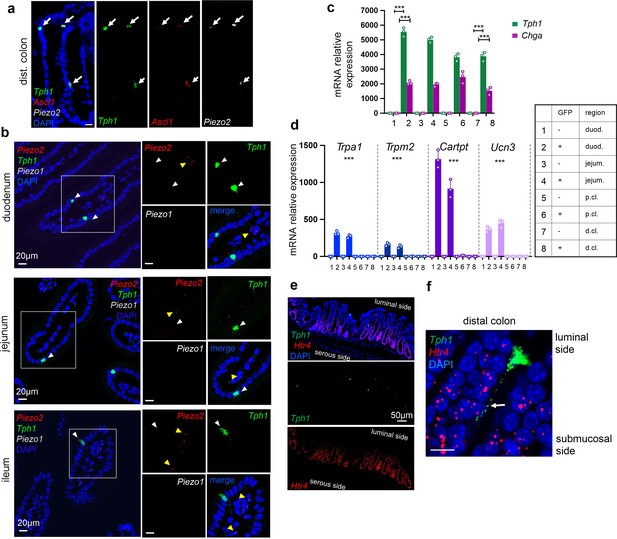
Piezo2 is highly enriched in distal colonic enterochromaffin (EC) cells that mediate colon motility.
(a) smRNA-FISH of Tph1, Ascl1, and Piezo2 in the distal colon. Arrows point to the triple-positive cells. Images are representative from three mice. (b) smRNA-FISH of Piezo2/Piezo1/Tph1 in the indicated regions of SI. White arrowheads point to the absence of Piezo2 in Tph1+ cells in the SI. Yellow arrowheads point to sparse staining of Piezo2 in the cells within the lamina propria. Images are representative from four mice. (c, d) qPCR validation of regional enriched genes in EC cells. GFP− and GFP+ cells were sorted from the duodenum, jejunum, and proximal and distal colon in the Tph1-bacTRAP mice. Relative gene expression was computed relative to the values in the GFP− cells isolated from the duodenum (indicated as 1), after normalization by the aggregates of three house-keeping genes (B2m, Gapdh, and Rpl13a). (c) Tph1 and Chga were highly enriched in the GFP+ cells from all regions. (d) Trpa1, Trpm2, Cartpt, and Ucn3 were highly enriched in small intestine. *** in (c) p < 0.001; unpaired two-tailed Student’s t-test (between the GFP− and GFP+ cells). *** in (d) p < 0.0001; two-way ANOVA test for both variables of GFP positivity and regions of gut, as well as the interaction of the two variables. Data are representative from three independent experiments. (e, f) smRNA-FISH assay for Htr4 and Tph1 in the distal colon. (e) Luminal side of the crypts is at the top of the image. An arrow points to the basal extension of a representative Tph1+ cell projecting toward Htr4+ epithelial cells (f). Image are representative from three wild-type mice. For all relevant panels ***p < 0.001. Scale bars in panels a, b, f: 10 µm; in panel e: 50 µm.
-
Figure 5—figure supplement 1—source data 1
Serum serotonin levels, bead expulsion.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig5-figsupp1-data1-v1.xlsx
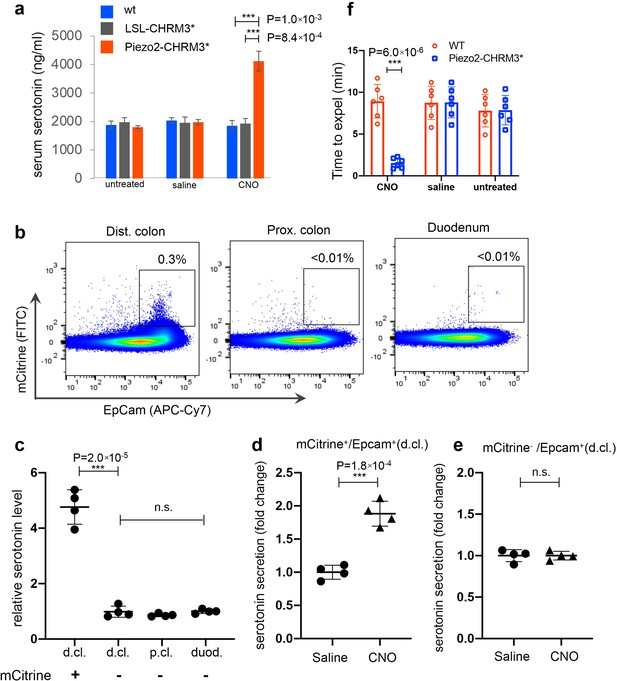
Chemogenetic activation of Piezo2 cells leads to serotonin release and accelerated colon motility.
(a) Serum serotonin level examined 15 min after clozapine-N-oxide (CNO) administration (60 ng/kg) by ELISA. n = 3 mice per group. (b) Representative flow cytometry results of dissociated epithelial cells from the indicated regions of the gut in the Piezo2-CHRM3*/mCitrine line. The Rosa26-LSL-CHRM3*/mCitrine locus is activated in a Piezo2-cre-specific manner. Percentages within indicated gates are shown. EpCam antibody was used to identify epithelial cells. Note that mCitrine+/EpCam+ cells were only observed in the distal colon of the Piezo2-CHRM3*/mCitrine mice. (c) Total serotonin levels shown for the sorted mCitrine+/EpCam+ cells from the distal colon (d.cl.), or mCitrine−/EpCam+ cells from the distal colon, proximal colon (p.cl.) and duodenum (duod.). Total serotonin was calculated as the sum of secreted serotonin and lysate serotonin, which was then normalized to the sum observed in the duodenal mCitrine−/EpCam+ cells. Elevated serotonin secretion as observed in response to CNO treatment (10 ng/µl) in the mCitrine+/EpCam+ cells (d) but not the mCitrine−/EpCam+ cells isolated from the distal colon (e). Percentage of secreted serotonin/total serotonin was normalized to that of saline-treated cells. Serotonin level was assessed by ELISA. Data in (b–e) are based on four mice from two independent experiments. (f) Bead expulsion assay (BEA) performed 20 min after CNO administration. For all relevant panels ***p < 0.001. n = 6 mice per group.
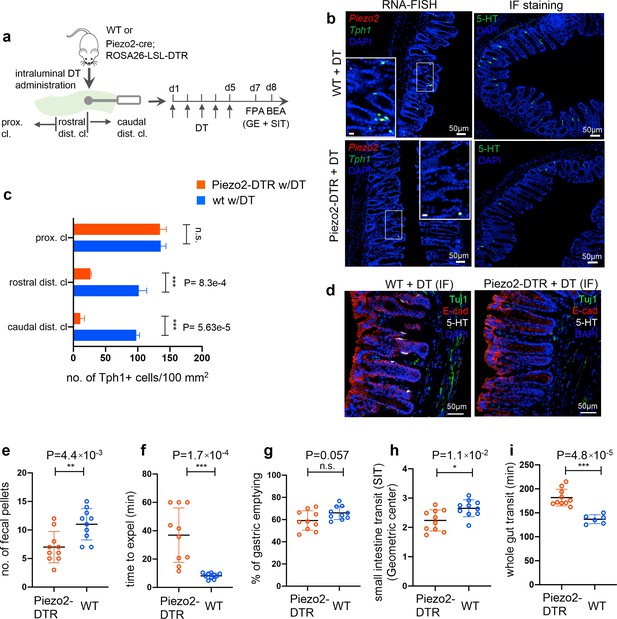
Piezo2+/Ascl1+/Tph1+ cells are required for normal colon motility.
(a) Schematic of intraluminal administration of diphtheria toxin (DT) followed by fecal pellet assay (FPA) and bead expulsion assay (BEA) in the same cohort. In a separate cohort, gastric emptying (GE) and small intestine transit (SIT) time assays were conducted following DT administration. Either wild-type or Piezo2-cre;ROSA26-LSL-DTR (referred to as Piezo2-DTR) mice were treated with DT at 50 µg/kg twice a day for 5 consecutive days. smRNA-FISH assay of Piezo2/Tph1 (left) and IF staining for 5-hydroxytryptamine (5-HT) (right) from the distal colon in wild-type (upper) and Piezo2-DTR (lower) mice after DT administration (b). Dashed boxes are enlarged in the insets. Images are representative from five animals per group. Scale bars: 50 µm. (c) Tph1+ cell counts based on smRNA-FISH experiments from the caudal distal colon (caudal dist. cl.), rostral distal colon (rostral dist. cl.), and the proximal colon (proximal cl.) for animals shown in (a) and (b). Five sections per animal were examined in five animals per group. (d) IF staining of 5-HT, E-cadherin, and Tuj1 in the distal colon of WT (left) and Piezo2- DTR (right) mice after DT administration. Images are representative of five different animals per group. Scale bars: 50 µm. FPA showing the number of fecal pellets collected in 2 hr (e), and BEA measuring the time to expel a glass bead inserted 2 cm into the distal colon (f). n = 10 per group, representative data from four independent experiments. Gastric emptying time (g) and SIT time (h) examined after 5 days of consecutive treatment of DT. Animals were orally gavaged with methylcellulose supplemented with rhodamine B dextran (10 mg/ml). Fifteen minutes after gavage, the remaining rhodamine B dextran was determined from the stomach and segments of intestine to assess upper GI motility. SIT was estimated by the position of the geometric center of the rhodamine B dextran in the small bowel. The geometric center values are distributed between 1 (minimal motility) and 10 (maximal motility). n = 10 in each group, representative data from three independent experiments are shown. (i) Whole gut transit time examined after 5 days of consecutive treatment of DT. An unabsorbable dye (carmine red) was administered by gavage and the time interval of first observance of the dye in stool was considered as whole gut transit time. n = 10 in Piezo2-DTR and 6 in wild-type control. Data are representative from two independent experiments. Error bars in panels e–i denote standard deviation of the mean; *p < 0.05, **p < 0.01, ***p < 0.001; unpaired two-tailed Student’s t-test.
-
Figure 6—source data 1
RNA-FISH cell counts and motility data.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig6-data1-v1.xlsx
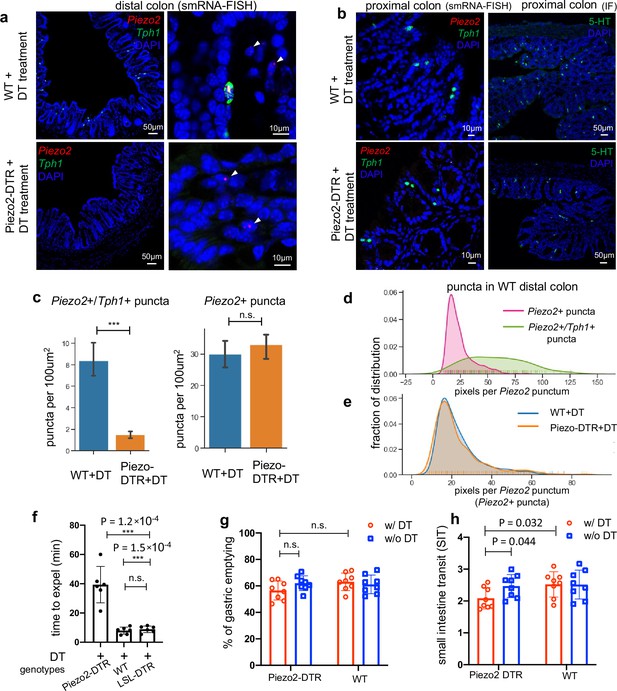
Piezo2+/Ascl1+/Tph1+ cells required for normal colon motility.
(a) Piezo2 and Tph1 transcripts determined by smRNA-FISH in the distal colon of wild-type (WT, upper) or Piezo2-DTR (lower) mice after DT administration. Arrowheads point to the remaining submucosal staining of Piezo2 in both WT and Piezo2-DTR mice. Images are representative from five animals per group. Scale bars: 10 or 50 µm, as indicated. (b) smRNA-FISH of Piezo2/Tph1 (left) and IF staining for 5-hydroxytryptamine (5-HT) (right) in the proximal colon of the WT (upper) or Piezo2-DTR (lower) mice after DT administration. Data are representative from five animals per group. Scale bars: 10 or 50 µm, as indicated. (c) Signals of Piezo2 single positivity (mostly in lamina propria) or Tph1/Piezo2 double positivity (in the epithelial layer) were identified by smRNA-FISH in the distal colon of WT (blue) or Piezo2- DTR (orange) mice after DT administration and quantitated using a custom pipeline in Cellprofiler. The numbers of Tph1/Piezo2 double-positive puncta (right) and Piezo2 single-positive puncta (left) were based on eight mice from three independent experiments (four of Piezo2-DTR + DT and four of WT + DT). n.s.: not significant, ***p < 0.001; unpaired two-tailed Student’s t-test. (d) Distribution of pixels per punctum in the Piezo2 channel quantitated in Piezo2 single-positive puncta (pink) versus Piezo2/Tph1 double-positive puncta (green). Pixels per punctum indicates expression levels of examined genes. (e) Distribution of pixels per punctum quantitated in Piezo2 single-positive puncta identified in WT + DT (blue) versus Piezo2-DTR + DT (orange). (f) Bead expulsion assay (BEA) performed after 5 days of consecutive treatment of DT in indicated groups. n = 6 in each group. WT mice and mice with only LSL-DTR transgene were included as controls. All animals received the same DT treatment. Gastric emptying and (g) small intestine transit (SIT) time (h). Animals were orally gavaged with methylcellulose supplemented with rhodamine B dextran (10 mg/ml). Fifteen minutes after gavage, the remaining rhodamine B dextran was determined in stomach and segments of intestine to assess upper GI motility. SIT was estimated by the position of the geometric center of the rhodamine B dextran in the small bowel. The geometric center values are distributed between 1 (minimal motility) and 10 (maximal motility). n = 8 in each group. Scale bars: 10 µm or as indicated otherwise. Error bars denote standard deviation of the mean. *p < 0.05, **p < 0.01, ***p < 0.001; unpaired two-tailed Student’s t-test.
-
Figure 6—figure supplement 1—source data 1
Bead expulsion, gastric emptying and transit data.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig6-figsupp1-data1-v1.xlsx
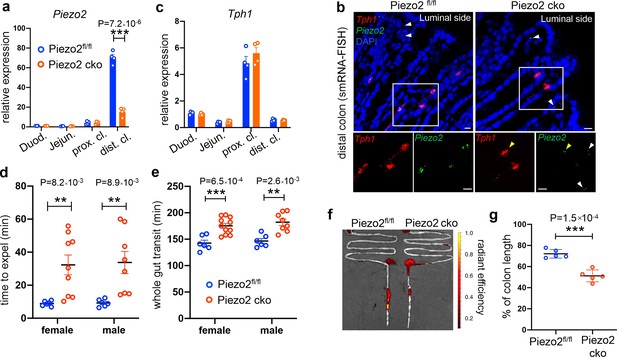
Epithelium Piezo2 is required for efficient colon motility.
(a) qPCR analysis of Piezo2 depletion in Villin-cre;Piezo2fl/fl mice (Piezo2 cko) and Piezo2fl/fl. qPCR was performed on the RNA prepared from the epithelial extracts of the indicated regions of the gut. Gene expression was computed relative to the values in the Piezo2fl/fl duodenum, after normalization by the aggregates of three house-keeping genes (B2m, Gapdh, and Rpl13a). Each circle represents one animal. Data were summarized from n = 4 in each group. (b) smRNA-FISH assay of Piezo2 and Tph1 in either Piezo2fl/fl (left) or Piezo2 cko (right) mice. Dashed boxed are enlarged and presented in individual channels. Data are representative from five different animals per group. White arrowheads point to the submucosal signals of Piezo2. Yellow arrowheads point to the residual Piezo2 signals in the Tph1+ cells of the Piezo2 cko animals. Scale bars: 10 µm. (c) As in (a), qPCR analysis of Tph1 in the Piezo2 cko and Piezo2fl/fl epithelium. (d) BEA. Each circle represents one animal. Representative data from two independent experiments. (e) Whole gut transit time. Each circle represents one animal. Representative data from two independent experiments. (f) Example fluorescent images of Piezo2 cko and Piezo2fl/fl intestine, 120 min after gavage of a fluorescent dye. (g) Summary data of fluorescent dye transit in the colon at 120 min after gavage. % of colon length = dye travel distance in colon ÷ full length of colon × 100%. Each circle represents one animal. Representative data from two independent experiments. Error bars in panels a–e, g denote standard deviation of the mean; *p < 0.05, **p < 0.01, ***p < 0.001; unpaired two-tailed Student’s t-test.
-
Figure 7—source data 1
Quantitative PCR data.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig7-data1-v1.xlsx
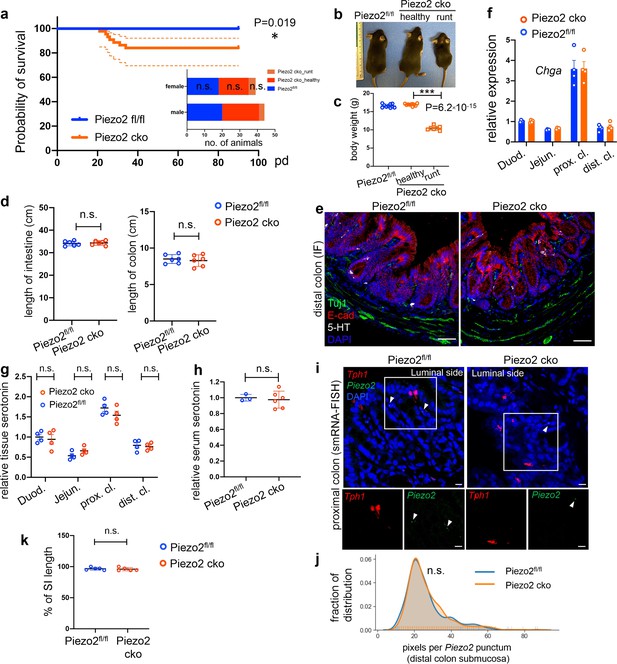
Epithelium Piezo2 is required for efficient colon motility.
(a) Kaplan–Meier survival curve of Vilin-cre;Piezo2fl/fl (Piezo2 cko, orange, n = 44) in comparison to littermate controls of Piezo2fl/fl (blue, n = 36). p = 0.019, Mantel–Cox test. Dashed lines represent 95% confidence interval (CI). Inset: sex breakdown of the early mortality phenotype. n.s., not significant; Chi-square test. (b) Representative image of littermates (all male) of indicated genotypes before humane euthanasia of Piezo2 cko-runt animals. (c) Summary of body weights before humane euthanasia of the Piezo2 cko-runt animals. Both sexes are included. Each circle represents a datapoint. (d) Length of intestine (left) and colon (right) of the indicated animals at 8 weeks of age. Both sexes are included. Each circle represents a datapoint. (e) IF staining of 5-hydroxytryptamine (5-HT), E-cadherin, and Tuj1 in the distal colon of Piezo2fl/fl (left) and Piezo2 cko (right) mice. Images are representative of three animals per group. (f) qPCR analysis of Chga in Piezo2 cko and Piezo2fl/fl epithelium. Gene expression was computed relative to the values in the Piezo2fl/fl duodenum, after normalization by the aggregates of three house-keeping genes (B2m, Gapdh, and Rpl13a). Each circle represents one animal. n = 4 per group. (g) Tissue serotonin level determined by ELISA and normalized by the serotonin level detected in the duodenum of the Piezo2fl/fl mice. n = 4 for both Piezo2 cko (Vilin-cre;Piezo2fl/fl) and Piezo2fl/fl mice. n = 4 per group. (h) Serum serotonin level determined by ELISA and normalized to the levels observed in Piezo2fl/fl mice. n = 6 for Piezo2 cko and 3 for Piezo2fl/fl mice. (i) smRNA-FISH of Piezo2 and Tph1 in the proximal colon of Piezo2fl/fl (left) or Piezo2 cko (right) mice. Data are representative of three animals per group. (j) Distribution of pixels per punctum quantitated in Piezo2 single-positive puncta identified in Piezo2fl/fl (blue) or Piezo2 cko (orange) distal colon. Pixels per punctum indicates expression levels of examined genes. (k) Summary data of fluorescent dye transit at 70 min after gavage. % of CI length = dye travel distance in SI ÷ full length of SI × 100%. Each circle represents one animal. Representative data from two independent experiments. Scale bars: e, 50 µm; i, 10 µm. Error bars stand for standard deviation of the mean. *p < 0.05, ***p < 0.001; unpaired two-tailed Student’s t-test.
-
Figure 7—figure supplement 1—source data 1
Serotonin levels and transit times.
- https://cdn.elifesciences.org/articles/90596/elife-90596-fig7-figsupp1-data1-v1.xlsx
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain (Mus musculus) C57Bl6 | Neurod1-Cre | Jackson Labs | JAX 028364 | |
| Strain (M. musculus) C57Bl6 | Rosa26-LSL- tdTomato | Jackson Labs | JAX 007914 | |
| Strain (M. musculus) C57Bl6 | Piezo2-IRES-cre | Jackson Labs | JAX 027719 | |
| Strain (M. musculus) C57Bl6 | Rosa26-LSL- CHRM3 | Jackson Labs | JAX 026220 | |
| Strain (M. musculus) C57Bl6 | Rosa26-LSL-DTR | Jackson Labs | JAX 007900 | |
| Strain (M. musculus) C57Bl6 | Piezofl/fl | Jackson Labs | JAX 027720 | |
| Strain (M. musculus) C57Bl6 | Vilin-cre | Jackson Labs | JAX 021504 | |
| Antibody | anti-5-HT (Goat; polyclonal) | ImmunoStar, | Cat# 20079, RRID:AB_572262 | IF(1:10,000) |
| Antibody | anti-5-HT (Rabbit; polyclonal) | ImmunoStar, | Cat# 20080, RRID:AB_572263 | IF(1:1000) |
| Antibody | anti-Secretin (Rabbit; polyclonal) | Phoenix Pharmaceuticals | Cat# H-067–04, RRID:AB_2650428 | IF(1:2000) |
| Antibody | anti-CCK/ gastrin (Mouse monoclonal) | Gift | Code: 28.2 RRID:AB_2650429 | IF(1:2000) |
| Antibody | anti-CCK/ gastrin (Rabbit; polyclonal) | Gift | Code: 8007/5 RRID:AB_2884979 | IF(1:8000) |
| Antibody | anti-GIP (Rabbit; polyclonal) | Santa Cruz | Cat# SC23554, RRID:AB_2247481 | IF(1:500) |
| Antibody | anti-Ghrelin (Rabbit; polyclonal) | Santa Cruz | Code MF1601, RRID:Ab_2767291 | IF(1:10,000) |
| Antibody | anti-GFP (Rabbit; polyclonal) | Abcam | Cat# AB13970, RRID:AB_300798 | IF(1:2000) |
| Antibody | anti-Oxyntomodulin (Mouse monoclonal) | Anshlabs | Cat# 323AO010, RRID:AB_2934156 | IF(1:2000) |
| Antibody | anti-Substance P (Rabbit; polyclonal) | Gift | Cat# SK1, RRID:AB_2814842 | IF(1:1600) |
| Antibody | anti-Neurotensin (Goat; polyclonal) | Santa Cruz | Cat# SC-7592, RRID:AB_650372 | IF(1:50) |
| Antibody | anti-IFT88 (Rabbit; polyclonal) | Proteintech | Cat# 13967-1AP, RRID:AB_2121979 | IF(1:500) |
| Antibody | anti-Tuj1 (Rabbit; polyclonal) | Abcam | Cat# Ab18207, RRID:AB_444319 | IF(1:1000) |
| Antibody | anti-e-cadherin (Rat; polyclonal) | Abcam | Cat# Ab11512, RRID:AB_298118 | IF(1:1000) |
| Antibody | anti-Dclk1 (Rabbit; polyclonal) | Abcam | Cat# Ab31704, RRID:AB_873537 | IF(1:1000) |
| Antibody | anti-Mouse IgG Alexa 488 (Rabbit; polyclonal) | Molecular Probes | Cat# 21202, RRID:AB_141607 | IF(1:500) |
| Antibody | anti-Chicken IgG Alexa 488 (Goat; polyclonal) | Jackson ImmunoResearch | Cat# 103–545155, RRID:AB_2337390 | IF(1:500) |
| Antibody | anti-Rabbit IgG Alexa 568 (Donkey; polyclonal) | Molecular Probes | Cat# A10042, RRID:AB_2534017 | IF(1:800) |
| Antibody | anti-Sheep IgG Alexa 488 (Donkey; polyclonal) | Molecular Probes | Cat# A11015, RRID:AB_141362 | IF(1:500) |
| Antibody | anti-Goat IgG Alexa 555 (Donkey; polyclonal) | Molecular Probes | Cat# A21432, RRID:AB_141788 | IF(1:800) |
| Sequence-based reagent | Mm-Tph1 | ACDBio/Bio-Techne | 318701 C2 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Piezo2 | ACDBio/Bio-Techne | 400191 C3 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Gpx3 | ACDBio/Bio-Techne | 400191 C3 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Cartpt | ACDBio/Bio-Techne | 432001 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Stra6 | ACDBio/Bio-Techne | 450321 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Cck | ACDBio/Bio-Techne | 402271 C3 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Foxj1 | ACDBio/Bio-Techne | 317091 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Dnah9 | ACDBio/Bio-Techne | 556771 C3 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Mc4r | ACDBio/Bio-Techne | 402741 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Casr | ACDBio/Bio-Techne | 423451 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Asic5 | ACDBio/Bio-Techne | 588601 C3 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Pyy | ACDBio/Bio-Techne | 420681 C3 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Gpbar1 | ACDBio/Bio-Techne | 318451 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Trpm2 | ACDBio/Bio-Techne | 313291 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Ascl1 | ACDBio/Bio-Techne | 313291 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Olfr558 | ACDBio/Bio-Techne | 316131 C2 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Htr4 | ACDBio/Bio-Techne | 408241 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Piezo1 | ACDBio/Bio-Techne | 500511 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Onecut3 | ACDBio/Bio-Techne | 583241 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Crp | ACDBio/Bio-Techne | 583251 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Cpb2 | ACDBio/Bio-Techne | 583261 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Tlr2 | ACDBio/Bio-Techne | 317521 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Tlr5 | ACDBio/Bio-Techne | 451601 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-F5 | ACDBio/Bio-Techne | 502411 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Gip | ACDBio/Bio-Techne | 451601 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Cnr1 | ACDBio/Bio-Techne | 420721 C2 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Ucn3 | ACDBio/Bio-Techne | 464861 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Il12a | ACDBio/Bio-Techne | 414881 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Iapp | ACDBio/Bio-Techne | 512571 C2 | Probe used for smRNA-FISH |
| Sequence-based reagent | Mm-Olfr78 | ACDBio/Bio-Techne | 436601 | Probe used for smRNA-FISH |
| Sequence-based reagent | Foxj1-F | IDT DNA | N/A | qPCR primer 5’ AGCCCAGAAGACTGGGAACT 3’ |
| Sequence-based reagent | Foxj1-R | IDT DNA | N/A | qPCR primer 5’ AATCCTTGGGCTTGAGGGAAC 3’ |
| Sequence-based reagent | Ascl1-F | IDT DNA | N/A | qPCR primer 5’ GAATGGACTTTGGAAGCAGGATG 3’ |
| Sequence-based reagent | Ascl1-R | IDT DNA | N/A | qPCR primer 5’ TGCCCCTGTAGGTTGGCTG 3’ |
| Sequence-based reagent | Piezo2-a-F | IDT DNA | N/A | qPCR primer 5’ GCACTCTACCTCAGGAAGACTG 3’ |
| Sequence-based reagent | Piezo2-a-R | IDT DNA | N/A | qPCR primer 5’ CAAAGCTGTGCCACCAGGTTCT 3’ |
| Sequence-based reagent | Piezo2-b-F | IDT DNA | N/A | qPCR primer 5’ TCAAACACGCCAGTGACAAT 3’ |
| Sequence-based reagent | Piezo2-b-R | IDT DNA | N/A | qPCR primer 5’ TGTCTCTGAACAAAATGATGGTGA 3’ |
| Sequence-based reagent | Trpa1-F | IDT DNA | N/A | qPCR primer 5’ GAGGATTGCTATGCAGGTGGA 3’ |
| Sequence-based reagent | Trpa1-R | IDT DNA | N/A | qPCR primer 5’ CGTGCCTGGGTCTATTTGGA 3’ |
| Sequence-based reagent | Chga-F | IDT DNA | N/A | qPCR primer 5’ CCAAGGTGATGAAGTGCGTC 3’ |
| Sequence-based reagent | Chga-R | IDT DNA | N/A | qPCR primer 5’ GGTGTCGCAGGATAGAGAGGA 3’ |
| Sequence-based reagent | Tph1-F | IDT DNA | N/A | qPCR primer 5’ TGTTGACTGCGACATCAGCCGA 3’ |
| Sequence-based reagent | Tph1-R | IDT DNA | N/A | qPCR primer 5’ GGAAACCAAGGGACAGTCTCCA 3’ |
| Sequence-based reagent | Trpm2-F | IDT DNA | N/A | qPCR primer 5’ AAGGATGTGGCTCTCACAGAC 3’ |
| Sequence-based reagent | Trpm2-R | IDT DNA | N/A | qPCR primer 5’ CGGGAACCCATACTCGACC 3’ |
| Sequence-based reagent | B2m-F | IDT DNA | N/A | qPCR primer 5’ CACTGAATTCACCCCCACTGA 3’ |
| Sequence-based reagent | B2m-R | IDT DNA | N/A | qPCR primer 5’ TGTCTCGATCCCAGTAGACGG 3’ |
| Sequence-based reagent | Rpl13a-F | IDT DNA | N/A | qPCR primer 5’ AGCAGATCTTGAGGTTACGGA 3’ |
| Sequence-based reagent | Rpl13a-R | IDT DNA | N/A | qPCR primer 5’ GGAGTCCGTTGGTCTTGAGG 3’ |
| Sequence-based reagent | Gapdh-F | IDT DNA | N/A | qPCR primer 5’ CTGGAGAAACCTGCCAAGTATG 3’ |
| Sequence-based reagent | Gapdh-R | IDT DNA | N/A | qPCR primer 5’ AGAGTGGGAGTTGCTGTTGAAG 3’ |
| Sequence-based reagent | Cartpt-F | IDT DNA | N/A | qPCR primer 5’ AAGAAGTACGGCCAAGTCCC 3’ |
| Sequence-based reagent | Cartpt-R | IDT DNA | N/A | qPCR primer 5’ CAGTCACACAGCTTCCCGAT 3’ |
| Sequence-based reagent | Ucn3-F | IDT DNA | N/A | qPCR primer 5’ AAGGCCAAGAATTTGCGAGC 3’ |
| Sequence-based reagent | Ucn3-R | IDT DNA | N/A | qPCR primer 5’ TGTCTTGATGTGCCACCCTC 3’ |
| Sequence-based reagent | Il12a-F | IDT DNA | N/A | qPCR primer 5’ CCACTGGAACTACACAAGAACG 3’ |
| Sequence-based reagent | Il12a-R | IDT DNA | N/A | qPCR primer 5’ ATGCTACCAAGGCACAGGGT 3’ |
| Software, algorithm | ImageJ | NIH | RRID:SCR_003070 | https://imagej.net/ij/ |
| Software, algorithm | Zen Blue | Zeiss | RRID:SCR_013672 | |
| Software, algorithm | Prism | Graphpad | RRID:SCR_002798 | https://www.graphpad.com/ |
| Software, algorithm | CellRanger | 10 X Genomics | RRID:SCR_023221 | v. 4.0.0 |
| Software, algorithm | Python | Python Software Foundation | RRID:SCR_008394 | v. 3.6.2 |
| Software, algorithm | Scanpy | Scanpy development team | RRID:SCR_018139 | v. 2.1.4 |
| Commercial assay or kit | RNeasy Micro Kit | Qiagen | 74104 | |
| Commercial assay or kit | SuperScript III First-Strand Synthesis System | ThermoFisher | 18080051 | |
| Commercial assay or kit | FastStart Universal SYBR Green Master Mix | Sigma | 4913850001 | |
| Commercial assay or kit | Serotonin ELISA Assay Kit | Eagle Biosciences | EA602 | |
| Other | DAPI stain | Invitrogen | D1306 | (1 µg/mL) |
Additional files
-
MDAR checklist
- https://cdn.elifesciences.org/articles/90596/elife-90596-mdarchecklist1-v1.docx
-
Supplementary file 1
Summary of EC clusters and their potential physiological roles.
- https://cdn.elifesciences.org/articles/90596/elife-90596-supp1-v1.docx