Redox regulation and dynamic control of brain-selective kinases BRSK1/2 in the AMPK family through cysteine-based mechanisms
Figures
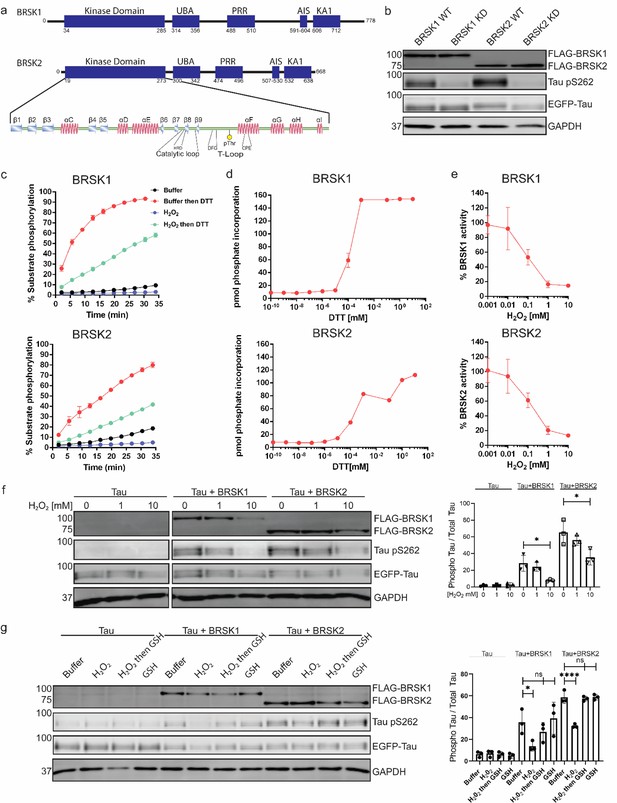
BRSK1/2 are redox sensitive.
(a) Schematic representation of BRSK domain architecture, including kinase domain, ubiquitin-associated (UBA) domain, proline-rich region (PRR), kinase-associated domain (KA1), and autoinhibitory sequence (AIS). (b) Immunoblotting showing BRSK-dependent phosphorylation of Tau at Ser262 (pS262), from lysates of HEK-293T cells overexpressing full-length FLAG-BRSK1 or 2 (wild type [WT] or kinase dead [KD]) and EGFP-Tau. (c) Real-time phosphorylation of fluorescent AMARA peptide by full-length BRSK1 and 2 (200 ng) purified from Sf21 cells. BRSK proteins were incubated with buffer or 1 mM H2O2 for 10 min, reactions were then initiated with the addition of ATP and peptide substrate in the presence (where indicated) of 10 mM DTT. Dose–response curves for (d) DTT and (e) H2O2 with 200 ng full-length BRSK1 and BRSK2. All kinases assays are shown as mean and SD of three experiments. (f) Immunoblot (left) of pS262 in transiently co-transfected HEK-293T cells incubated with the indicated concentration of H2O2 for 10 min. Signal density for phospho Tau S262 and total Tau (GFP) was obtained using ImageStudio software (LI-COR) and results from at least three biological replicates were analyzed with GraphPad Prism software using one-way ANOVA to determine significance. Data shown is mean and SE. (g) Representative immunoblot (left) of transiently co-transfected HEK-293T cells treated with 10 mM H2O2 for 10 min before the addition of 20 mM GSH. Whole-cell lysates were harvested after a further 15 min. Normalized densitometry of Tau pS262 signal (right) was calculated from three independent experiments. Data shown is mean and SD. *p<0.05, **p<0.01, ***p<0.001.
-
Figure 1—source data 1
Original western blots for Figure 1, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig1-data1-v1.zip
-
Figure 1—source data 2
Original files for western blot analysis displayed in Figure 1.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig1-data2-v1.zip
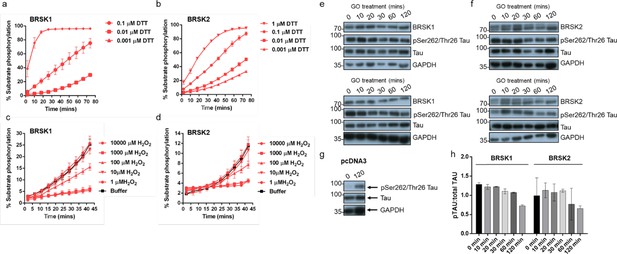
Redox regulation of BRSK1 and 2.
(a–d) Real-time phosphorylation of fluorescent AMARA peptide by full-length BRSK1 and 2 (200 ng). BRSK proteins were incubated with buffer or the indicated concentrations of DTT or H2O2. Rates of BRSK activity were calculated as pmol per min phosphate incorporation and are presented in Figure 1. Data shown here is a subset of the conditions shown in Figure 1 (mean and SD from three repeats). (e–h) Time dependent loss of pTAU by incubation of HEK-293T cells with 2 U/ml glucose oxidase (GO). Cells were transiently co-transfected with EGFP-Tau and either (e) BRSK1, (f) BRSK2, or (g) empty vector (pcDNA3). Data shown is western blot analysis from two independent repeats. (h) pTau:Tau signals calculated with ImageJ. Data shown is mean and SD, calculated from (e) and (f).
-
Figure 1—figure supplement 1—source data 1
Original western blots for Figure 1—figure supplement 1, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig1-figsupp1-data1-v1.zip
-
Figure 1—figure supplement 1—source data 2
Original files for western blot analysis displayed in Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig1-figsupp1-data2-v1.zip
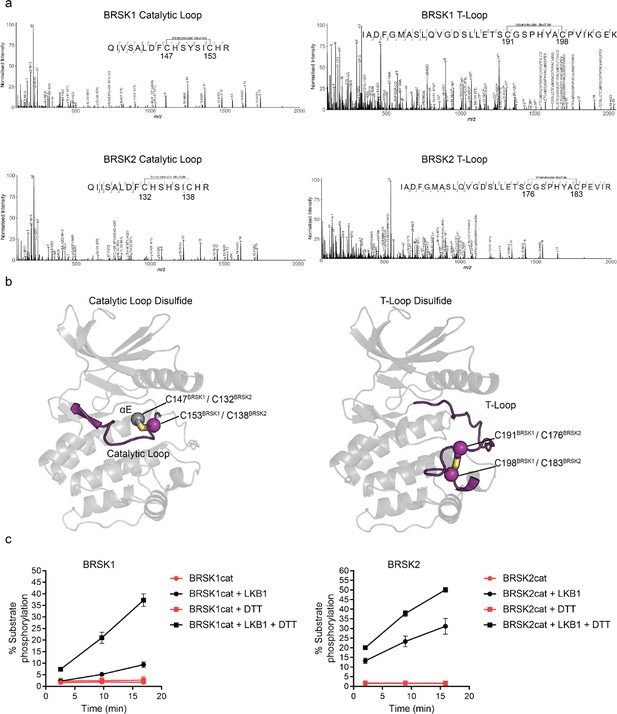
Intramolecular disulfide bonds form in the kinase domains of BRSK1 and 2.
(a) Full-length BRSK1 and 2 were affinity-purified from HEK-293T cells and subjected to LC-MS/MS analysis. LC-MS/MS spectrum mapping revealed disulfide bridges formation between C147BRSK1–C153BRSK1, C191BRSK1–C198BRSK1, C132BRSK2–C138BRSK2, and C176BRSK2–C183BRSK2. Peptide coverage was 74 and 78% for BRSK1 and BRSK2 respectively. (b) AlphaFold structures demonstrating the location of disulfide bonds within the kinase domains of BRSK1 and BRSK2. (c) Real time phosphorylation of fluorescent AMARA peptide by the kinase domains of BRSK1 and 2 (100 ng) purified from E. coli. BRSK1 (29–358) and BRSK2 (14–341) were activated by incubation with LKB1 and assayed in the presence or absence of 1 mM DTT.
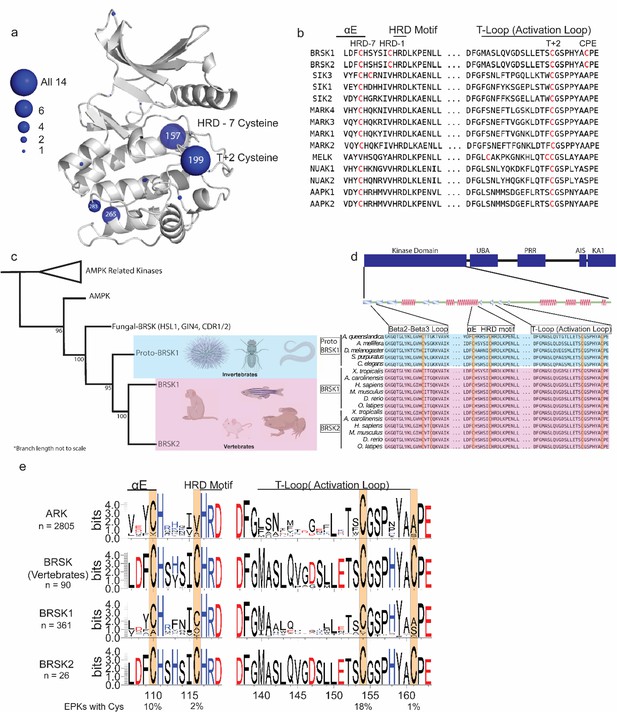
Cysteine pairs are highly conserved within the activation segments of BRSKs.
(a) Mapping of Cys residues (spheres) in the kinase domains of human ARK family members. Numbers represent the corresponding amino acid position in PKA. Sphere size is proportional to the number of ARKs that contain a Cys at a specific site. (b) Activation segment sequence alignment of the 14 human ARKs. (c) Phylogenetic analysis showing divergence and grouping of BRSKs sub-families in different taxonomic groups. Bootstrap values are included for each clade. (d) Sequence alignment of the kinase domains of invertebrate and vertebrate BRSKs. (e) Analysis of relative amino acid conservation in ARKs and BRSKs, centered on the HRD containing catalytic loop, and the T-loop (between the DFG and APE motifs). Data is presented as hidden Markov models (HMM) Sequence Logos. The % of ePKs that possess a specific Cys is shown at the bottom.
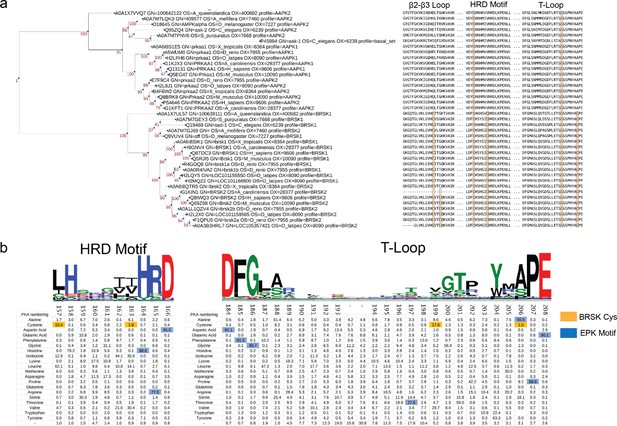
Phylogenetic analysis of BRSK and the ARK family.
(a) Phylogenetic analysis of the ARK family reveals that the closest relative of BRSK kinases is AMPK. The number of cysteines in the kinase domain of BRSKs increases relative to AMPK. (b) Sequence alignment and relative amino acid composition of the activation segment of ePKs (top). Data is presented as hidden Markov models (HMM) Sequence Logos. Table (bottom) depicts the frequency of an amino acid at each position along the catalytic and T-loop. Key Cys residues are highlighted in orange; residues highly conserved in ePK canonical kinase motifs are highlighted in blue.
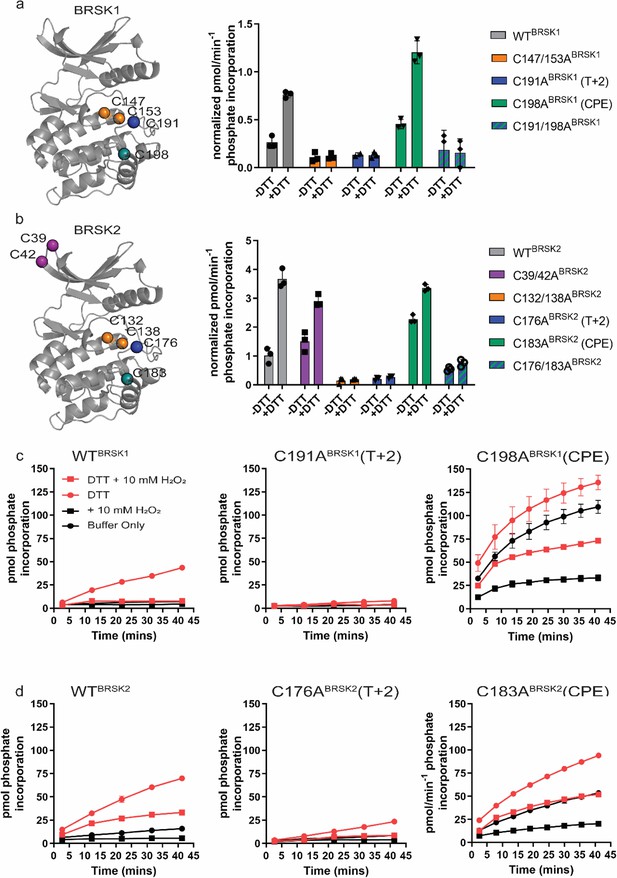
Cysteine residues within the kinase domain fine-tune BRSK activity.
In vitro kinase assays (right panels) showing normalized rates of peptide phosphorylation by WT and Cys-to-Ala variants of (a) BRSK1 and (b) BRSK2. 100 ng of LKB1-activated BRSK kinase domain was assayed in the presence or absence of 1 mM DTT. The positions of mutated Cys residues are modeled on the kinase domain as colored spheres (left panel). Real-time in vitro assays using (c) 50 ng BRSK1 and (d) 20 ng BRSK2. LKB1-activated BRSK proteins were incubated on ice in the presence or absence of 250 µM DTT for 30 min. Assays were initiated by the addition of ATP and fluorescent peptide substrate in the presence or absence of 1 mM H2O2. All data are mean and SD of three experiments and activities are normalized to LKB1-phosphorylated BRSK signal.
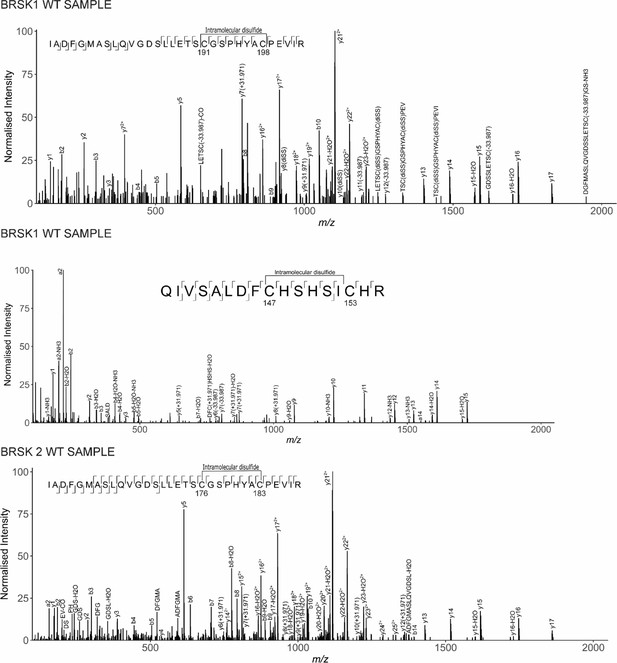
LC-MS/MS analysis of BRSK1/2 catalytic domains.
LC-MS/MS reveals intramolecular disulfide bonds in the kinase domains of BRSK1 and 2 purified from E. coli.
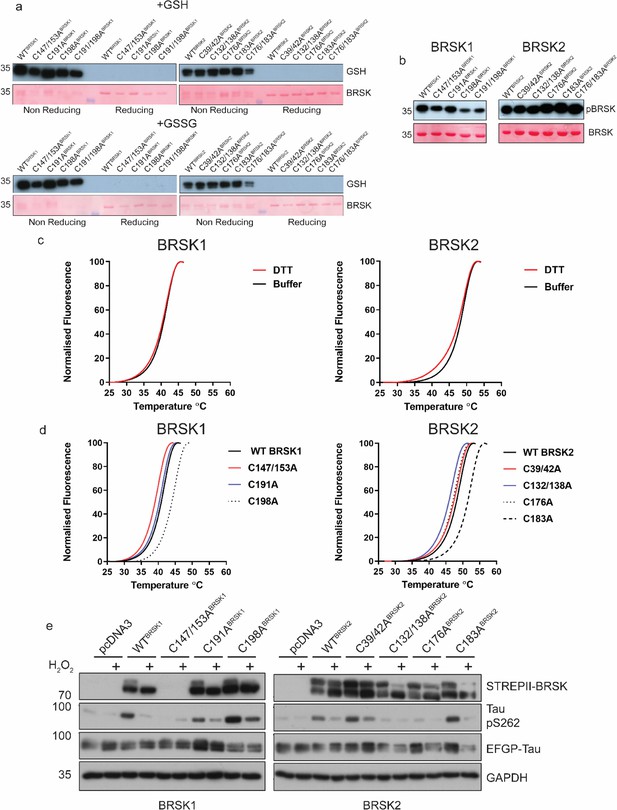
Biochemical analysis of BRSK Cys-to Ala mutants.
(a) Immunoblot of in vitro glutathionylation of BRSK kinase domains. (b) Immunoblot showing LKB1-dependent phosphorylation of BRSK kinase domain proteins. (c) Thermal denaturation curves of BRSK catalytic domain proteins in the presence or absence of 10 mM DTT. (d) Thermal denaturation curves of BRSK catalytic domain cysteine to alanine mutants. (e) Representative immunoblot of EGFP-Tau co-expressed with full-length, WT, and Cys-to-Ala mutants of BRSK1 and BRSK2. Transiently transfected HEK-293T cells were treated with or without 10 mM H2O2 for 10 min.
-
Figure 4—figure supplement 2—source data 1
Original western blots for Figure 4—figure supplement 2, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig4-figsupp2-data1-v1.zip
-
Figure 4—figure supplement 2—source data 2
Original files for western blot analysis displayed in Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig4-figsupp2-data2-v1.zip
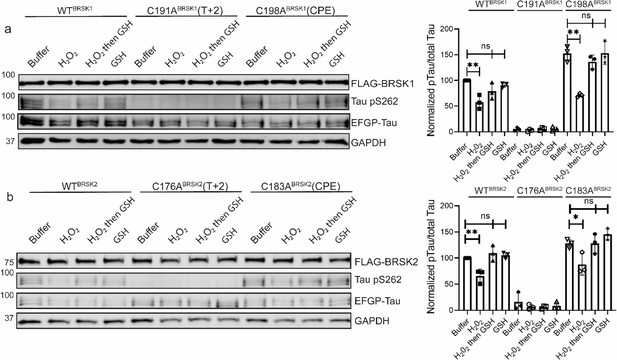
Impact of T-Lloop and CPE Cys-to-Ala mutations on BRSK redox sensitivity in a cellular EGFP-Tau HEK-293T co-expression system.
Representative immunoblot of EGFP-Tau co-expressed with WT and Cys-to-Ala mutants of (a) BRSK1 and (b) BRSK2 (left panels). Transiently transfected HEK-293T cells were treated with or without 10 mM H2O2 for 10 min before the addition of 20 mM GSH. Whole-cell lysates were harvested after a further 15 min. Signal density for phospho Tau S262 and total Tau (GFP) was obtained using ImageStudio software (LI-COR) and results from at least three biological replicates were analyzed with GraphPad Prism software using one-way ANOVA to determine significance (right panels). Data shown is mean and SE. All values are normalized to Tau pS262 signals from control (buffer only treatment) WT BRSK and Tau co-transfections. Data shown is mean and SD. *p<0.05, **p<0.01, ***p<0.001.
-
Figure 5—source data 1
Original western blots for Figure 5, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig5-data1-v1.zip
-
Figure 5—source data 2
Original files for western blot analysis displayed in Figure 5.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig5-data2-v1.zip
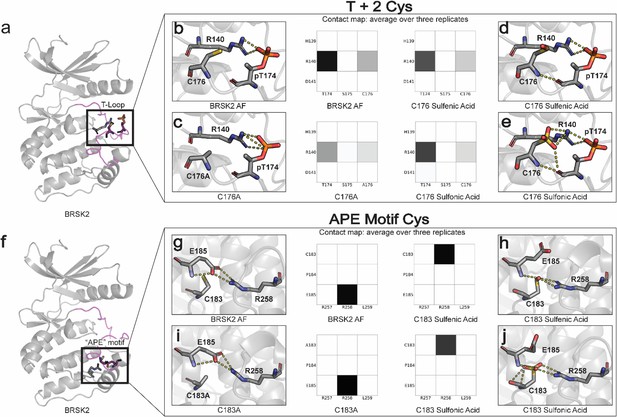
Oxidative cysteine modifications alter critical structural interactions required for BRSK allosteric regulation.
Three replicates of 100 ns GROMACS molecular dynamics simulations were performed to evaluate the effects of cysteine mutation and oxidation. Salt bridge disruption was analyzed by generating contact maps representing the percentage of the simulation time in which residues were within appropriate distance (3 Å). (a) T +2 Cys is located in proximity to the activation loop threonine in the T loop. (b–e) Evaluation of pT174-R140 salt bridge formation in wild-type, C176A, and oxidized C176 BRSK2. (f) Location of CPE salt bridge within BRSK2. (g–j) Evaluation of E185-R258 salt bridge formation in wild-type, C183A, and oxidized C183 BRSK2.
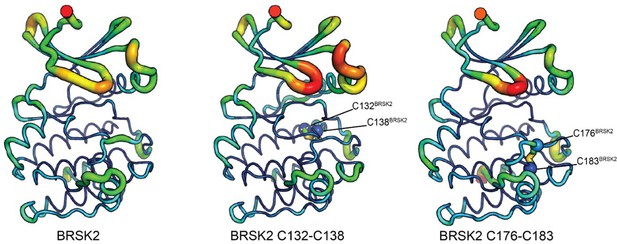
Molecular dynamics simulations of intramolecular disulfide bonds.
Simulations incorporating disulfide bonds identified in MS/MS experiments. Root Mean Square Fluctuation (RMSF) was calculated based on three 100 ns GROMACS molecular dynamics simulations. Higher mobility is indicated by warmer colors and thickness of representation.
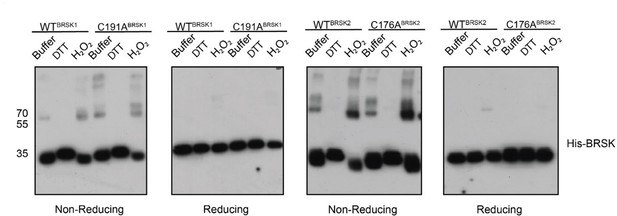
BRSK1/2 form limited disulfide-mediated multimers.
Western blot analysis of BRSK1/2 kinase domain purified from E. coli and incubated with buffer, H2O2, or DTT and subjected to non-reducing or reducing PAGE to evaluate the formation of intramolecular disulfide bonds.
-
Figure 7—source data 1
Original western blots for Figure 7, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig7-data1-v1.zip
-
Figure 7—source data 2
Original files for western blot analysis displayed in Figure 7.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig7-data2-v1.zip
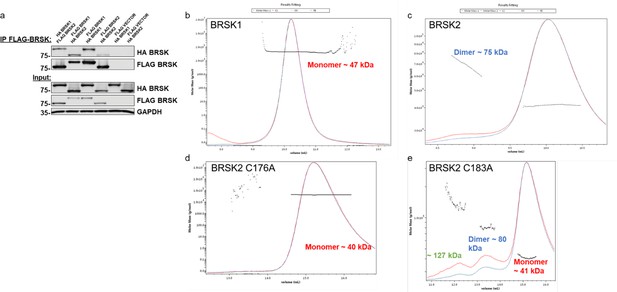
Evidence for limited BRKS dimer species.
(a) Co-immunoprecipitation of HA-BRSK1/2 with immunoprecipitated FLAG-BRSK1/2 expressed in HEK-293T cells. SEC-MALS analysis of WT (b) BRSK1 and (c) 2, and (d) C176A and (e) C183A BRSK2 kinase domains in solution, performed in the absence of reducing agents.
-
Figure 7—figure supplement 1—source data 1
Original western blots for Figure 7—figure supplement 1, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig7-figsupp1-data1-v1.zip
-
Figure 7—figure supplement 1—source data 2
Original files for western blot analysis displayed in Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/92536/elife-92536-fig7-figsupp1-data2-v1.zip
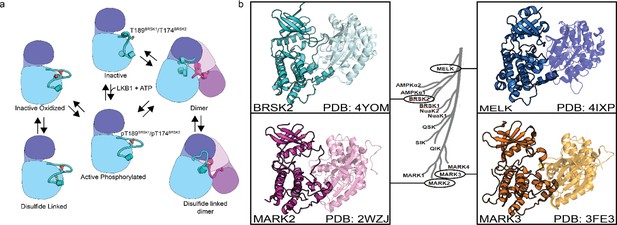
Model of BRSK1/2 regulation.
(a) Schematic diagram demonstrating ways in which residues within BRSK kinases permit fine-tuning of catalytic activity through a variety of oxidative modifications, potentially including inter and intramolecular disulfide bonds. Cartoon representation of kinase domain with N-lobe colored dark blue/purple and the C-lobe colored light blue/purple. (b) ARK family member BRSK2, MELK, and MARK2/3 crystal structures demonstrate the ability to form asymmetric dimers, bringing T +2 cys into proximity. Crystal structures for MARK2, and MELK both contain intermolecular disulfide bonds between T +2 cys (Marx et al., 2010; Marx et al., 2006; Murphy et al., 2007; Cao et al., 2013).
Additional files
-
Supplementary file 1
Proximal cysteine pairs in the human kinome predicted from structural analysis of human protein kinases in the AlphaFold database.
Cysteine pairs within 10 Å are considered proximal.
- https://cdn.elifesciences.org/articles/92536/elife-92536-supp1-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/92536/elife-92536-mdarchecklist1-v1.docx





