Uncovering the BIN1-SH3 interactome underpinning centronuclear myopathy
Figures
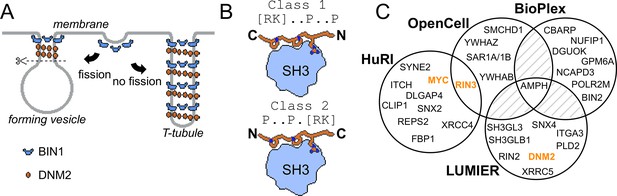
Involvement of BIN1 in membrane remodeling and a compendium of known BIN1 interaction partners.
(A) Models of vesicle and T-tubule formation in the context of BIN1 and DNM2. BIN1 interplays in both processes, through its membrane bending/tubulating BAR domain and its SH3 domain. In clathrin-, or caveolin-coated vesicle formation, as well as during the formation of recycling endosomes, the recruitment of DNM2 by BIN1 is critical for vesicle scission. During T-tubule formation, DNM2 is also recruited, but in this case less scission occurs. (B) Schematic illustration of binding of PRMs to SH3 domains. Due to the twofold pseudo-symmetry of PPII helices, class 1 and class 2 PxxP motifs bind in different orientations to SH3 domains (Lim et al., 1994). (C) Known interaction partners of BIN1 identified by high-throughput qualitative interactomic studies and the experimental overlap between the different sources (Cho et al., 2022; Ellis et al., 2012; Huttlin et al., 2021; Luck et al., 2020). Note that the known SH3-domain mediated interaction partners, that were studied by low-throughput methods, were only detected on a few occasions (DNM2, MYC, RIN3, marked in orange), or not detected at all (TAU/MAPT, CAVIN4).
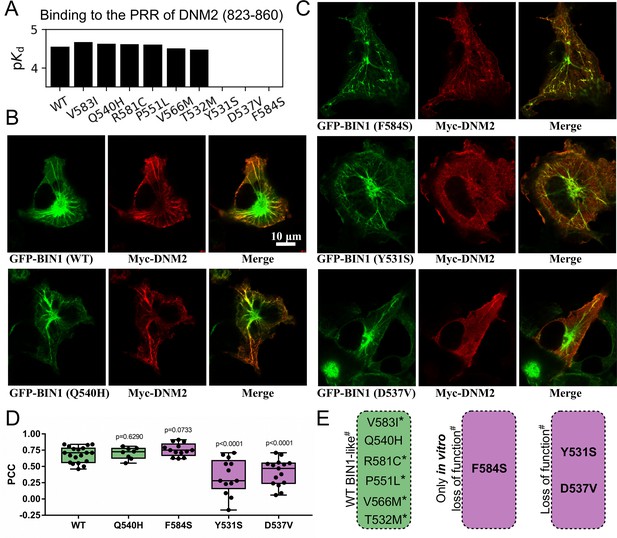
Several BIN1 variants of unknown clinical significance have a strong impact on the binding of DNM2 and display altered cellular phenotype.
(A) Measured affinities of the PRR of DNM2 against a set of natural BIN1_SH3 variants. Most variants interact with DNM2 with similar affinities, but Y531S, D537V and F584S variants disrupt this interaction. Affinities are expressed as negative logarithm of dissociation constants, i.e. pKd 4 equals to 100 μM Kd. (B) Membrane tubulation assay performed with WT BIN1 and DNM2, as well as Q540H variant which binds DNM2 with the same affinity as WT BIN1. (C) Membrane tubulation assay performed with the variants displaying decreased affinities to DNM2. Cos-1 cells were transfected with GFP-BIN1 and Myc-DNM2. The effect of F584S seems to be apparently rescued in the context of FL BIN1, but both Y531S and D537V variants are unable to efficiently recruit DNM2 to membrane tubules in cells. (D) Statistical analysis of single-cell co-localization experiments between the BIN1 variants and DNM2 (n[WT]=19, n[Q540H]=8, n[F584S]=13, n[Y531S]=13, n[D537V]=15). P values were calculated between Pearson correlation coefficients (PCC) of WT and missense variants using a two-tailed unpaired Student’s T-test. Box plots indicate the median and upper and lower quartiles, and whiskers label the minimal and maximal measured PCC values. Individual data points representing measurements of single cells are also indicated. (E) A summary of the effects of the BIN1 variants. Asterisk indicates that the variants were only tested in vitro, and # indicates that the effects were measured based on the BIN1-DNM2 interaction phenotype. See Figure 2—figure supplement 1 for additional images.
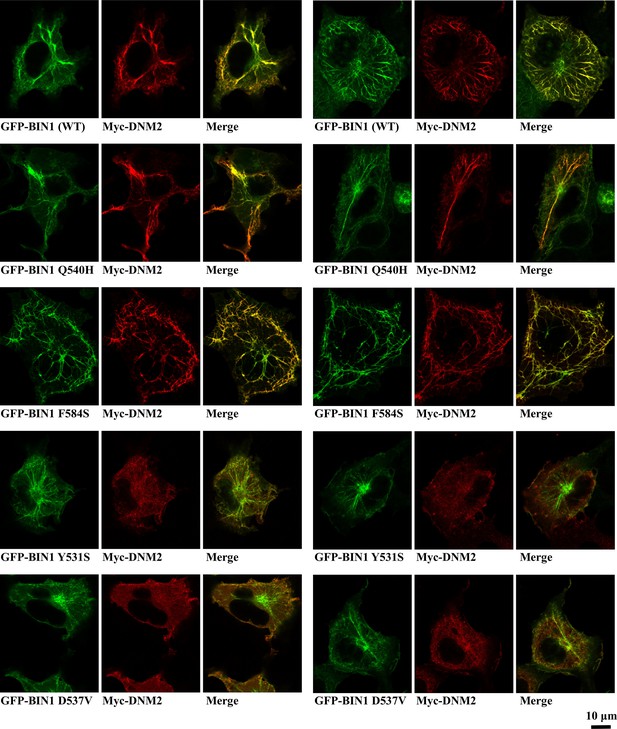
Additional images of membrane tubulation assay performed with BIN1 variants.
Cos-1 cells were transfected with GFP-BIN1 and Myc-DNM2. Confocal images were taken from fixed cells. Rare BIN1 variants Y531S and D537V decrease the amount of DNM2 recruited to BIN1-induced membrane tubules. F584S variant shows altered mesh-like organization of the membrane tubules.
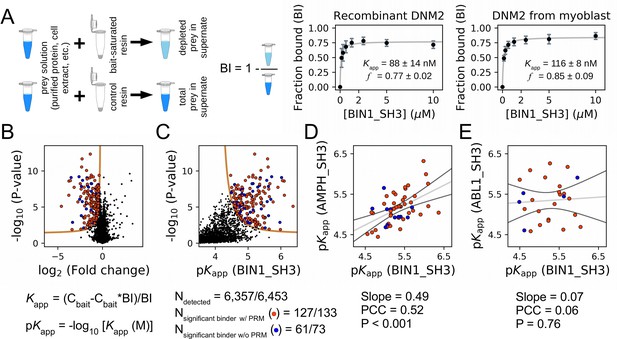
Affinity measurements between the SH3 domain of BIN1 and full-length proteins from cell extracts using nHU-MS.
(A) Outline of the holdup assay and benchmarking of nHU. Holdup is a simple tool to measure the fraction bound of prey molecules. This prey solution can be either a single purified protein, or a complex mixture of molecules and the prey depletion can be monitored with a multitude of analytical approches, such as western blot, or mass spectrometry. Titration holdup experiments were used to further characterize the interactions of BIN1_SH3 and full-length DNM2 using either recombinant, purified DNM2, or total myoblast extract containing endogenous DNM2 as a prey. The two experiments show nearly identical binding affinity and partial activity. (The average and standard deviation of three nHU western blot experiments are shown.) (B) Results of single point nHU-MS experiments carried out with the SH3 domain of BIN1 and total Jurkat extracts. Interaction partners above the significance threshold (tan line) are colored in orange if putative PRMs were identified in their sequence and blue if not. (C) Measured depletion values were converted to affinities using the functions indicated below panel B, assuming a simple binding mechanism and 10 μM estimated bait concentration. The number of unique affinity measurements performed and the identified BIN1 interaction partners found in a single experiment/in all measurements are indicated below panel C. (D, E) We also performed nHU-MS experiments with a set of closely or distantly related SH3 domains and compared their affinity profiles with BIN1. This way, we could quantify that related SH3-domains, for example the one found in AMPH, show similarities in their affinity interactomes, displaying statistically significant correlation between the measured affinity constants. In contrast, unrelated SH3 domains, such as the one found in ABL1, bind targets with dissimilar affinities. A linear fit (grey line) and a 95% confidence band (black line) is shown on all affinity comparisons. The statistical significance of correlation was determined by two-tailed, unpaired T-test. See Figure 3—figure supplements 1–3 and Supplementary file 1 for further details. Mass spectrometry experiments were performed with three injection replicates.
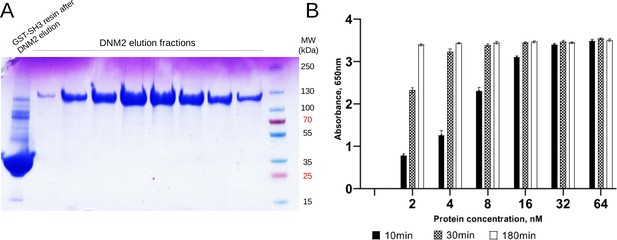
Quality control of purified DNM2.
(A) FL human DNM2 was produced in insect cells and was purified on GST column, preloaded with GST-fused BIN1_SH3. DNM2 was eluted at acidic pH and the pooled elution fractions were dialized to a basic pH. (B) Malachite green GTPase activity assay was used to verify the catalytic activity of recombinant DNM2 (n = 3).
-
Figure 3—figure supplement 1—source data 1
Original SDS-PAGE for panel A.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp1-data1-v1.zip
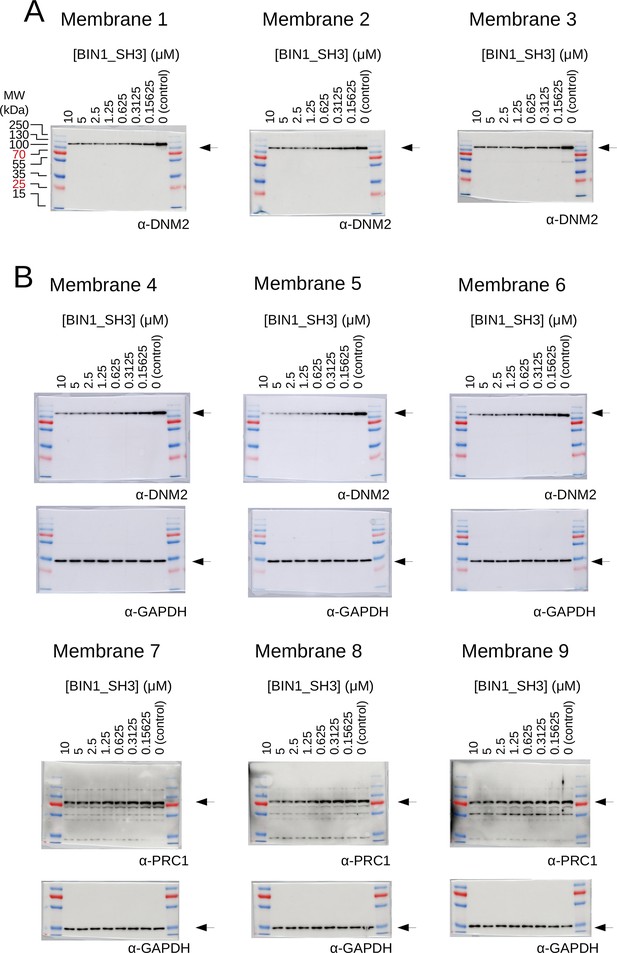
Raw results of the titration nHU and titration HU experiments.
(A) Titration HU-WB experiments were carried out using purified DNM2. (B) Titration nHU-WB experiments were carried out using myoblast extracts. The recovered supernatants were assayed using western blot with DNM2, PRC1 and GAPDH antibodies. The recovered supernatants were assayed using western blot with DNM2 antibody. All western blots were repeated three times and the measured luminescent signals overlaid with colorimetric images are shown.
-
Figure 3—figure supplement 2—source data 1
Original western blot image (overlayed with colorimetric image) of Membrane 1.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data1-v1.zip
-
Figure 3—figure supplement 2—source data 2
Original western blot image (overlayed with colorimetric image) of Membrane 2.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data2-v1.zip
-
Figure 3—figure supplement 2—source data 3
Original western blot image (overlayed with colorimetric image) of Membrane 3.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data3-v1.zip
-
Figure 3—figure supplement 2—source data 4
Original western blot image (overlayed with colorimetric image) of Membrane 4, DNM2.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data4-v1.zip
-
Figure 3—figure supplement 2—source data 5
Original western blot image (overlayed with colorimetric image) of Membrane 4, GAPDH.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data5-v1.zip
-
Figure 3—figure supplement 2—source data 6
Original western blot image (overlayed with colorimetric image) of Membrane 5, DNM2.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data6-v1.zip
-
Figure 3—figure supplement 2—source data 7
Original western blot image (overlayed with colorimetric image) of Membrane 5, GAPDH.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data7-v1.zip
-
Figure 3—figure supplement 2—source data 8
Original western blot image (overlayed with colorimetric image) of Membrane 6, DNM2.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data8-v1.zip
-
Figure 3—figure supplement 2—source data 9
Original western blot image (overlayed with colorimetric image) of Membrane 6, GAPDH.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data9-v1.zip
-
Figure 3—figure supplement 2—source data 10
Original western blot image (overlayed with colorimetric image) of Membrane 7, PRC1.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data10-v1.zip
-
Figure 3—figure supplement 2—source data 11
Original western blot image (overlayed with colorimetric image) of Membrane 7, GAPDH.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data11-v1.zip
-
Figure 3—figure supplement 2—source data 12
Original Western blot image (overlayed with colorimetric image) of Membrane 8, PRC1.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data12-v1.zip
-
Figure 3—figure supplement 2—source data 13
Original western blot image (overlayed with colorimetric image) of Membrane 8, GAPDH.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data13-v1.zip
-
Figure 3—figure supplement 2—source data 14
Original western blot image (overlayed with colorimetric image) of Membrane 9, PRC1.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data14-v1.zip
-
Figure 3—figure supplement 2—source data 15
Original western blot image (overlayed with colorimetric image) of Membrane 9, GAPDH.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig3-figsupp2-data15-v1.zip
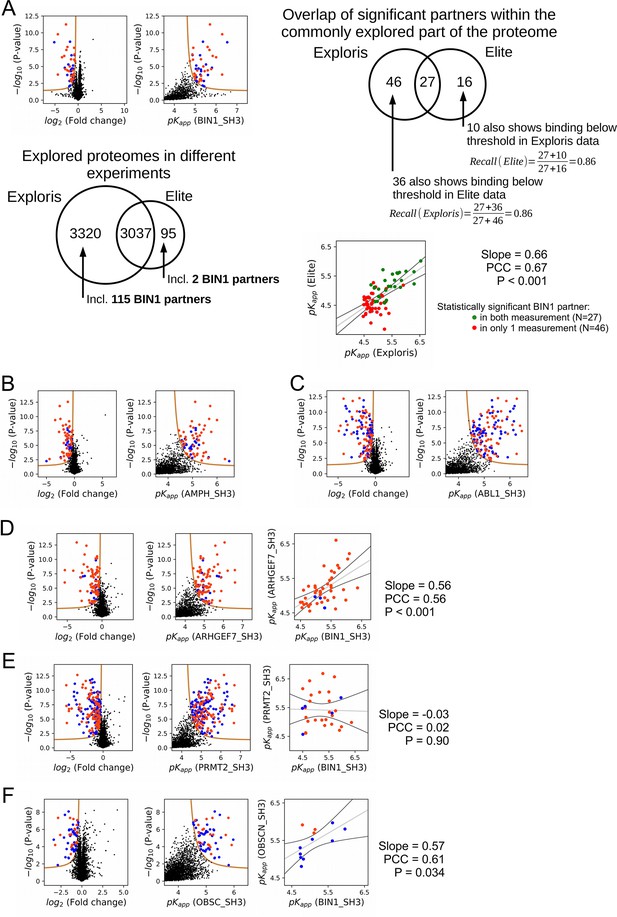
Additional results of nHU-MS experiments.
(A) Result of an independent single point nHU-MS experiment carried out with the SH3 domain of BIN1 measured on a different mass spectrometer (Orbitrap Elite). Below the volcano plot, the mapped proteomic space is shown of the two nHU measurements that were done using the two instruments (Orbitrap Elite and Orbitrap Exploris 480). On the top right, the statistical overlap of significant binders is shown, as well as their recall calculations. Below, the comparison of the independently measured BIN1 affinities is shown in the two measurements, differently coloring partners that were found to be significant in both measurement from partners that were found to be significant in only one measurement. (B–F) Results of single point nHU-MS experiments carried out with the SH3 domains of AMPH, ABL1, ARHGEF7, PRMT2, and OBSCN using total Jurkat extracts (left). Interaction partners that show deficiency in abundance above the significance threshold (tan line) are colored in orange in case we could identify putative class 1/2 PxxP motifs in their sequence and blue in case we cannot. Measured depletion values were converted to affinities using a 10 μM estimated bait concentration assuming simple binding mechanism (middle/right). (C, D, E) On panel C, D and E, an additional comparison is shown with the affinities of BIN1 (extreme right panel). A linear fit (grey line) and a 95% confidence band (black line) is shown on all affinity comparisons.
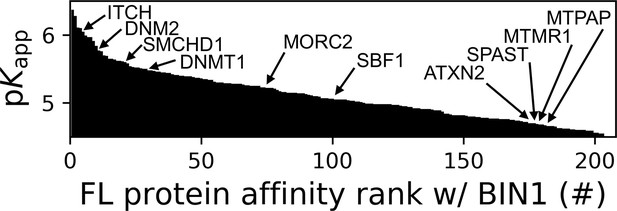

Affinity ranking of the 206 FL interaction partners of the BIN1 SH3 domain identified in nHU-MS experiments.
Interaction partners found in previous studies, as well as partners whose importance was found to be significant in neuromuscular disorders are indicated. See Supplementary file 1 for further details.
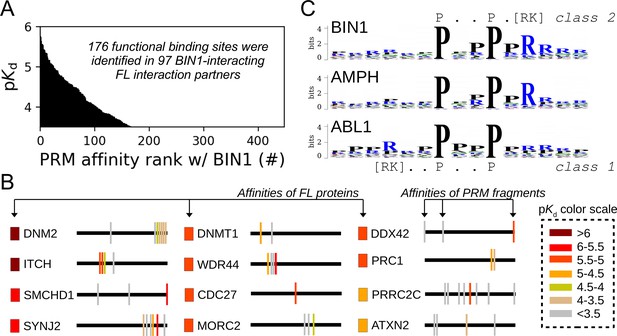
Affinity measurements between the SH3 domain of BIN1 and isolated PRM fragments.
(A) Affinity profile of BIN1_SH3 measured using fragmentomic holdup against 448 synthetic PRMs found in FL interaction partners previously identified by nHU-MS. 176 PRMs were found to bind to BIN1 displaying affinities ranging from low micromolar to a few hundreds of micromolar dissociation constants. These motifs were found in 97 proteins, matching at least a single functional binding site for half of the identified FL interaction partners. (B) The combination of native and fragmentomic holdup reveals biophysical properties of FL proteins and elementary binding sites. The measured affinities of intact proteins are indicated with colored boxes and site-specific affinities of individual PRMs are indicated with colored spikes, where colors were adjusted to measured steady-state affinities of FL proteins and PRM sites, respectively. Note that the protein schemes are not to scale to the actual protein length, but are the approximate relative positions of indicated PRMs. (C) Affinity-weighted specificity logo of the SH3 domains of BIN1, AMPH, and ABL1. BIN1 and AMPH nearly uniquely interacts with class 2 PxxP motifs, while ABL1 prefers to bind class 1 PxxP motifs. See Figure 5—figure supplement 1 and Supplementary file 2 for further details.
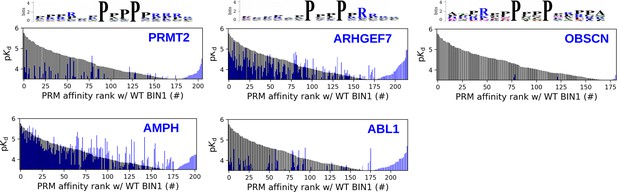
Site-specific affinity interactomes of other SH3 domains.
The affinity interactomes of the five tested SH3 domains are shown (blue) in comparision with the affinity profile of BIN1 (black). Note the highest similarity between the affinity profiles of BIN1 and AMPH. Only the motifs displaying detectable binding are shown. Affinity-weighted specificity logos are shown for PRMT2, ARHGEF7 and OBSCN above their affinity profiles.
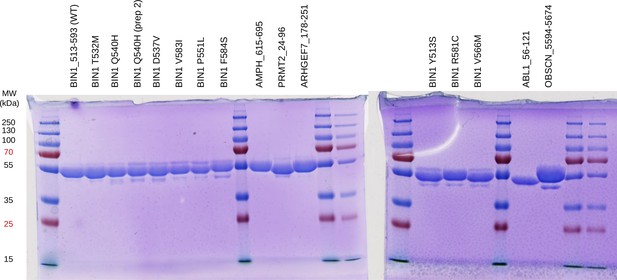
Quality control of purified MBP-fused SH3 domains used for fragmentomic holdup experiments.
The double-affinity purified His6-MBP-fused SH3 domains were loaded on SDS-PAGE and stained with Coomassie brilliant blue. No major contaminants or degradation products were detected.
-
Figure 5—figure supplement 2—source data 1
Original SDS-PAGE.
- https://cdn.elifesciences.org/articles/95397/elife-95397-fig5-figsupp2-data1-v1.zip
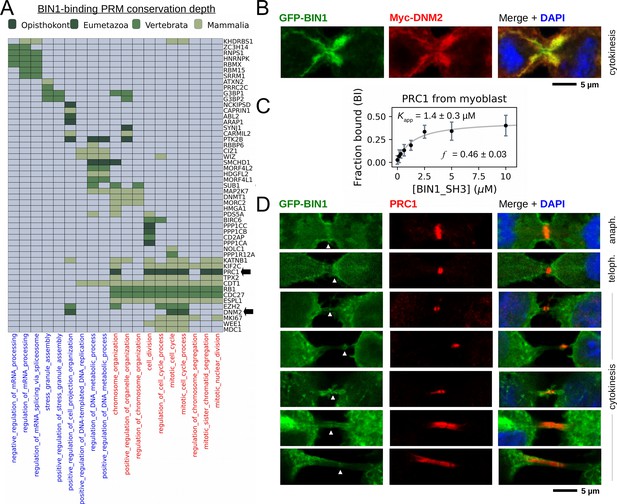
BIN1 interacts with multiple proteins involved in the mitotic phase and is localized at the membrane bridge formed between the daughter cells.
(A) Functional clustering of the identified BIN1 partners that contains BIN1-binding PxxP motifs. At the bottom of the panel, nuclear, or nucleic-acid-related processes are colored in blue and mitotic processes are colored in red. Heatmap color coding is according to the conservation depth of the highest affinity BIN1-interacting motif. See Supplementary file 3 for data. (B) During cytokinesis, BIN1 and DNM2 were found to localize at the cleavage furrow. A representative image of dividing Cos-1 cells, that were transfected with GFP-BIN1 and Myc-DNM2. (C) Titration nHU to further characterize the interactions of BIN1 with PRC1. PRC1 binding is measured from the same binding experiment using myoblast extract, that was used to characterize DNM2 binding (Figure 3A). (The average and standard deviation of three nHU western blot experiments are shown.) (D) Representative confocal images of the membrane bridge between daughter cells. Cos-1 cells were transfected with GFP-BIN1 and stained for endogenous PRC1. White arrows indicate the cleavage furrow or the midbody. For additional supporting confocal images see Figure 6—figure supplement 1.
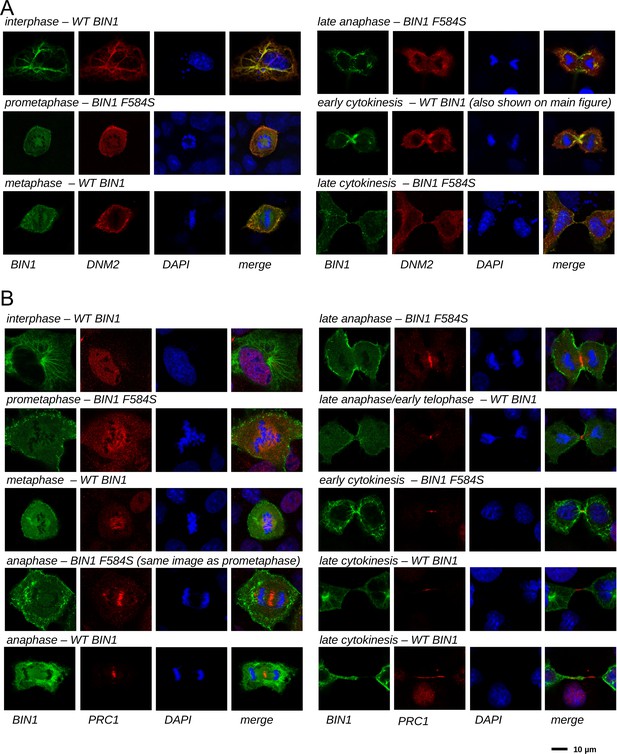
Cellular translocation of BIN1 during mitosis and cytokinesis.
Cos-1 cells were either transfected with GFP-BIN1 and Myc-DNM2 (A) or GFP-BIN1 alone (B). Confocal images were taken from fixed cells stained for Myc and DAPI (A) or PRC1 and DAPI (B). Both WT BIN1, as well as the F584S variants were used, where we have found larger number of dividing cells. Cell phase was determined based on the nuclear phase and the overall morphology.
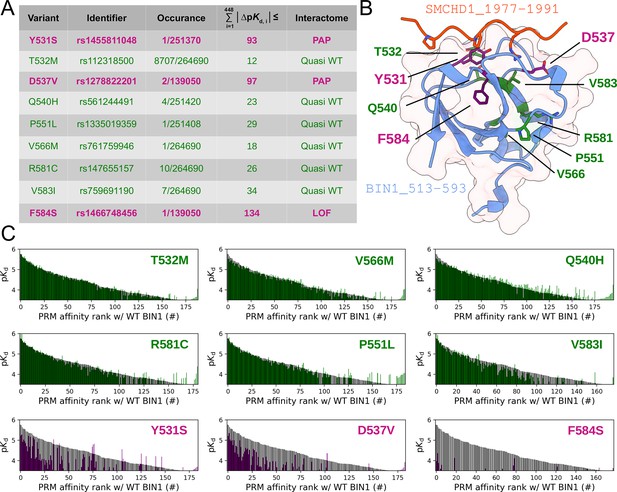
Natural variants of BIN1 can cause affinity perturbation at an interactomic scale.
(A) A summary of our affinity interactomic tests performed with 9 natural variants of the BIN1 SH3 domain. The cumulative Euclidean affinity distances to the WT BIN1, calculated based on the explored 448 dimensional affinity space, are indicated for each variant. For affinities where no detectable binding was observed the detection threshold was used for calculation, hence only the lower limit of the Euclidean distance could be estimated. Variants colored green have minor effect on affinity interactomes, while variants colored in purple displaying either perturbed affinity profiles (PAP) or general loss of functions (LOF). (B) Location of the assayed variants on the structure of WT BIN1. D537 and Y531 are positioned near the PRM binding interface, F584 is buried in the hydrophobic core of the SH3 domain. The structure of the BIN1 SH3 domain bound to a high affinity peptide taken from SMCHD1 was generated using AlphaFold2 (Tunyasuvunakool et al., 2021). (C) Affinity interactome profiles of the BIN1 variants (colored in green or purple) compared with the affinity profile of WT BIN1 (colored in black). Motifs in the affinity profiles were ranked according to their affinities measured with WT BIN1. Only the motifs displaying detectable binding out of the 449 assayed PRMs are shown. See Supplementary file 2 for further details.
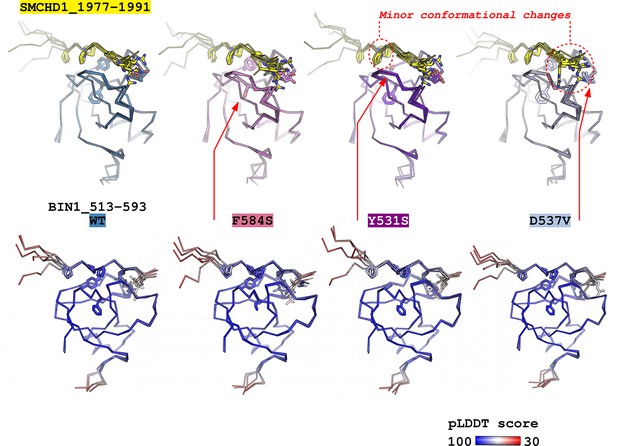
Predicting the structural consequences of the likely pathogenic variants of BIN1 with Alphafold2.
The structure of the BIN1_513–593 – SMCHD1_1977–1991 complex was predicted using ColabFold v1.5.2 – a user friendly implementation of AlphaFold2. The complexes of 4 BIN1 variants were predicted: WT, F584S, Y531S, D537V. No major perturbations in the bound PRM conformation could be observed and only minor local changes could be observed in the proximity of the Y531S, or D537V sequence variants. Only the side chains of the core optimal BIN1-binding PxPPxRR motif is shown on the SMCHD1 PRM and only the side chains of residues 531, 537, and 584 are shown on the BIN1_SH3. For each BIN1 variant, below the structures focusing on the binding sites, structures are shown colored by the pLDDT scores of AlphaFold2. No major change has been observed upon mutation in these scores.
Additional files
-
Supplementary file 1
Results of the nHU-MS experiments.
Each sheet contains the results of a single experiment performed with a given bait. Significant partners have numerical values in the “pKdsign” column.
- https://cdn.elifesciences.org/articles/95397/elife-95397-supp1-v1.xlsx
-
Supplementary file 2
Results of the fragmentomic holdup experiments.
Significant partners have non “0” values in the “pKd” column.
- https://cdn.elifesciences.org/articles/95397/elife-95397-supp2-v1.xlsx
-
Supplementary file 3
Results of the bioinformatic analysis.
The first sheet contains the evolutionary conservation of PxxP motifs. The second sheet contains the results of the GO enrichment analysis.
- https://cdn.elifesciences.org/articles/95397/elife-95397-supp3-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/95397/elife-95397-mdarchecklist1-v1.pdf





