Obox4 promotes zygotic genome activation upon loss of Dux
Figures
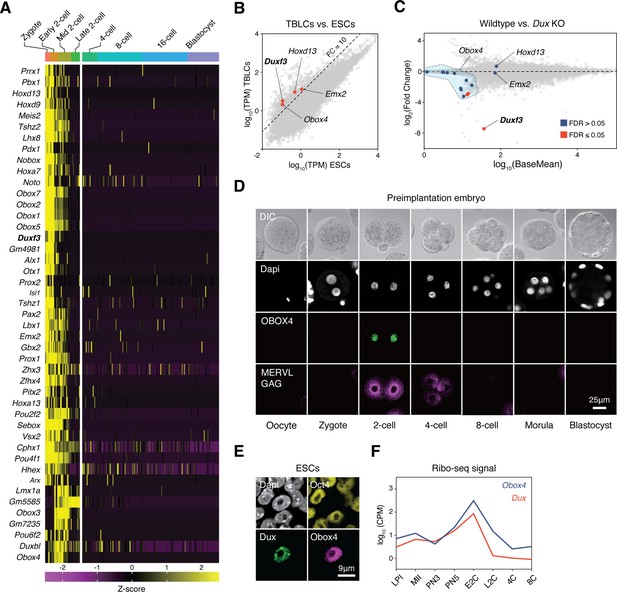
OBOX4 is expressed during zygotic genome activation (ZGA).
(A) Mouse homeobox genes that are specifically expressed during ZGA. The genes were identified by means of statistically determined k-means clustering based on their expression in preimplantation embryos. Dux is shown in a bold italic font. (B) Scatterplot showing per-gene normalized read counts in mouse totipotent totipotent blastomere-like cells (TBLCs) versus mouse embryonic stem cells (mESCs). Genes with more than tenfold normalized read counts (FC = 10) have been highlighted. Dux is shown in a bold italic font. (C) MA plot displaying gene expression in Dux knockout 2C-embryos versus WT 2C-embryos. Loss of Dux partially downregulates the multicopy homeobox gene Obox4. (D) Immunofluorescence staining of OBOX4 and MERVL GAG at different preimplantation embryo stages. (E) Immunofluorescence staining of DUX, OBOX4, and OCT4 in 2C-like mESCs. (F) Translation profile of DUX and OBOX4 during preimplantation characterized by Ribo-seq signal.
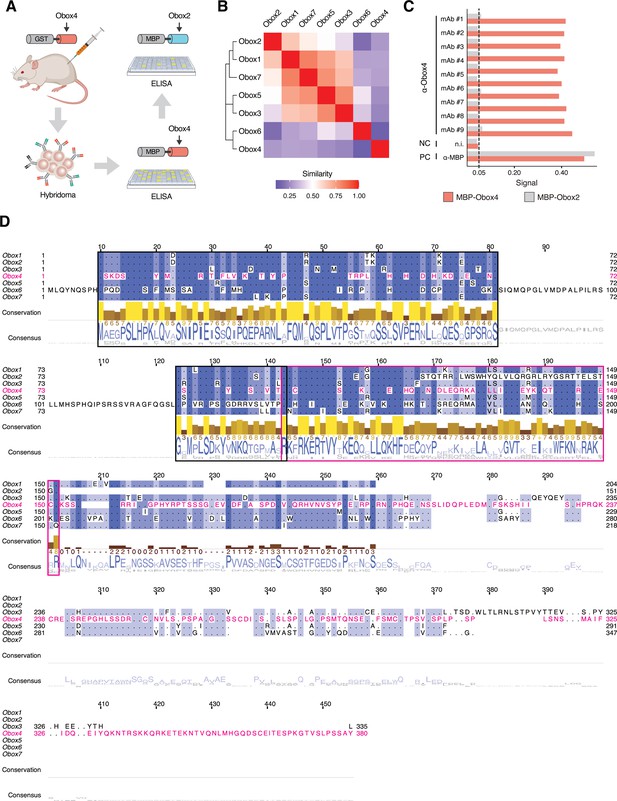
Production of anti-OBOX4 monoclonal antibodies.
(A) Schematic of the production of monoclonal antibodies. Mice were immunized with the GST-OBOX4 fusion protein. After fusing B-cells with myelomas, the antibodies produced by hybridomas were examined against MBP-OBOX4 fusion protein, followed by MBP-OBOX2 fusion protein, for identification of OBOX4-specific antibodies. (B) Analysis of protein sequence similarity among the OBOX family members. The Smith–Waterman local alignment method was used to align the amino acid sequences, and the divergence among different sequences was represented in the form of a block substitution matrix. (C) ELISA signal of anti-OBOX4 monoclonal antibodies against the MBP-OBOX4 and MBP-OBOX2 fusion proteins. Anti-OBOX4 monoclonal antibodies were not reactive to MBP-OBOX2. (D) Multiple-sequence alignment of protein sequence among the OBOX family members. The homeodomains are highlighted by a magenta box. The peptide sequences used for antibody production are highlighted by a black box.
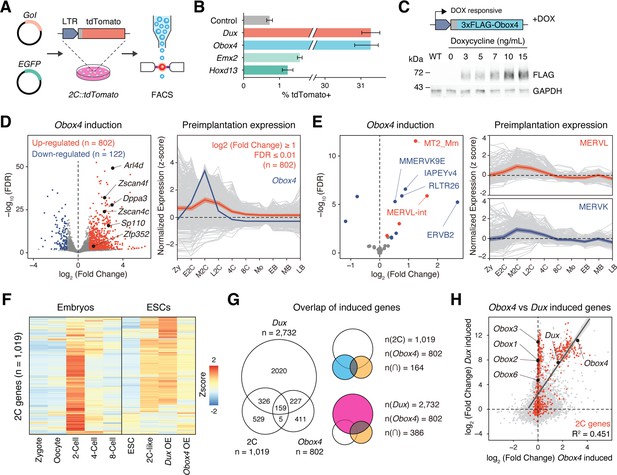
Obox4 and Dux induce 2C-gene expression in mouse embryonic stem cells (mESCs).
(A) Diagram of the 2C::tdTomato reporter assay. mESCs bearing the tdTomato expression cassette under the control of the MERVL long terminal repeat (LTR) promoter showed red fluorescence upon entering the 2C-like state. The expression of the transcription factor increased the 2C-like population, as detected using FACS. An EGFP expression plasmid was co-transfected with a gene of interest to normalize the transfection efficiency. (B) Boxplot showing normalized 2C-like cell percentage in 2C::tdTomato reporter mESCs overexpressing candidate pioneer factors. Dux and Obox4 potently induced a 2C-like state. n = 3 biological replicates. Error bars indicate standard deviations. (C) Upper panel: schematic of Obox4-inducible cell line construction. Lower panel: western blot showing OBOX4 level upon induction by different concentrations of doxycycline. Expression of OBOX4 was carried out in a dose-dependent manner. (D) Left panel: volcano plot of differentially expressed genes (DEGs) in mESCs with Obox4 induction for 48 hr. Representative 2C-genes are labeled with gene symbols. Right panel: expression profile of genes upregulated by Obox4 during embryogenesis. n = 3 biological replicates. (E) Left panel: volcano plot of differentially expressed transposable elements in mESCs with Obox4 induction for 48 hr. MERVL and MERVK elements were highlighted. Right panel: expression profile of MERVL and MERVK elements during preimplantation embryogenesis. n = 3 biological replicates. (F) Heatmaps of the expression of 2C-genes in preimplantation embryos, naturally occurred 2C-like mESCs, and induced 2C-like mESCs. (G) Venn diagram showing overlap of 2C-genes with genes induced by ectopic expression of Dux and Obox4 in mESCs. (H) Scatterplot showing per-gene expression changes in Dux-induced versus Obox4-induced mESCs. 2C-genes are highlighted in red.
-
Figure 2—source data 1
Original and uncropped blot for Figure 2C.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig2-data1-v2.zip
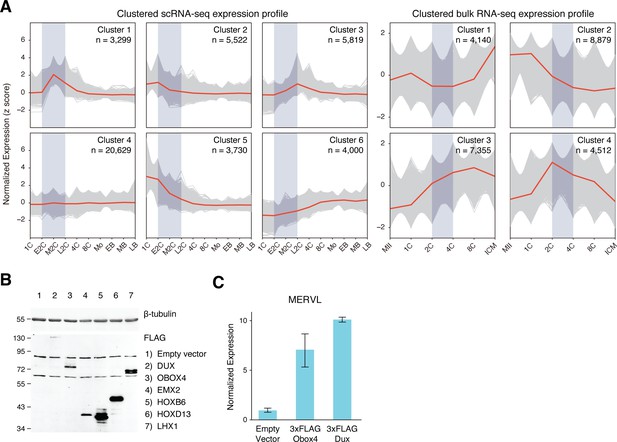
k-means clustering of reimplantation gene expression and MERVL induction by ectopic Obox4 and Dux expression.
(A) Expression profiles of k-means clustered gene sets during preimplantation development based on z-scored expression of scRNA-seq and bulk RNA-seq. Genes appeared in both scRNA-seq cluster 1 and bulk RNA-seq cluster 4 were designated as 2C-genes. (B) Western blot showing protein products of the candidate genes ectopically expressed in mouse embryonic stem cells (mESCs) at 18 hr post transfection. (C) Bar plot showing MERVL transcript level in mESCs at 18 hr post Dux or Obox4 transfection, detected by reverse transcription quantitative real-time PCR (RT-qPCR). n = 3 biological replicates. Error bars indicate standard deviations.
-
Figure 2—figure supplement 1—source data 1
Original and uncropped blot for Figure 2—figure supplement 1B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig2-figsupp1-data1-v2.zip
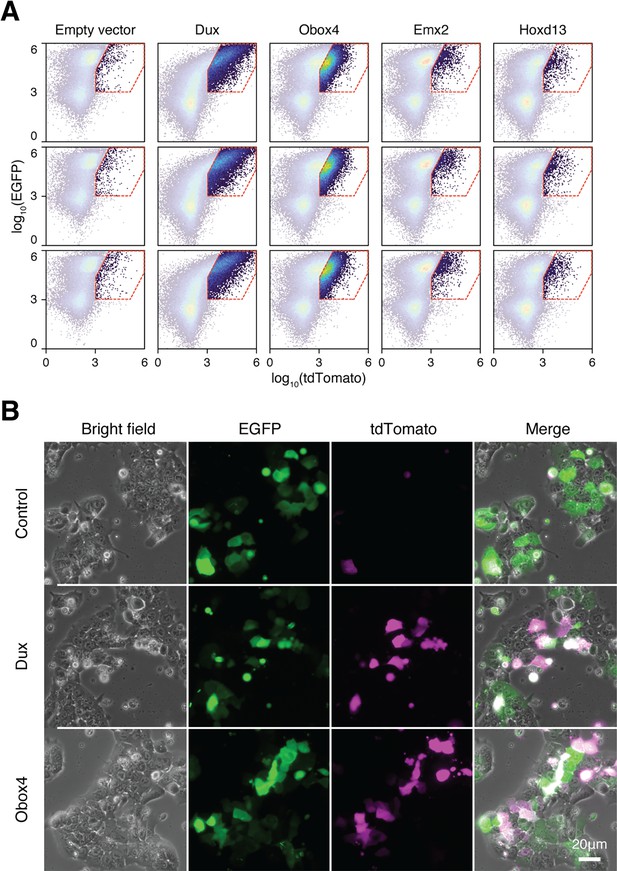
Ectopic expression of Obox4 and Dux activates MERVL reporter.
(A) FACS analysis of 2C::tdTomato cells co-transfected with candidate gene expression plasmids and EGFP. (B) Fluorescence microscopy of 2C::tdTomato mESCs transfected with empty plasmid, plasmid encoding DUX, or plasmid encoding OBOX4. Plasmids encoding EGFP were co-transfected to normalize the transfection efficiency in FACS analysis.
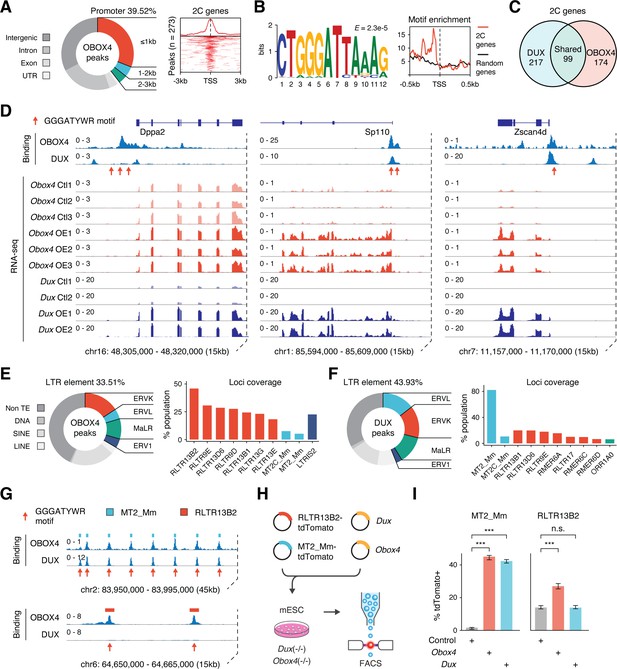
OBOX4 binds and activates 2C-specific long terminal repeat (LTR) elements.
(A) Left panel: pie chart displaying proportions of annotated genomic regions of the OBOX4 binding sites. Right panel: heatmap showing the OBOX4 binding site distribution near 2C-gene promoters. (B) Left panel: the predicted OBOX4 binding motif using the top 500 cleavage under targets and release using nuclease (CUT&RUN) peaks. Right panel: histograph showing the distribution of the predicted OBOX4 binding motif near 2C and random gene promoters. (C) Venn diagram showing the distinct and overlapping 2C-genes targeted by DUX and OBOX4. (D) Representative genomic track showing DUX and OBOX4 binding sites at the Dppa2, Sp110, and Zscan4d loci and their expression levels in Obox4 and Dux-induced mouse embryonic stem cells (mESCs). Read counts were CPM normalized. The OBOX4 binding sites overlapped with those of DUX. Dppa2, Sp110, and Zscan4d expression was upregulated upon Obox4 and Dux induction. (E) Left panel: pie- hart displaying proportions of annotated transposable elements (TEs) of the OBOX4 binding sites. Right panel: bar plot showing the top 10 OBOX4 covered LTR elements. (F) Left panel: pie chart displaying proportions of annotated TEs of the DUX binding sites. Right panel: bar plot showing the top 10 DUX covered LTR elements. (G) Representative genomic track showing DUX and OBOX4 binding sites at MT2_Mm and RLTR13B2. Read counts were CPM normalized. The OBOX4 binding sites overlapped with those of DUX at MT2_Mm loci, while RLTR13B2 was uniquely bound by OBOX4. (H) Schematic design of the LTR::tdTomato reporter assay. Plasmids bearing a tdTomato ORF downstream of MT2_Mm or RLTR13B2 were co-transfected with Dux or Obox4 expression plasmids. Activation of LTR elements resulted in an increased red fluorescence-positive mESC population, which was measured using FACS. EGFP expression plasmid was co-transfected into the culture, following which the green fluorescence-positive population was measured, to normalize the transfection efficiency. (I) Bar plots showing the percentage of red fluorescence-positive mESCs upon expression of Dux or Obox4. Both Dux and Obox4 activated MT2_Mm, whereas only Obox4 activated RLTR13B2.
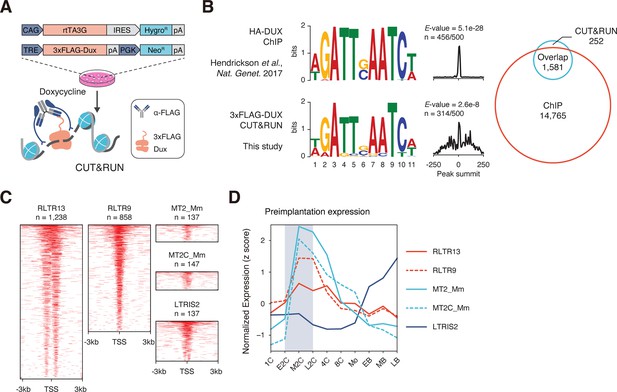
CUT&RUN captures DNA binding profiles of DUX and OBOX4.
(A) Schematic diagram of the cleavage under targets and release using nuclease (CUT&RUN) workflow using mouse embryonic stem cells (mESCs) with induced expression of FLAG-tagged DUX. mESCs bearing a doxycycline-inducible 3×FLAG-DUX expression cassette were induced with doxycycline for 24 hr. DUX-associated DNA was pulled down in the CUT&RUN assay using a high-affinity anti-FLAG antibody. (B) Left panel: comparison of DUX binding motif predicted with published ChIP data and CUT&RUN result from this study. Right panel: overlap of DUX binding peaks discovered with the two data sets. (C) Heatmaps showing the OBOX4 binding site distribution near top covered long terminal repeat (LTR) elements. (D) Expression profile of top 5 OBOX4 covered LTR elements during embryogenesis.
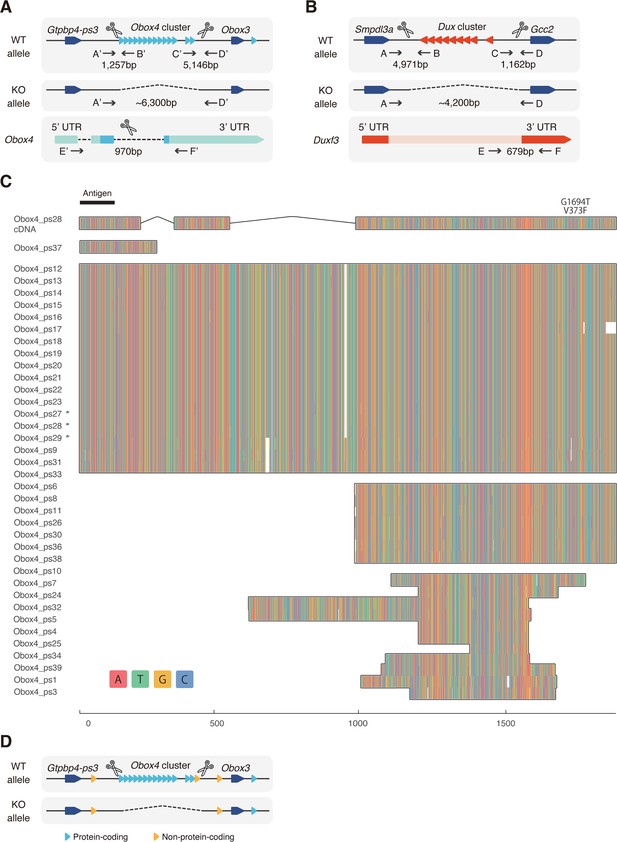
Knockout strategy of Obox4 and Dux clusters.
(A) Schematic diagram of producing Obox4 knockout mice using CRISPR-Cas9 (not drawn to scale). sgRNAs flanking the Obox4 cluster as well as the internal sequence of individual Obox4 were introduced into mouse zygotes with SpCas9, to knockout Obox4. Primers flanking Cas9 cutting sites (A’–D’) were designed to validate removal of target allele. Primers inside of Obox4 (E’–F’) were designed to quantify Obox4 copy number by qPCR. (B) Schematic diagram of producing Dux knockout mice using CRISPR-Cas9 (not drawn to scale). Previously reported sgRNAs flanking the Dux cluster were introduced into mouse zygotes with SpCas9, to knockout Dux. Primers flanking Cas9 cutting sites (A–D) were used to validate removal of target allele. Primers inside of Dux (E–F) were designed to detect translocation of Dux cluster. (C) Multiple sequence alignment of DNA sequence among the Obox4 copies. Full-length Obox4 with internal stop codons are marked by asterisks. (D) Schematic diagram of Obox4 cluster removal strategy and copy number quantification. Primers E’ and F’ targets all 15 protein-coding as well as 3 non-protein-coding Obox4 loci, among which 14 protein-coding and 1 non-protein-coding copies were subjected to knockout.
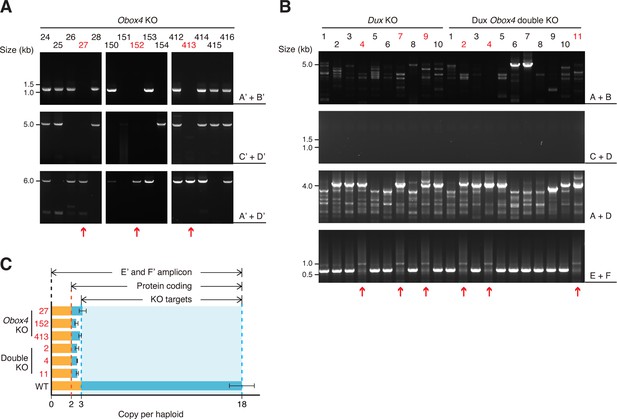
Validation of Obox4 and Dux knockout mouse embryonic stem cell lines.
(A) Agarose gel image of PCR-based genotyping of the Obox4 knockout allele in founder mouse embryonic stem cell (mESC) lines. The red arrows indicate cell lines bearing biallelic deletions of the Obox4 cluster. (B) Agarose gel image of PCR-based genotyping of the Dux knockout allele in founder mESC lines. The red arrows indicate cell lines without the genomic Dux sequences. (C) Bar plot showing copy number of Obox4 in the founder mESC lines analyzed in (A, B), as detected by qPCR using purified genomic DNA.
-
Figure 3—figure supplement 3—source data 1
Original and uncropped gels for Figure 3—figure supplement 3A.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig3-figsupp3-data1-v2.zip
-
Figure 3—figure supplement 3—source data 2
Original and uncropped gels for Figure 3—figure supplement 3B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig3-figsupp3-data2-v2.zip
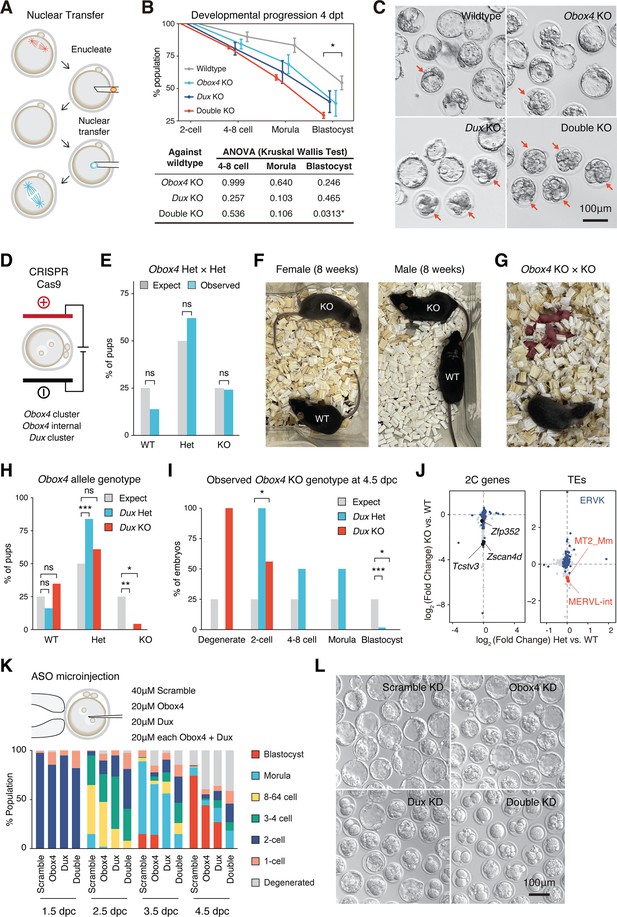
Concomitant loss of OBOX4 and DUX severely hinders zygotic genome activation (ZGA).
(A) Schematic representation of the somatic cell nuclear transfer (SCNT) experiment. The nuclei of knockout mouse embryonic stem cells (mESCs) were transferred into enucleated oocytes to generate zygotes with knockout genotype. (B) Upper panel: percent SCNT embryos developed to different stages at 4 days post nuclear transfer (dpt). Lower panel: p-value of non-parametric ANOVA among different genotypes and stages compared to wildtype SCNT embryos. Three independent experiments were conducted, with 150–200 embryos per condition. (C) Representative picture of SCNT embryos at 4 dpt. Morphologically abnormal blastocysts are highlighted. Blastocysts generated by double knockout mESCs were severely defective. (D) Schematic representation of CRISPR-Cas9-mediated Dux and Obox4 knockout mouse production. In vitro fertilized mouse zygotes were electroporated with pre-assembled CRISPR-Cas9 complex targeting Dux and Obox4 loci. (E) Bar plot showing genotype percentage of the pups delivered by Obox4Het intercrosses. Four litters delivered 29 pups, litter size 7.25 ± 1.26. ns p-value=0.9146, chi-square goodness-of-fit test. (F) Representative photos of Obox4KO and WT adult mice analyzed in (E). (G) Photo of Obox4KO intercross litter with live pups. (H) Bar plot showing genotype of Obox4 allele in the pups delivered by crossing of DuxKO/Obox4Het × DuxKO/Obox4Het or DuxKO/Obox4Het × DuxHet/Obox4Het. Nine litters delivered 54 pups, Dux heterozygous and knockout allele were present in 31 and 23 pups, respectively. **p-value=0.001306; *p-value=0.02218; chi-square goodness-of-fit test. (I) Bar plot showing observed percentages of different preimplantation stage embryos bearing Obox4 KO allele with Dux heterozygous or knockout allele at 4.5 days post coitum (dpc). Among the total 94 embryos assessed, 2 degenerated, 10 two-cell arrest, 2 4–8-cell arrest, 4 morula arrest embryos were observed at 4.5 dpc, whereas 76 developed to blastocyst. ***p-value=3.564 × 10–5; for two-cell; * p-value=0.04348; for blastocyst *p-value=0.02549; chi-square goodness-of-fit test. (J) Scatterplot showing expression log2 fold changes of genes (left panel) and transposable elements (TEs) (right panel) in DuxKO/Obox4KO and DuxKO/Obox4Het versus DuxKO/Obox4WT 2C embryos. 2C-genes and ERVK elements targeted by OBOX4 are highlighted in blue. MT2_Mm and MERVL-int are labeled. In total, 79 and 5 2C-genes were downregulated in DuxKO/Obox4KO and DuxKO/Obox4Het 2C embryos, respectively. (K) Upper panel: schematic representation of Dux and Obox4 knockdown experiments in preimplantation embryos. Male pronuclei of zygotes were microinjected with antisense oligonucleotide (ASO) targeting Dux or Obox4 transcripts. Lower panel: the percentages of embryonic stages observed at 1.5 dpc, 2.5 dpc, 3.5 dpc, and 4.5 dpc. (L) Representative picture of KD embryos at 4.5 dpc. No blastocyst was observed among double ASO knockdown embryos.
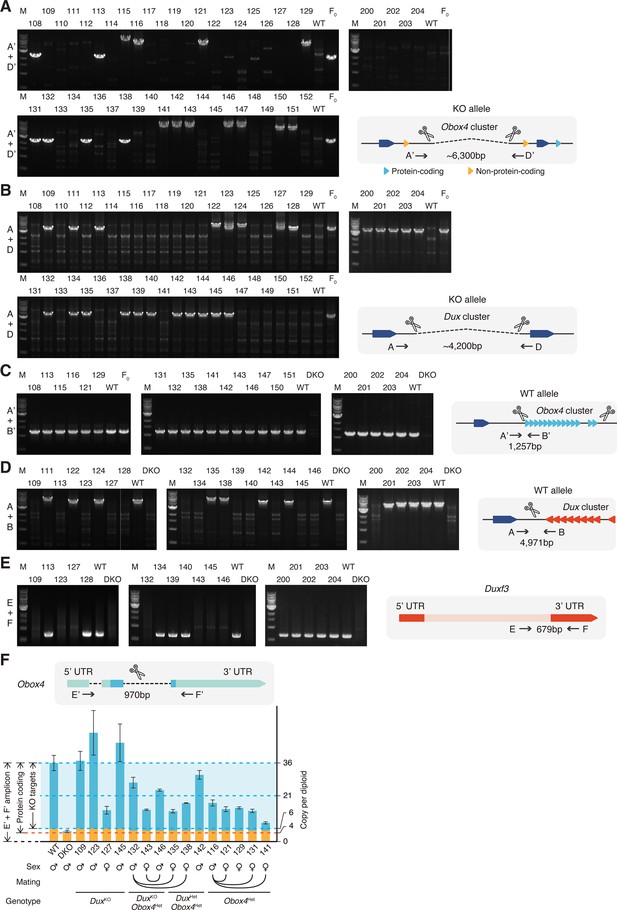
Genotyping of Obox4 and Dux knockout alleles in the F1 mice.
(A) Agarose gel image of PCR-based genotyping of the Obox4 knockout allele in the F1 mice. (B) Agarose gel image of PCR-based genotyping of the Dux knockout allele in the F1 mice. (C) Agarose gel image of PCR-based genotyping of the Obox4 cluster 5’ cutting site allele in the F1 mice. (D) Agarose gel image of PCR-based genotyping of the Dux cluster 5’ cutting site allele in the F1 mice. (E) Agarose gel image of PCR-based genotyping of the Dux internal sequence in the F1 mice. (F) Bar plot showing copy number of Obox4 in the F1 mice analyzed in (A–E), as detected by qPCR using purified genomic DNA.
-
Figure 4—figure supplement 1—source data 1
Original and uncropped gels of F1 mice 108–152 for Figure 4—figure supplement 1A–E.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp1-data1-v2.zip
-
Figure 4—figure supplement 1—source data 2
Original and uncropped gels of F1 mice 200–204 for Figure 4—figure supplement 1A–E.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp1-data2-v2.zip
Genotyping of Obox4 knockout allele in the F2 mice.
(A) Agarose gel image of PCR-based genotyping of the Obox4 knockout and Obox4 cluster 5’ cutting site wild-type allele in the F2 mice produced by intercrossing Obox4Het F1 mice. (B) Bar plot showing copy number of Obox4 in the F2 mice analyzed in (A), as detected by qPCR using purified genomic DNA.
-
Figure 4—figure supplement 2—source data 1
Original and uncropped gels of Figure 4—figure supplement 2A.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp2-data1-v2.zip
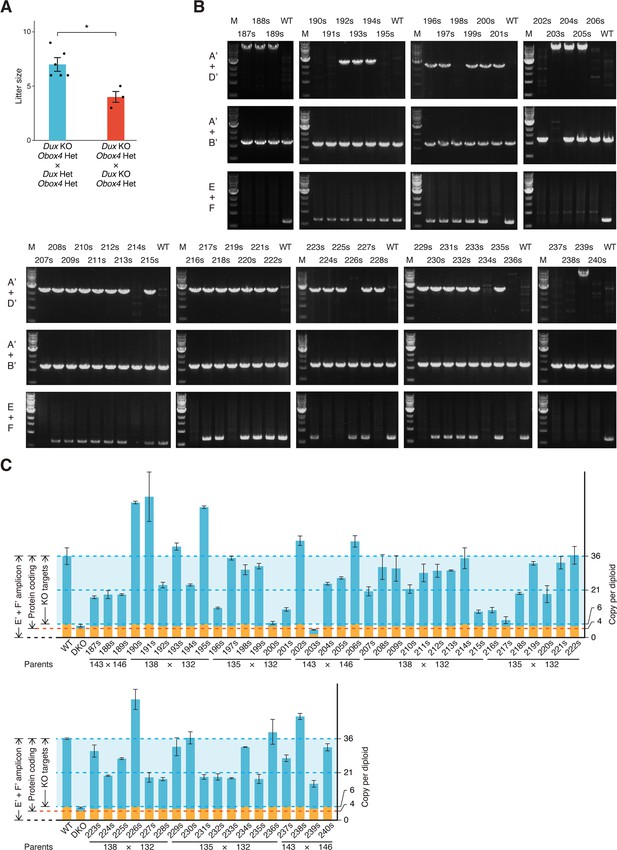
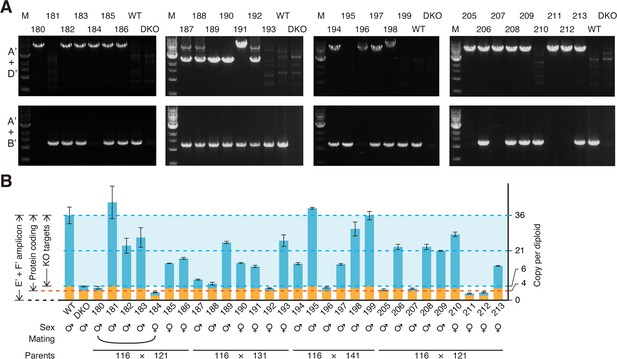
Genotyping of Obox4 and Dux knockout alleles in the F2 mice.
(A) Bar and dot plot showing litter sizes of intercross used to produce Dux/Obox4 double knockout mice. Each dot represents individual litter size. Six and three litters were analyzed for DuxKO/Obox4Het × DuxHet/Obox4Het and DuxKO/Obox4Het × DuxKO/Obox4Het mating, respectively. Crossing of DuxKO/Obox4Het × DuxKO/Obox4Het delivered reduced litter size than those of DuxKO/Obox4Het × DuxHet/Obox4Het. *p-value=0.01104, two-tailed Student’s t-test. Error bars indicate standard deviations. (B) Agarose gel image of PCR-based genotyping of the Obox4 knockout, Obox4 cluster 5’ cutting site allele, and Dux internal sequence in the F2 mice produced by intercrossing DuxKO/Obox4Het and/or DuxHet/Obox4Het F1 mice. (C) Bar plot showing copy number of Obox4 in the F2 mice analyzed in (A), as detected by qPCR using purified genomic DNA.
-
Figure 4—figure supplement 3—source data 1
Original and uncropped gels of F2 mice 187s–201s for Figure 4—figure supplement 3B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp3-data1-v2.zip
-
Figure 4—figure supplement 3—source data 2
Original and uncropped gels of F2 mice 202s–206s for Figure 4—figure supplement 3B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp3-data2-v2.zip
-
Figure 4—figure supplement 3—source data 3
Original and uncropped gels of F2 mice 207s–215s for Figure 4—figure supplement 3B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp3-data3-v2.zip
-
Figure 4—figure supplement 3—source data 4
Original and uncropped gels of F2 mice 216s–240s for Figure 4—figure supplement 3B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp3-data4-v2.zip
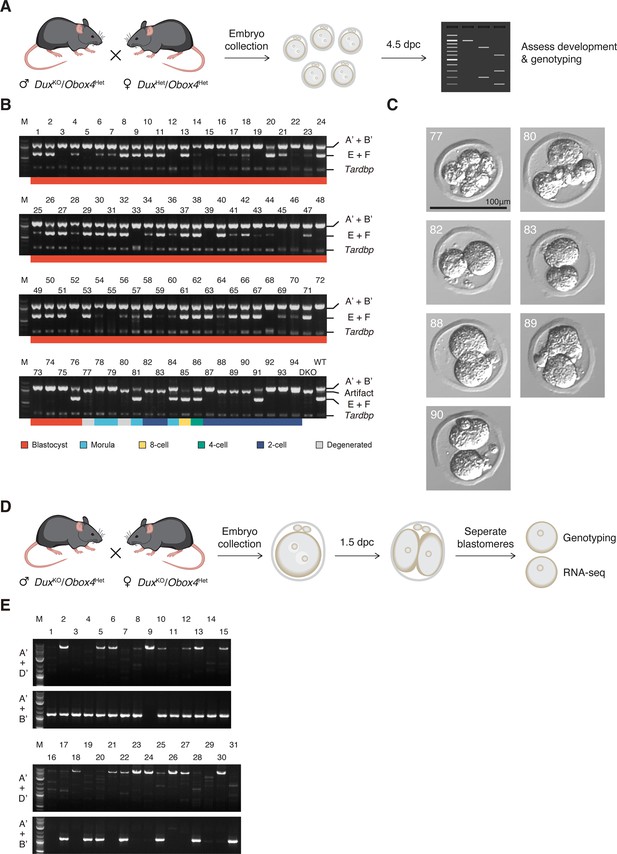
Genotyping of Obox4 and Dux knockout alleles in the embryos.
(A) Schematic diagram of knockout embryo production for development monitoring. DuxHet/Obox4Het female mice were mated with DuxKO/Obox4Het male mice. Embryos were collected at 0.5 days post coitum (dpc) and cultured in vitro until 4.5 dpc, followed by developmental assessment and genotyping. (B) Agarose gel image of multiplex PCR-based genotyping of Obox4 cluster 5’ cutting site allele, Dux internal sequence, and Tardbp internal sequence in the embryos produced by intercrossing DuxKO/Obox4Het and DuxHet/Obox4Het F1 mice. Note that multiplexing three sets of PCR primers produced artifact band in DuxKO/Obox4KO embryos. (C) Representative pictures of DuxKO/Obox4KO embryos. (D) Schematic diagram of knockout embryo production for RNA-seq. Embryos were collected at 0.5 dpc from DuxKO/Obox4Het mating pairs and cultured in vitro until two-cell stage at 1.5 dpc, blastomeres of each embryo were separated and subject to genotyping and RNA-seq respectively. (E) Agarose gel image of genotyping of Obox4 cluster 5’ cutting site allele in the embryos produced by intercrossing DuxKO/Obox4He mice.
-
Figure 4—figure supplement 4—source data 1
Original and uncropped gels for Figure 4—figure supplement 4B.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp4-data1-v2.zip
-
Figure 4—figure supplement 4—source data 2
Original and uncropped gels for Figure 4—figure supplement 4E.
- https://cdn.elifesciences.org/articles/95856/elife-95856-fig4-figsupp4-data2-v2.zip
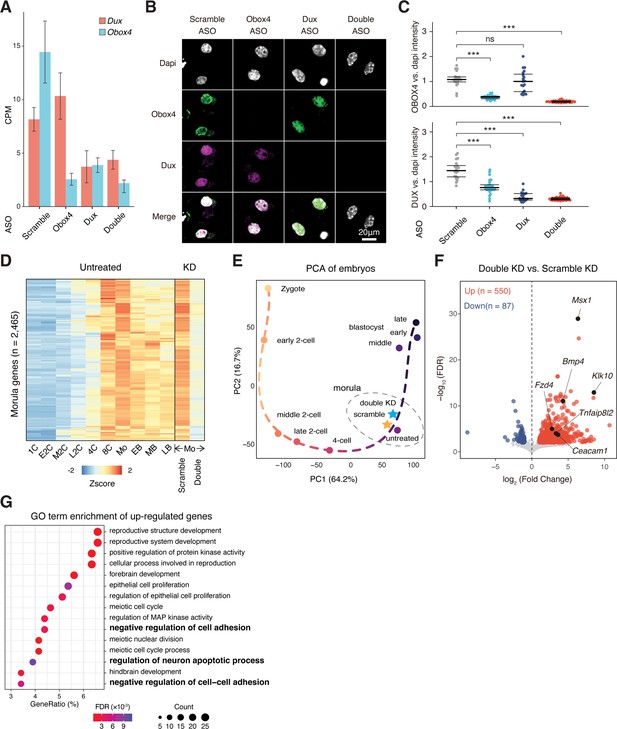
Obox4/Dux double knockdown embryos showed developmental defects.
(A) Transcript levels of Dux and Obox4 in antisense oligonucleotide (ASO)-injected embryos, as measured using RNA-seq. CPM, counts per million. (B) Representative images of immunofluorescence co-staining of OBOX4 and DUX in 2C embryos subjected to ASO-mediated knockdown. (C) Dot plots showing the per-nucleus intensity of OBOX4 and DUX normalized to DAPI. (D) Heatmaps of the expression of morula-genes in preimplantation embryos and knockdown morulae. Genes appeared in both scRNA-seq cluster 6 and bulk RNA-seq cluster 3 in Supplementary file 1a were designated as morula-genes. (E) Transcriptome-based PCA of knockdown morulae and preimplantation embryos. (F) Volcano plot of differentially expressed genes (DEGs) in double knockdown morulae compared with scramble knockdown. Selected apoptosis-associated genes are highlighted. (G) The enrichment of Gene Ontology (GO) terms of upregulated genes in double knockdown morulae.
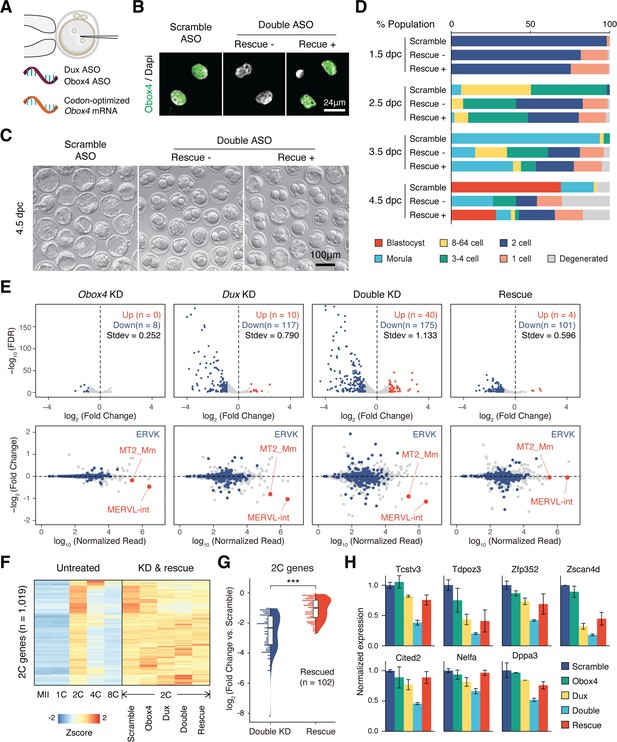
OBOX4 promotes zygotic genome activation (ZGA) in the absence of DUX.
(A) Schematic of the double knockdown rescue experiment. Male pronuclei of zygotes were injected with antisense oligonucleotide (ASO) targeting Obox4 and Dux transcripts as well as in vitro-transcribed codon-optimized Obox4 mRNA. (B) Immunofluorescence staining of OBOX4 in early two-cell embryos microinjected with scrambled ASO, double ASO (Obox4 and Dux), or double ASO with codon-optimized Obox4 mRNA. (C) Representative picture of knockdown and rescue embryos at 4.5 days post coitum (dpc). Codon-optimized Obox4 mRNA injection rescued blastocyst formation in ASO knockdown embryos. (D) The percentages of embryonic stages observed at 1.5 dpc, 2.5 dpc, 3.5 dpc, and 4.5 dpc. The plot represents the sum of three independent experiments, with 80–100 embryos per condition. (E) Upper panel: volcano plot showing the results of differentially expressed gene (DEG) analysis of 2C-genes in knockdown and rescue embryos compared to that in scramble ASO-injected embryos. Standard deviations of log2(fold-change) were used to represent the degree of transcriptome dysregulation. Stdev, standard deviation. Lower panel: MA plot of differential expression of transposable elements (TEs) in knockdown and recue embryos compared to that in scramble ASO-injected embryos. MERVK elements and MERVL are highlighted. n = 3 biological replicates. (F) Heatmaps of the expression of 2C-genes in preimplantation embryos, knockdown 2C-embryos, and rescue 2C-embryos. (G) Rain plot displaying the expression change distribution of rescued 2C-genes in double knockdown and recue 2C-embryos. (H) Bar plots showing the expression of representative 2C-genes in knockdown and recue embryos; n = 3 biological replicates.
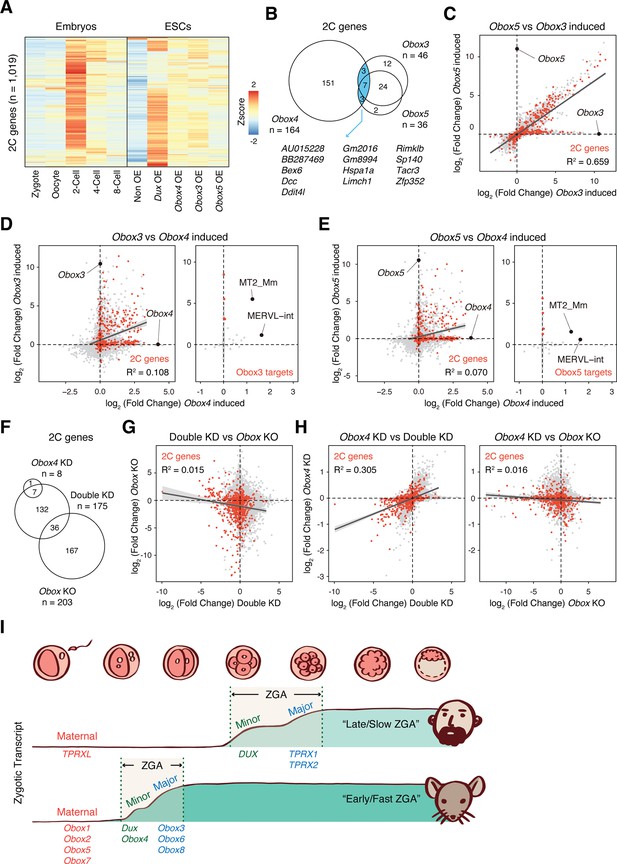
OBOX4-mediated DUX redundancy is distinct among the OBOX family.
(A) Heatmaps of the expression of 2C-genes in preimplantation embryos, non-induced mouse embryonic stem cells (mESCs), and Dux/Obox4/Obox3/Obox5-induced mESCs. (B) Venn diagram showing overlap of 2C-genes induced by ectopic expression of Obox3, Obox4, and Obox5 in mESCs. (C) Scatterplot showing per-gene expression changes in Obox3 versus Obox5-induced mESCs. 2C-genes are highlighted in red. (D) Scatterplot showing per-gene (left panel) and per-transposable element (right panel) expression changes in Obox3 and induced versus Obox4-induced mESCs. 2C-genes and Obox3-induced transposable elements (TEs) are highlighted in red. (E) Scatterplot showing per-gene (left panel) and per-transposable element (right panel) expression changes in Obox5 and induced versus Obox4-induced mESCs. 2C-genes and Obox3-induced TEs are highlighted in red. (F) Venn diagram showing overlap of 2C-genes downregulated upon Obox4 knockdown, Dux/Obox4 double knockdown, and Obox maternal-zygotic knockout. (G) Scatterplot showing per-gene expression changes in Obox maternal-zygotic knockout versus Dux/Obox4 double knockdown 2C embryos. 2C-genes are highlighted in red. (H) Scatterplot showing per-gene expression changes in Obox4 knockdown versus Dux/Obox4 double knockdown (left panel) and Obox maternal-zygotic knockout (right panel) 2C embryos. 2C-genes are highlighted in red. (I) Schematic model of different zygotic genome activation (ZGA) strategy employed by human and mouse. Obox family members with redundant functions are expressed in high dose at peri-ZGA stages to ensure rapid ZGA. Obox4 evolved as a divergent Obox family member that provides functional redundancy to Dux.
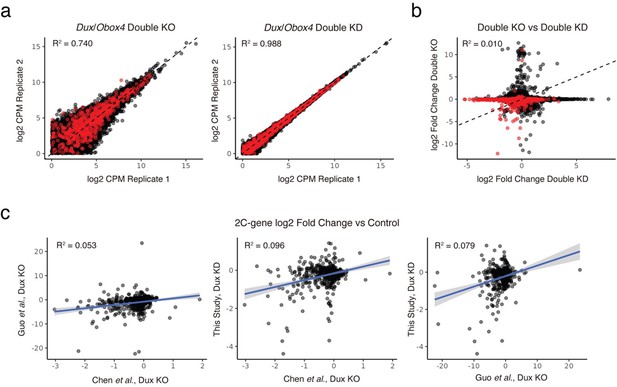
Comparison of transcriptome data in different KO and KD experiments.
a) Correlations between biological replicates of Dux/Obox4 DKO pseudo-bulk scRNA-seq (left panel) and ASO mediated Dux/Obox4 DKD RNA-seq (right panel). DKO samples bear higher inter-sample variance than DKD samples. 2C-genes are highlighted, inter-sample transcriptome correlations are measured by coefficient of determination (R2). b) Correlations of gene expression changes between Dux/Obox4 DKO and DKD RNA-seq. RNA-seq of DKD provides higher statistic power at discovering down-regulated 2C genes. 2C-genes are highlighted. c) Correlations of 2C-gene expression changes between published Dux KO RNA-seq and our ASO-mediated Dux KD RNA-seq. Our KD transcriptome shows higher agreement with the two KOs produced by Guo et al. and Chen et al. than between them each other.
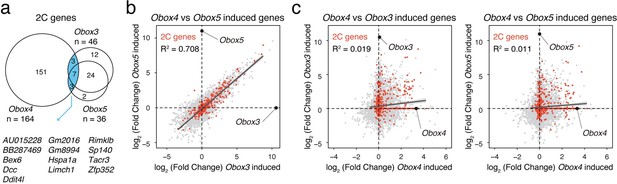
Comparison of Obox3/Obox4/Obox5 induced transcriptome.
a) Venn diagram showing overlap of 2C genes induced by ectopic expression of Obox3, Obox4, and Obox5 in mESCs. b) Scatterplot showing per-gene expression changes in Obox3 versus Obox5 induced mESCs. 2C-genes are highlighted in red. c) Scatterplot showing per-gene expression changes in Obox3 (left panel) and Obox5 (right panel) induced versus Obox4 induced mESCs. 2C-genes are highlighted in red.
Tables
Expression levels of all Obox family members in mouse pluripotent stem cell (PSC) and totipotent blastomere-like cell (TBLC).
| PSC (TPM) | TBLC (TPM) | ZGA Expression | |
|---|---|---|---|
| Oboxl Obox2 Obox3 Obox4 Obox5 Obox6 Obox7 Obox8 | 0.00 0.00 0.00 0.09 0.00 0.78 0.00 0.90 | 0.00 0.00 0.26 2.09 0.11 7.24 0.00 1.02 | TRUE TRUE TRUE TRUE TRUE FALSE TRUE FALSE |
Additional files
-
Supplementary file 1
List of Z-score and k-means clustering of genes based on preimplantation expression profiling data by Wu et al., 2016 (GSE66390) and Deng et al., 2014 (GSE45719).
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp1-v2.xlsx
-
Supplementary file 2
Deferential expression analysis results of genes and transposable elements in Obox4 and Dux overexpressing mESCs.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp2-v2.xlsx
-
Supplementary file 3
List OBOX4 and DUX bond genes.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp3-v2.xlsx
-
Supplementary file 4
Total numbers of annotated loci in each transposable element family and numbers of loci bond by OBOX4.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp4-v2.xlsx
-
Supplementary file 5
List of mating pairs and genotyping results of Dux/Obox4 knockout mice.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp5-v2.xlsx
-
Supplementary file 6
Genotyping and development monitoring results of preimplantation embryos produced by DuxKO/Obox4Het × DuxHet/Obox4Het mating pair.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp6-v2.xlsx
-
Supplementary file 7
Deferential expression analysis results of genes and transposable elements in DuxKO/Obox4Het and DuxKO/Obox4KO versus DuxKO/Obox4WT 2C embryos.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp7-v2.xlsx
-
Supplementary file 8
Combined deferential expression analysis results of genes and transposable elements in Obox3/4/5 overexpressing mESCs.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp8-v2.xlsx
-
Supplementary file 9
Combined deferential expression analysis results of genes and transposable elements in Dux/Obox4 knockdown and Obox family knockout 2C embryos.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp9-v2.xlsx
-
Supplementary file 10
Primer sequences used in this study.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp10-v2.xlsx
-
Supplementary file 11
Antisense oligonucleotides sequences used in this study.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp11-v2.xlsx
-
Supplementary file 12
sgRNA sequences used in this study.
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp12-v2.xlsx
-
Supplementary file 13
- https://cdn.elifesciences.org/articles/95856/elife-95856-supp13-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/95856/elife-95856-mdarchecklist1-v2.pdf





