Post-meiotic mechanism of facultative parthenogenesis in gonochoristic whiptail lizard species
Figures
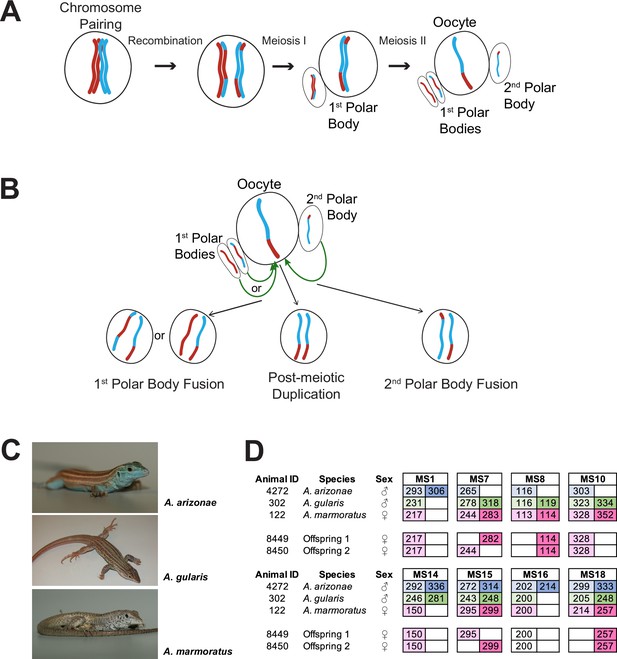
Overview.
(A) Schematic of canonical meiosis. Only one pair of homologous chromosomes is shown using red and blue to distinguish homologs. (B) Schematic of main mechanisms by which a diploid oocyte may be produced in the context of facultative parthenogenesis. First polar body fusion, second polar body fusion, or post-meiotic duplication of chromosomes in the haploid gamete. (C) Photographs of Aspidoscelis arizonae with characteristic blue ventral coloration (top), A. gularis with light spots in dark fields that separate light stripes on dorsum (middle), and A. marmoratus with light and dark reticulated pattern on dorsum (bottom). (D) Microsatellite analysis for the three co-housed animals and two offspring (ID 8449 and 8450) produced in this enclosure. Alleles are color-coded as follows: A. arizonae male (blue), A. gularis male (green), and A. marmoratus female (red). Differences in shading highlight the two alleles at heterozygous loci. Both offspring are homozygous at all loci with most alleles matching only maternal alleles. For MS16, offspring alleles are not shaded because of this allele being shared between the mother and the A. gularis male. Single nucleotide differences in size are common binning artifacts and, therefore, are not scored as different alleles.
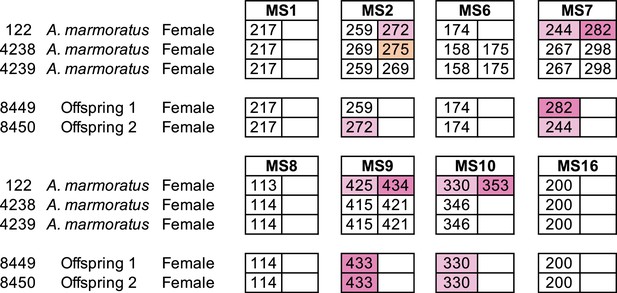
Microsatellite analysis of eight loci for the three female Aspidoscelis marmoratus within the enclosure and the two offspring hatched in January 2009.
Only alleles that are unique to a potential mother are colored. Single nucleotide differences in size are common binning artifacts and, therefore, are not scored as different alleles. The two offspring were homozygous for all eight markers. At three markers (MS7, MS9, MS10), both offspring inherited alleles only found in animal 122.
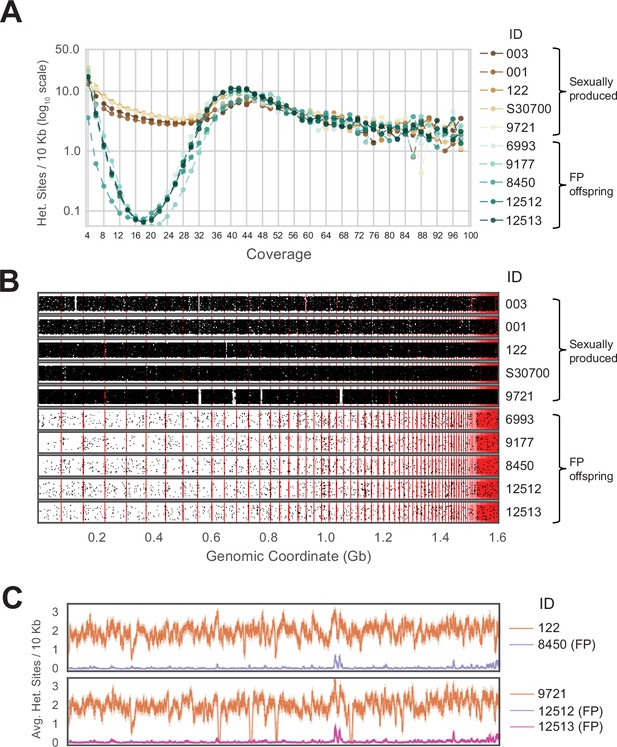
Genome-wide homozygosity in animals produced by facultative parthenogenesis.
(A) Effect of coverage on the apparent rate of heterozygosity based on evenly split read counts supporting two alleles. Analysis of whole-genome sequencing data for five sexually produced animals (ID 003, 001, 122, S30700, 9721) and five individuals produced by facultative parthenogenetic (FP) animals (ID 6993, 9177, 8450, 12512, 12513) were aligned to the reference genome. In FP animals, the number of heterozygous sites approaches zero for sites with mean coverage (x̄=18.37). (B) Scaffolds are ordered from largest to smallest on the x-axis. Red lines indicate borders between ordered scaffolds. Each black dot represents a heterozygous position in the genome defined by having a sequencing coverage equal to the average and equal support for only two alleles. The y-axis position of each data point is a random value between bounds of area shown to spread the data and better illustrate the density of heterozygous sites. (C) Average heterozygous sites, as defined in (B), per 10 Kb window for mothers (orange) and respective FP daughters (purple and pink).
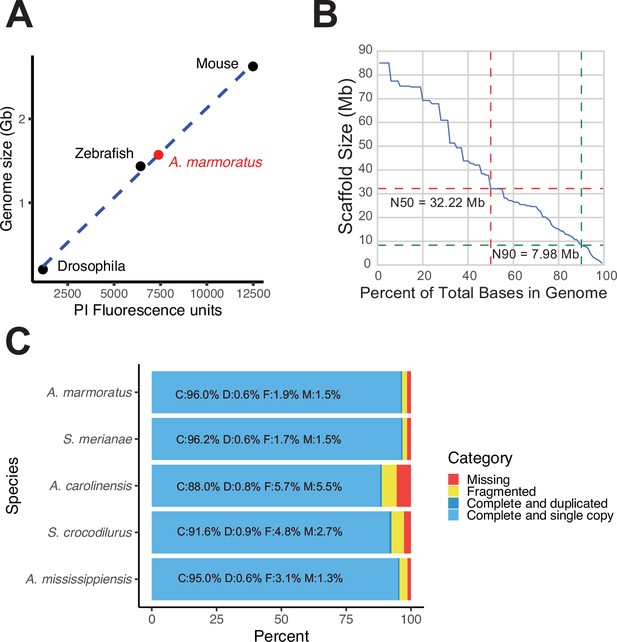
Genome assembly of Aspidoscelis marmoratus.
(A) Genome size estimation using fluorescence-activated cell sorting (FACs). The standard curve was generated using known genome sizes from fruit flies, zebrafish, and mouse, and then comparing fluorescent intensity with that of erythrocytes from A. marmoratus. (B) N(x)% plot shows the ordered scaffold lengths plotted against the cumulative fraction of the genome. Dashed lines mark the N50 and N90 values. (C) BUSCO analysis of 2586 conserved vertebrate coding genes for A. marmoratus, Salvator merianae, Anolis carolinensis, Shinisaurus crocodilurus, and Alligator mississippiensis. Values for S. crocodilurus and A. mississippiensis were taken from Gao et al., 2017.
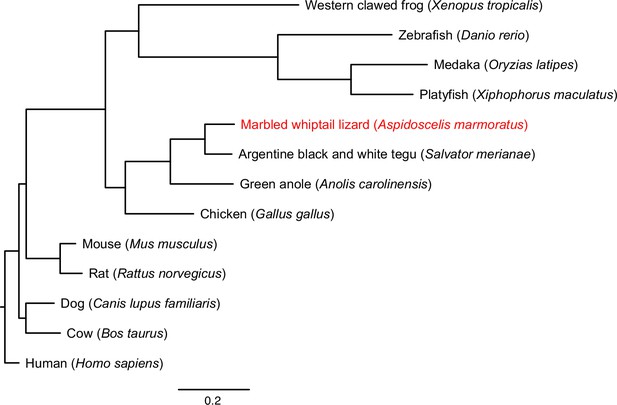
Maximum likelihood tree for 13 vertebrate genomes, based on 1333 single-copy BUSCOs detected across all species analyzed.
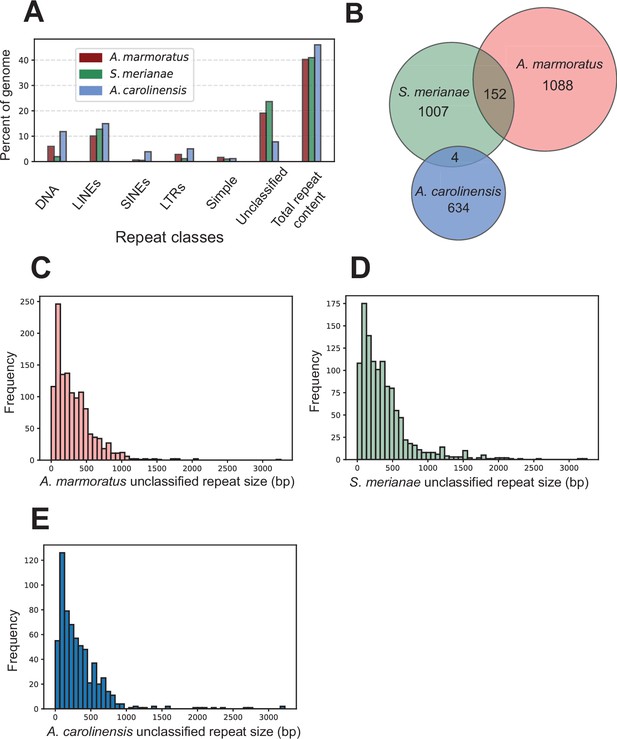
Identification of unclassified repeat elements in the A. marmoratus genome.
(A) Repetitive DNA content in Aspidoscelis marmoratus, Salvator merianae, and Anolis carolinensis genomes separated by repeat classes as defined by RepeatMasker. (B) Overlap of unclassified repetitive elements between the three genomes as in (A). (C–E) Distribution of the sizes of unclassified repetitive elements, respectively, for A. marmoratus, S. merianae, and A. carolinensis.
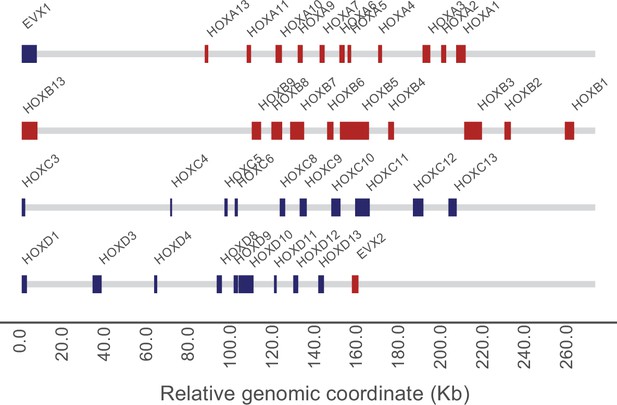
Aspidoscelis marmoratus HOX gene clusters.
Red blocks indicate a region of homology was found on the sense strand, blue indicates the antisense strand. EVX1 and EVX2 are also shown, due to their relevance to HOX cluster evolution.
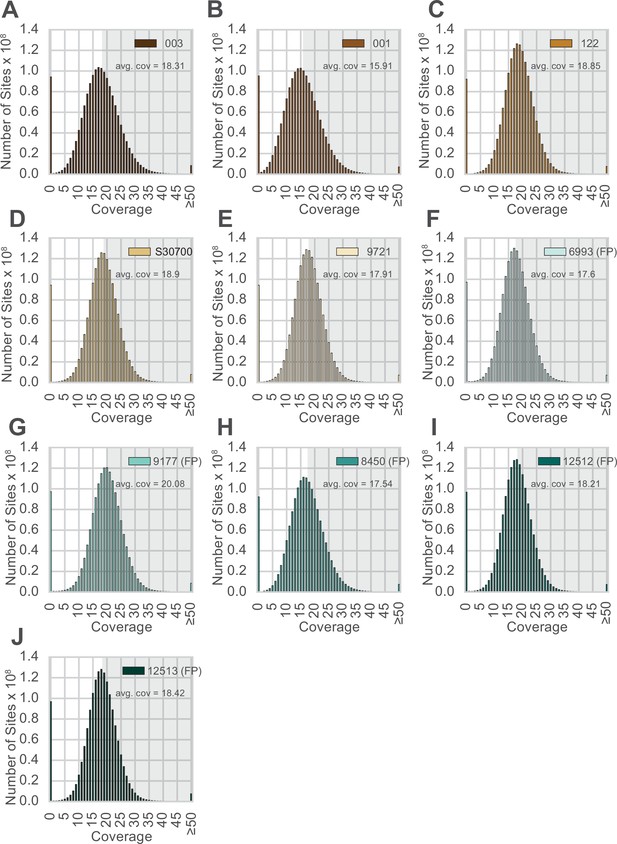
Distribution of sequence coverage across the genome for each animal.
(A–J) Animal IDs are shown in the top right corner of each panel, with animals produced by facultative parthenogenesis denoted with ‘facultative parthenogenesis (FP).’ The distributions were generated by examining the coverage at every position in the reference assembly. Regions with coverage greater than the mean are shaded in gray in each panel.
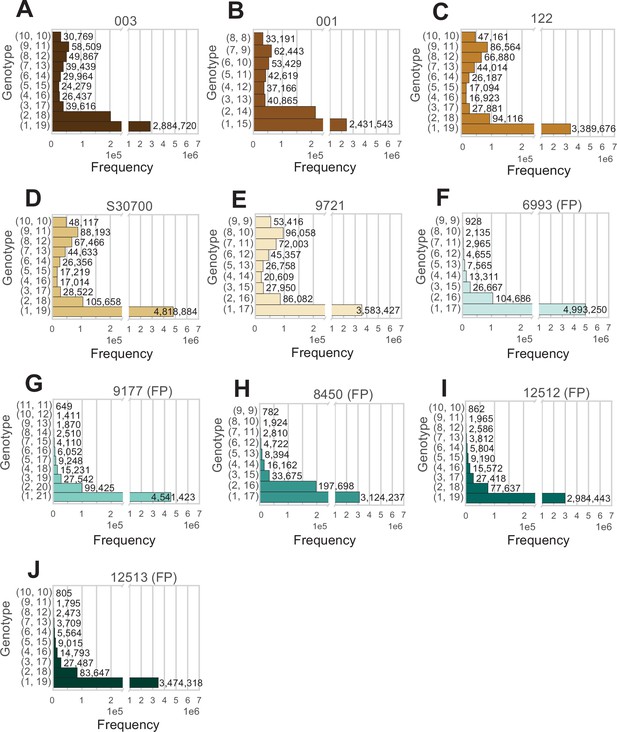
Analysis of all positions in the genome at average coverage, for which reads support exactly two alleles.
(A–J) The two numbers to the left of each row indicate how many reads support each allele. Animal IDs are shown above each panel, with animals produced by facultative parthenogenesis denoted with ‘facultative parthenogenesis (FP)’.
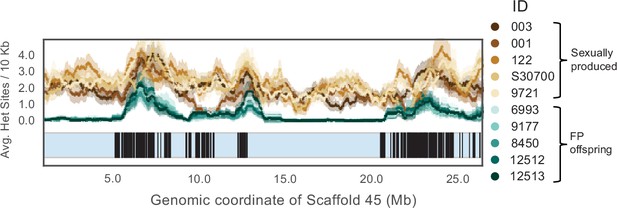
A 1Mb sliding window for Scaffold 45 showing the annotated vomeronasal 2 receptors homologs.
A scaffold ideogram is shown in blue and coordinates of 167 annotated V2r26 homologs are shown in black.
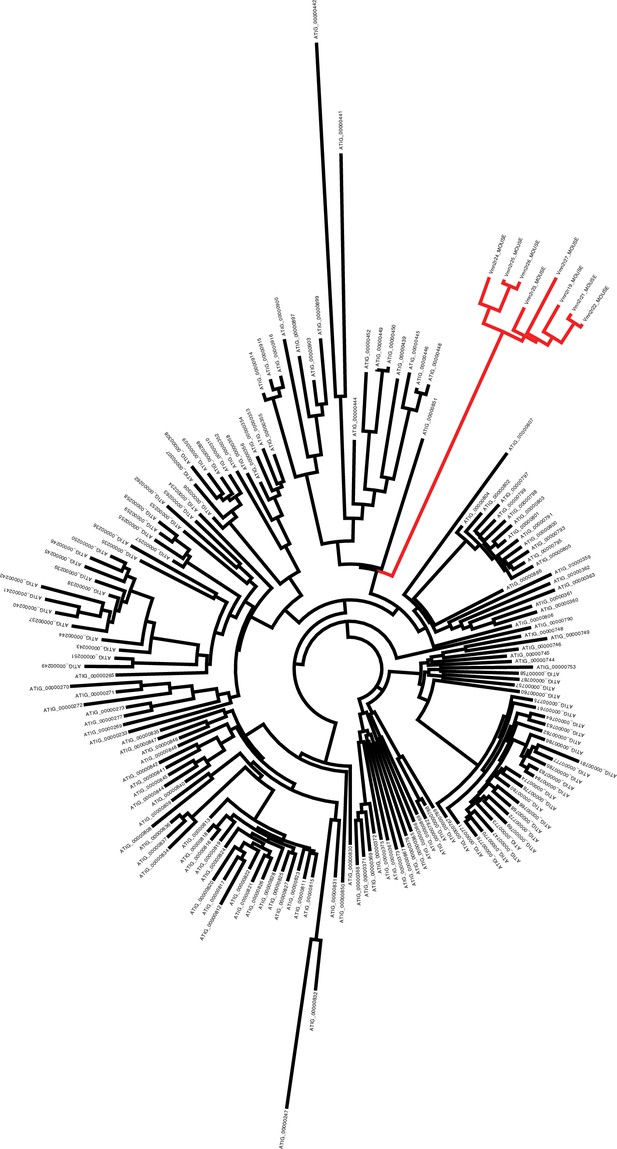
Phylogenetic tree showing the evolutionary distance between mouse V2Rs (red branches) and Aspidoscelis marmoratus homologs (black branches).
There are 10 additional copies of Vmn2r26 homologs found on Scaffold 45 not identified by MAKER2 (177 total), while genome-wide, we annotated 323 copies. Phylogenetic analysis of the 323 copies with eight other mouse vomeronasal two receptors show that the mouse receptors form their own distinct clade from A. marmoratus and there is substantial sequence diversity among the A. marmoratus homologs. A manual search for additional copies of Vmn2r26 resulted in 478 further hits. Vmn2r26 belongs to a family of receptors known as V2Rs and this estimate on the number of Vmn2r26 homologs ranks A. marmoratus as one of the species with the largest expansion of V2Rs (Shi and Zhang, 2007; Brykczynska et al., 2013).
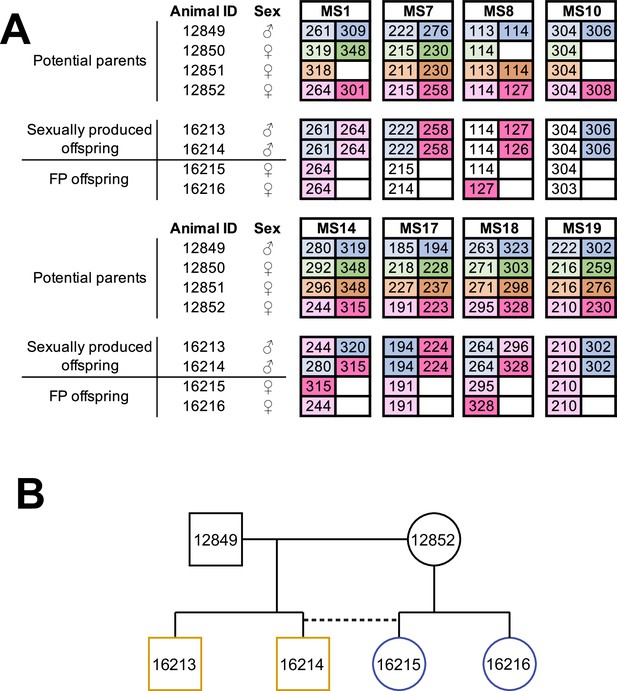
Facultative parthenogenesis is also found in Aspidoscelis arizonae.
(A) Microsatellite analysis for the four co-housed adult animals (ID 12849, 12850, 12851, 12852) and the four hatchlings (ID 16213, 16214, 16215, 16216) produced in this enclosure. Alleles are color-coded for each potential parent: 12849 male (blue), 12850 female (green), 12851 female (orange), 12852 female (red). Differences in shading highlight the two alleles at heterozygous loci. Offspring 16213 and 16214 are heterozygous at all loci, with most loci having one allele matching 12849 and one allele matching 12852. Offspring 16215 and 16216 are homozygous at all loci, with most alleles matching only the 12852 female. Non-shaded offspring alleles indicate ambiguous inheritance as multiple adult animals share the same allele. Single nucleotide differences in size are common binning artifacts and, therefore, are not scored as different alleles. (B) Pedigree shows the relationship between the four offspring. The single clutch of four contains both sexually (yellow) and facultative parthenogenetically (blue) produced offspring.
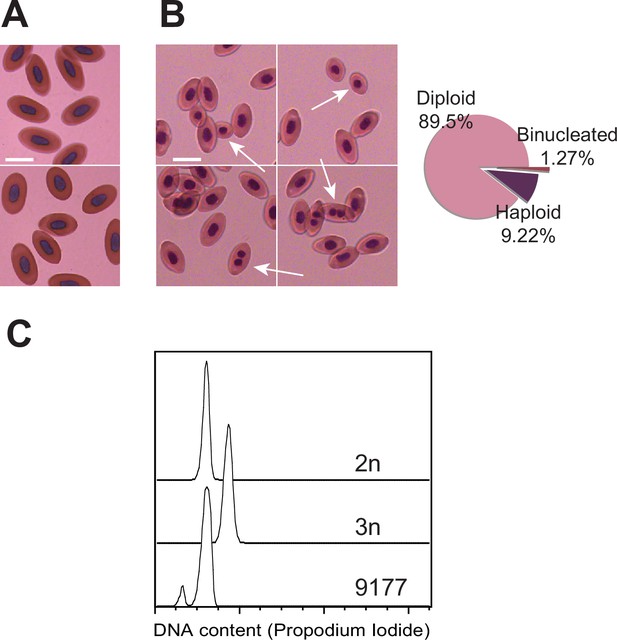
Detection of mixoploidy associated with facultative parthenogenesis.
(A) Giemsa staining of erythrocytes from a sexually produced Aspidoscelis marmoratus (ID 14744). All cells are diploid (n=601). Scale bar corresponds to 10 µm. (B) Giemsa staining of erythrocytes from a newly hatched facultative parthenogenetic A. marmoratus (ID 9177). Diploid (n=844), smaller haploid (n=87), and binucleated (n=12) cells are evident. Scale bar corresponds to 10 µm. (C) DNA content from erythrocytes determined by propidium iodide staining and detection by flow cytometry. Samples are from a sexually produced A. marmoratus (2n, ID 5358), an obligate triploid parthenogen A. exanguis (3n, ID 4950), and facultative parthenogenesis (FP) A. marmoratus (ID 9177). Number of events scored by flow cytometry were 44,145 (2n), 44,043 (3n), and 44,060 (9177). The FP 9177 sample contained an additional peak to the left of the 2 n peak (90.04%), indicating the presence of haploid cells (9.62%).
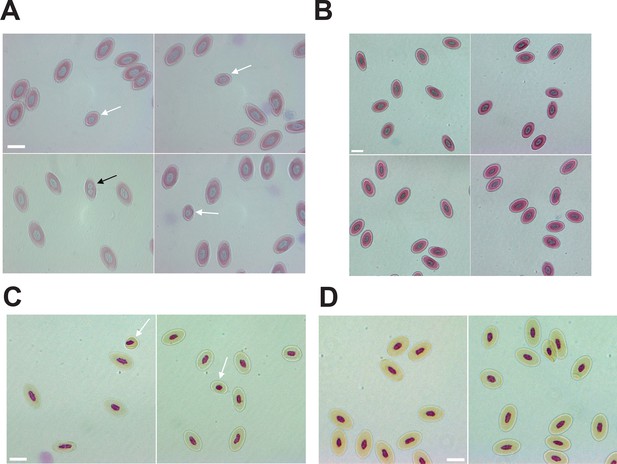
Mixoploidy detected in both A. marmoratus and A. arizonae.
(A) Giemsa staining of erythrocytes from a one-year-old facultative parthenogenetic Aspidoscelis marmoratus (ID 25384). Diploid (n=1661), smaller haploid (white arrow, n=17), and binucleated (black arrow, n=1) cells are evident. Scale bar corresponds to 10 μm. (B) Giemsa staining of erythrocytes from a sexually produced A. marmoratus (ID 23880). All cells are diploid (n=924). Scale bar corresponds to 10 μm. (C) Feulgen staining of erythrocytes from a one-year-old facultative parthenogenetic A. arizonae (ID 23507). Diploid (n=130) and smaller haploid (white arrow, n=2) cells are evident. Scale bar corresponds to 10 μm. (D) Feulgen staining of erythrocytes from a sexually produced A. arizonae (ID 26714). All cells are diploid (n=1506). Scale bar corresponds to 10 μm.
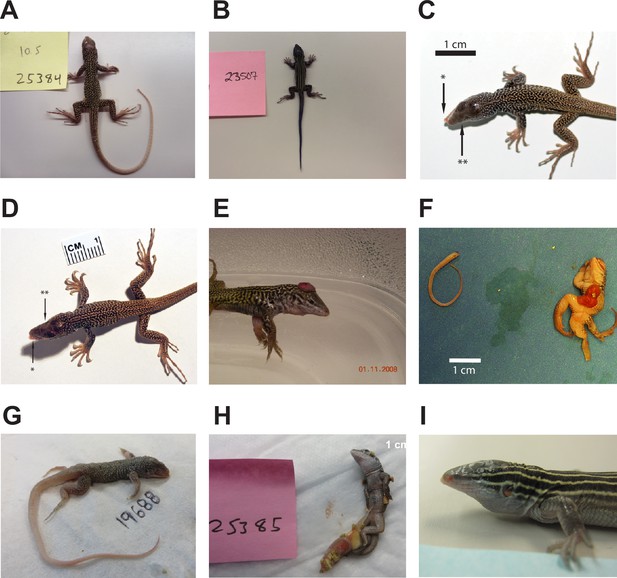

Animals produced by facultative parthenogenesis.
(A) Aspidoscelis marmoratus 25384. No abnormal phenotypes noted. (B) Aspidoscelis arizonae 23507. No abnormal phenotypes noted. (C) A. marmoratus 12512. Misalignment of jaws (*). Agenesis of left eye (**). (D) A. marmoratus 12513. Misalignment of jaws (*). Agenesis of right eye (**). (E) A. marmoratus 6993. Animal was cut from egg with exposed brain. (F) A. marmoratus 8394. Did not hatch. Missing a leg, failure of abdomen closure leading to exposed organs, face abnormalities, and hunched back. (G) A. marmoratus 19688. Did not hatch. Multiple craniofacial deformities. (H) A. marmoratus 25385. Partially emerged from egg with egg yolk still attached. (I) A. arizonae 16216. Agenesis of left eye.
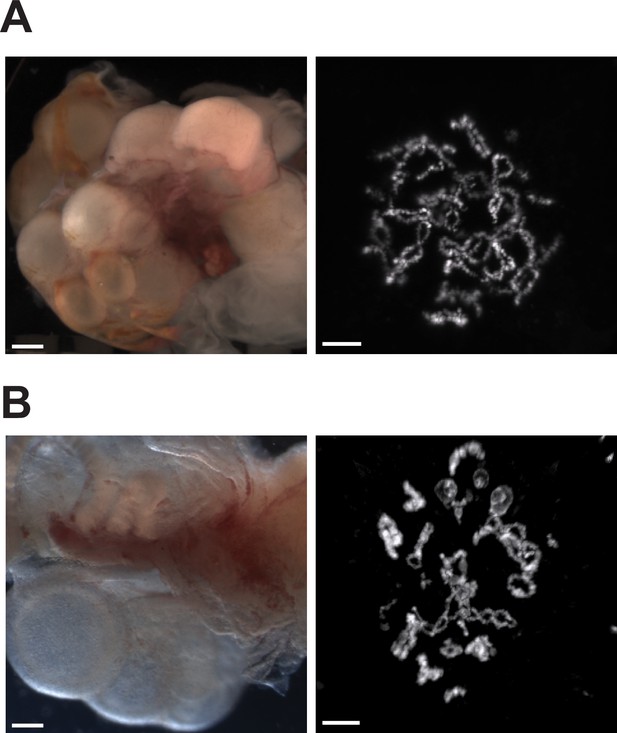
Ovaries of Aspidoscelis marmoratus facultative parthenogenesis (FP) animal 8450 and germinal vesicles of FP sister 8449 revealed no differences in structure and anatomy compared to fertile sexually reproducing animals.
(A) Ovary (left; ID 8450) and diplotene stage germinal vesicle (right; ID 8449). The lizard ovary image is taken with a Leica M205FA dissection microscope. Scale bar corresponds to 1 mm. The germinal vesicle is stained with DAPI. Scale bar corresponds to 10 μm. (B) Ovary (left; ID 13103) and diplotene stage germinal vesicle (right ID 5359) from sexually produced A. marmoratus. The lizard ovary image is taken with a Leica as described previously. Scale bar corresponds to 1 mm. The germinal vesicle is stained with Acridine Orange (0.01%). Scale bar corresponds to 10 μm.

Heterozygosity estimates of whiptail lizards collected in nature.
Percent heterozygosity estimates from reduced-representation sequencing (RAD-seq) for 321 whiptail lizards from 15 species. All individuals had an average coverage of at least 20. Each point is an individual, and percent heterozygosity was calculated only for sites where the coverage is equal to the average sequencing coverage. Five points with black borders indicate individuals (one angusticeps and four deppii) with low levels of heterozygosity. The heterozygosity of Aspidoscelis deppii ID LDOR30 (marked with arrow) is far less than that observed for individuals of the same species (Rosner’s Test for Outliers within deppii individuals, p<0.001), having only one called heterozygous position. Species (sample size): angusticeps (Booth et al., 2023), arizonae (Lenk et al., 2005), burti (Germano and Smith, 2010), costatus (Shibata et al., 2017), deppii (Ryba and Tirindelli, 1997), gularis (Olsen and Marsden, 1954), guttatus (Reeder et al., 2002), inornatus (Ryder et al., 2021), lineattissimus (Groot et al., 2003), marmoratus (Allen et al., 2018), occidentalis (Dudgeon et al., 2017), sackii (Groot et al., 2003), scalaris (Streisinger et al., 1981), sexlineatus (Germano and Smith, 2010), tigris (Card et al., 2021).
-
Figure 5—source data 1
Source data of Figure 5.
- https://cdn.elifesciences.org/articles/97035/elife-97035-fig5-data1-v1.xlsx
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Biological sample (Aspidoscelis spp.) | Erythrocytes, tail clippings, liver isolation | This paper, wild populations | See Ethics Statement | |
| Chemical compound, drug | Giemsa stain | Sigma | GS500 | 0.40% |
| Chemical compound, drug | Schiff’s reagent | Fisher Scientific | #SS32-500 | |
| Commercial assay or kit | Roche gDNA Isolation Kit | Roche | #11814770001 | |
| Commercial assay or kit | KAPA HTP kit | KAPA | KK8234 | |
| Commercial assay or kit | Nextera Mate-Pair Library Prep Kit | Illumina | FC-132–1001 | |
| Commercial assay or kit | TruSeq RNA Library Prep Kit v2 | Illumina | RS-122–2001 | |
| Sequence-based reagent | Oligos for MS analysis | This paper | See Supplementary file 7 | |
| Software, algorithm | Meraculous 2.0 | https://jgi.doe.gov/data-and-tools/software-tools/meraculous/ | RRID:SCR_010700 | |
| Software, algorithm | BUSCO 3.0.1 | https://busco.ezlab.org/ | RRID:SCR_015008 | |
| Software, algorithm | RAxML 8.2.11 | https://cme.h-its.org/exelixis/web/software/raxml/ | RRID:SCR_006086 | |
| Software, algorithm | RepeatModeler 1.0.11 | https://www.repeatmasker.org/RepeatModeler/ | RRID:SCR_015027 | |
| Software, algorithm | RepeatMasker 4.0.9 | https://www.repeatmasker.org/ | RRID:SCR_012954 | |
| Software, algorithm | BLAST 2.6.0 & 2.9.0+ | https://blast.ncbi.nlm.nih.gov/Blast.cgi | RRID:SCR_004870 | |
| Software, algorithm | BWA 0.7.15 | https://bio-bwa.sourceforge.net/ | RRID:SCR_010910 | |
| Software, algorithm | Picard 1.119 | https://broadinstitute.github.io/picard/ | RRID:SCR_006525 | |
| Software, algorithm | GATK 3.5 | https://gatk.broadinstitute.org/hc/en-us | RRID:SCR_001876 | |
| Software, algorithm | seqtk 1.2-r94 | https://github.com/lh3/seqtk Li, 2016 | 1.2-r94 | |
| Software, algorithm | pysam 0.12.0.1 | https://github.com/pysam-developers/pysam pysam-developers, 2017 | 0.12.0.1 | |
| Software, algorithm | pysamstats 0.24.3 | https://github.com/alimanfoo/pysamstats Miles, 2015 | 0.24.3 | |
| Software, algorithm | Trinity | https://github.com/trinityrnaseq/trinityrnaseq trinityrnaseq, 2015 | 2.0.6 | |
| Software, algorithm | seqclean | https://sourceforge.net/projects/seqclean | ||
| Software, algorithm | MAKER2 2.31.8 | https://www.yandell-lab.org/software/maker.html | RRID:SCR_005309 | |
| Software, algorithm | Interproscan 5.13–52.0 | https://interproscan-docs.readthedocs.io/en/latest/ | RRID:SCR_005829 | |
| Software, algorithm | Exonerate 2.4.0 | https://www.ebi.ac.uk/about/vertebrate-genomics/software/exonerate | RRID:SCR_016088 | |
| Software, algorithm | Geneious 10.1.3 | https://www.geneious.com/ | RRID:SCR_010519 | |
| Software, algorithm | Stacks 2.62 | https://catchenlab.life.illinois.edu/stacks/ | RRID:SCR_003184 | |
| Software, algorithm | Micromanager 1.4 | https://micro-manager.org/ | RRID:SCR_000415 | |
| Software, algorithm | Flowjo treestar | https://www.flowjo.com/ | RRID:SCR_008520 |
Additional files
-
Supplementary file 1
A. marmoratus genome assembly statistics.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp1-v1.docx
-
Supplementary file 2
Trinity assembly statistics.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp2-v1.docx
-
Supplementary file 3
MAKER2 summary.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp3-v1.docx
-
Supplementary file 4
Information for all A. marmoratus animals sequenced.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp4-v1.docx
-
Supplementary file 5
Whole genome sequencing alignment statistics.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp5-v1.docx
-
Supplementary file 6
Animals confirmed by microsatellite analysis to be of facultative parthenogenesis (FP) origin.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp6-v1.docx
-
Supplementary file 7
Microsatellite primer information.
- https://cdn.elifesciences.org/articles/97035/elife-97035-supp7-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/97035/elife-97035-mdarchecklist1-v1.docx





