DTX3L ubiquitin ligase ubiquitinates single-stranded nucleic acids
Figures
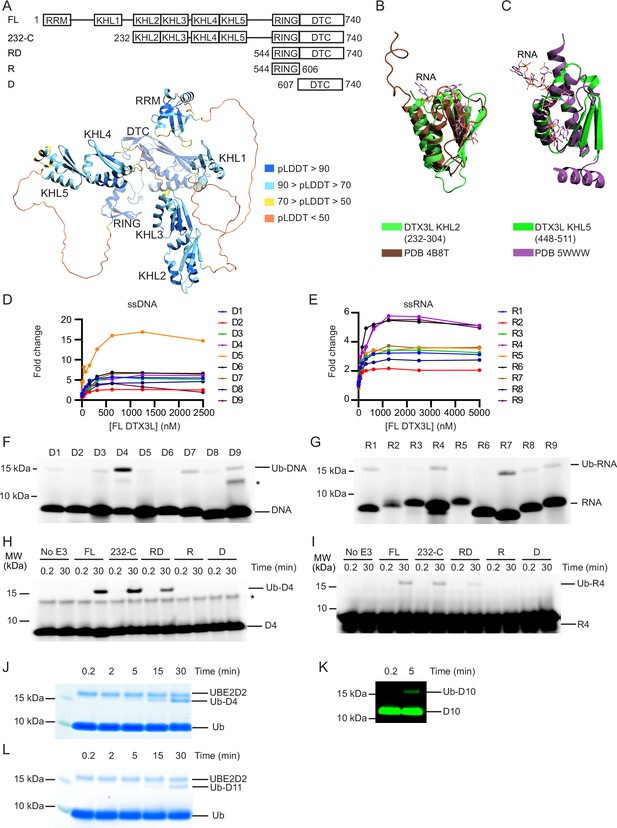
DTX3L catalyses the ubiquitination of single-stranded nucleic acids.
(A) Cartoon representation of the AlphaFold model of DTX3L. Domains are coloured according to model confidence. Domain architecture of DTX3L constructs is shown above the model. (B) DTX3L KHL2 (232-304) prediction shown in green overlaid with the third K Homology (KH) domain of KSRP in complex with AGGGU RNA sequence (PDB 4B8T) shown in brown. (C) DTX3L KHL5 (448-511) prediction shown in green overlaid with the MEX-3C KH1 domain in complex with GUUUAG RNA sequence (PDB 5WWW) shown in purple. (D) Fold change of fluorescence polarisation of 6-FAM-labelled ssDNA D1-9 upon titrating with full-length DTX3L. (E) As in (D) but with 6-FAM-labelled ssRNA R1-9. Data points for (D) and (E) are shown in Figure 1—source data 1 and 2, respectively. (F) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D1-9 oligonucleotides by FL DTX3L in the presence of E1, UBE2D2, Ub, Mg2+-ATP. (G) As in (F) but with 6-FAM-labelled ssRNA R1-9 oligonucleotides. (H) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L variants (FL, full length; RD, RING-DTC domains; R, RING domain; D, DTC domain) in the presence of E1, UBE2D2, Ub, Mg2+-ATP. (I) As in (H) but with 6-FAM-labelled ssRNA R4. (J) Coomassie-stained SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L-RD in the presence of E1, UBE2D2, Ub, and Mg2+-ATP. (K) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 5’ IRDye 800 ssDNA D10 by DTX3L-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP. (L) Coomassie-stained SDS-PAGE gel of in vitro ubiquitination of ssDNA D11 by DTX3L-RD in the presence of E1, UBE2D2, Ub, and Mg2+-ATP. Asterisks in (F) and (H) indicate contaminant bands from single-stranded DNA (ssDNA) or single-stranded RNA (ssRNA). Raw unedited and uncropped gel images for (F–L) are shown in Figure 1—source data 3 and 4, respectively.
-
Figure 1—source data 1
Fluorescence polarisation data related to Figure 1D.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-data1-v1.xlsx
-
Figure 1—source data 2
Fluorescence polarisation data related to Figure 1E.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-data2-v1.xlsx
-
Figure 1—source data 3
Raw unedited gels for Figure 1.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-data3-v1.zip
-
Figure 1—source data 4
Uncropped and labelled gels for Figure 1.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-data4-v1.zip
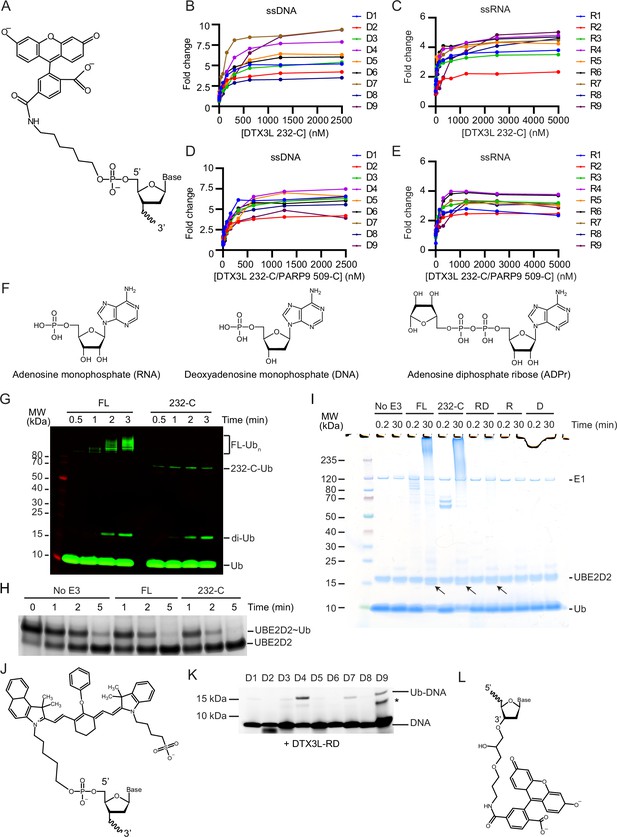
DTX3L binds and ubiquitinates single-stranded nucleic acids (ssNAs).
(A) Schematic of the 5’ 6-FAM modification. (B) Fold change of fluorescence polarisation of 6-FAM-labelled single-stranded DNA (ssDNA) D1-9 oligonucleotides upon titrating with DTX3L 232-C. (C) As in (B) but with 6-FAM-labelled ssRNA R1-9 oligonucleotides. (D) As in (B) but with DTX3L 232-C/PARP9 509-C. (E) As in (D) but with 6-FAM-labelled single-stranded RNA (ssRNA) R1-9. Data points for (B–E) are shown in Figure 1—figure supplement 1—source data 1–4, respectively. (F) Schematic of RNA, DNA, and ADPr. (G) Fluorescently detected SDS-PAGE gel of in vitro autoubiquitination of DTX3L FL and 232-C in the presence of E1, UBE2D2, fluorescent-labelled Ub and Mg2+-ATP. (H) Coomassie-stained SDS-PAGE gel of lysine discharge reactions showing the disappearance of UBE2D2~Ub over time in the presence of FL, 232-C or absence of E3. (I) Coomassie-stained SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L variants in the presence of E1, UBE2D2, Ub, Mg2+-ATP (from Figure 1H). Arrows indicate potential ~15 kDa Ub-DNA band. (J) Schematic of IRDye 800 modification. (K) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D1-9 in the presence of E1, UBE2D2, Ub, Mg2+-ATP by DTX3L-RD. (L) Schematic of the 3’ 6-FAM modification. Asterisks in (K) indicate contaminant band from ssDNA. Raw unedited and uncropped gel images of (G–I), (K) are shown in Figure 1—figure supplement 1—source data 5 and 6, respectively.
-
Figure 1—figure supplement 1—source data 1
Fluorescence polarisation data related to Figure 1—figure supplement 1B.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-figsupp1-data1-v1.xlsx
-
Figure 1—figure supplement 1—source data 2
Fluorescence polarisation data related to Figure 1—figure supplement 1C.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-figsupp1-data2-v1.xlsx
-
Figure 1—figure supplement 1—source data 3
Fluorescence polarisation data related to Figure 1—figure supplement 1D.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-figsupp1-data3-v1.xlsx
-
Figure 1—figure supplement 1—source data 4
Fluorescence polarisation data related to Figure 1—figure supplement 1E.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-figsupp1-data4-v1.xlsx
-
Figure 1—figure supplement 1—source data 5
Raw unedited gels for Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-figsupp1-data5-v1.zip
-
Figure 1—figure supplement 1—source data 6
Uncropped and labelled gels for Figure 1—figure supplement 1.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig1-figsupp1-data6-v1.zip
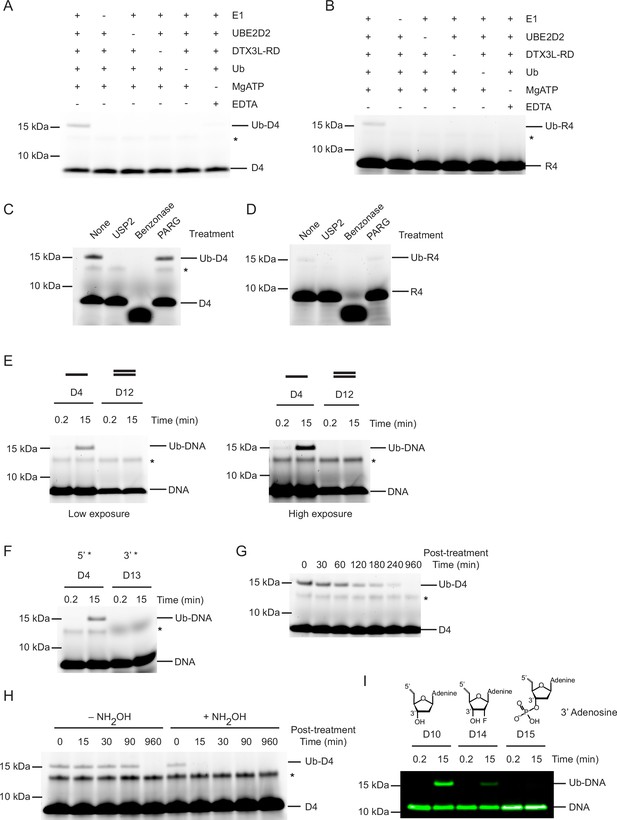
Ubiquitin (Ub) modification of single-stranded DNA (ssDNA) occurs at the 3’ hydroxyl at the 3’ end.
(A) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of ssDNA D4 in which E1, UBE2D2, DTX3L-RD, Ub, or Mg2+-ATP have been omitted. (B) As in (A) but with 6-FAM-labelled single-stranded RNA (ssRNA) R4. (C) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP subsequently treated with USP2, Benzonase, Poly(ADP-ribose) glycohydrolase (PARG) or not treated (None). (D) As in (C) but with 6-FAM-labelled ssRNA R4. (E) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 and dsDNA D12 oligonucleotides by DTX3L-RD (left panel) and at increased exposure (right panel) in the presence of E1, UBE2D2, Ub, Mg2+-ATP. (F) As in (E) but with 6-FAM-labelled ssDNA D4 (5’ label) and D13 (3’ label) oligonucleotides. (G) Fluorescently detected SDS-PAGE gel of in vitro ubiquitinated 6-FAM-labelled ssDNA D4 subsequently treated with pH 9.5 buffer for the times indicated. (H) Fluorescently detected SDS-PAGE gel of in vitro ubiquitinated 6-FAM-labelled ssDNA D4 subsequently treated with 1.5 M NH2OH at pH 9 for the times indicated. (I) As in (E) but with 5’ IRDye 800 ssDNA D10, D14 and D15 oligonucleotides. Asterisks in (A–C) and (E–H) indicate contaminant bands from ssDNA or ssRNA. Raw unedited and uncropped gel images are shown in Figure 2—source data 1 and 2, respectively.
-
Figure 2—source data 1
Raw unedited gels for Figure 2.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig2-data1-v1.zip
-
Figure 2—source data 2
Uncropped and labelled gels for Figure 2.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig2-data2-v1.zip
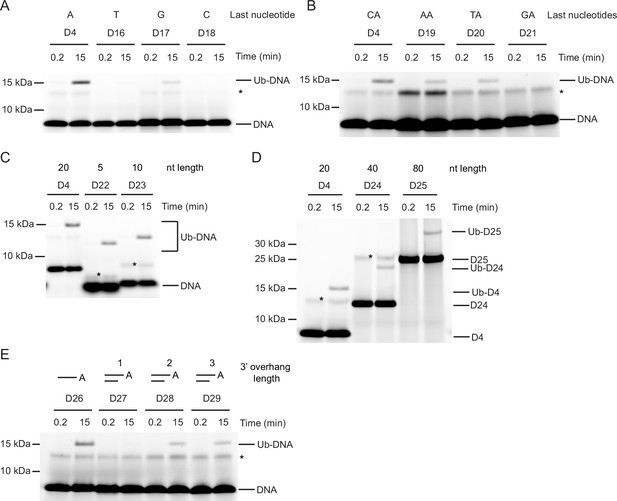
Nucleotide sequence requirements for ubiquitin (Ub)-DNA formation.
Fluorescently detected SDS-PAGE gel of in vitro ubiquitination (in the presence of E1, UBE2D2, Ub, and Mg2+-ATP) of (A) 6-FAM-labelled single-stranded DNA (ssDNA) D4, D16, D17 and D18 by DTX3L-RD. (B) 6-FAM-labelled ssDNA D4, D19, D20, and D21 by DTX3L-RD. (C) 6-FAM-labelled ssDNA D4, D22, and D23 by DTX3L-RD. (D) 6-FAM-labelled ssDNA D4, D24 and D25 by DTX3L-RD. (E) 6-FAM-labelled ssDNA D26, dsDNA D27, D28 and D29 by DTX3L-RD. Asterisks indicate contaminant bands from ssDNA. Raw unedited and uncropped gel images are shown in Figure 3—source data 1 and 2, respectively.
-
Figure 3—source data 1
Raw unedited gels for Figure 3.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig3-data1-v1.zip
-
Figure 3—source data 2
Uncropped and labelled gels for Figure 3.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig3-data2-v1.zip
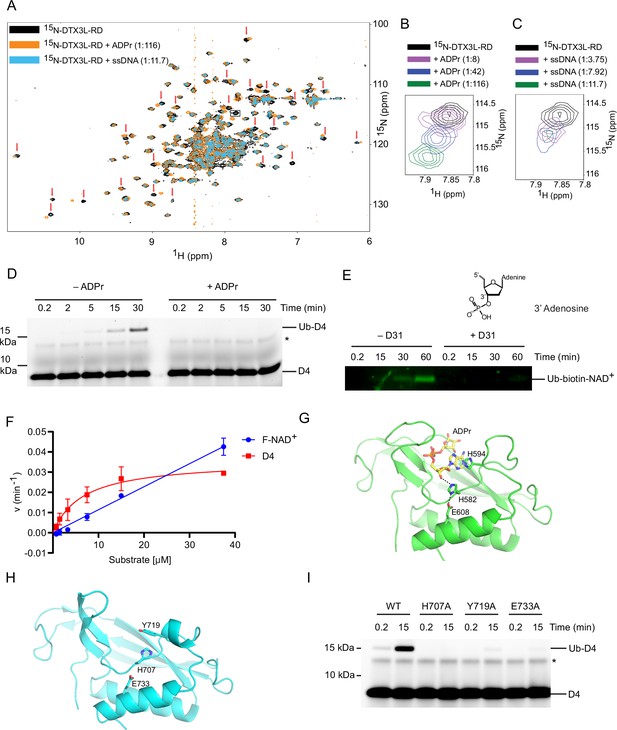
DTX3L DTC domain binds and facilitates ubiquitin (Ub)-DNA formation.
(A) 1H-15N heteronuclear single-quantum coherence (HSQC) spectra of 15N-DTX3L-RD (black), ADPr-15N-DTX3L-RD (orange), and single-stranded DNA (ssDNA) D30-15N-DTX3L-RD (blue). Red arrows indicate cross peaks that shift upon titrating with adenosine 5′-diphosphate (ADP)–ribose (ADPr) or ssDNA. (B) Close-up view of the cross peak indicated by the black box in (A) upon titration of specified molar ratios of ADPr with 15N-DTX3L-RD. (C) Close-up view of the cross peak indicated by the black arrow in (A) upon titration of specified molar ratios of ssDNA D30 with 15N-DTX3L-RD. (D) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP and treated with excess ADPr. (E) Western blot of in vitro ubiquitination of biotin-NAD+ by DTX3L-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP and treated with excess ssDNA D31. (F) Kinetics of Ub-D4 and Ub-F-NAD+ formation catalysed by DTX3L-RD. Data from two independent experiments (n=2) were fitted with the Michaelis–Menten equation and kcat/Km value for D4 (5457 M–1 min–1) was calculated. kcat/Km value for F-NAD+ (1190 M–1 min–1) was estimated from the slope of the linear portion of the curve. (G) Structure of DTX2-DTC domain (green) bound to ADPr (yellow) (PDB: 6Y3J). The sidechains of H582, H594, and E608 are shown in sticks. Hydrogen bonds are indicated by dotted lines. (H) Structure of DTX3L-DTC domain (cyan; PDB: 3PG6). The sidechains of H707, Y719, and E733 are shown in sticks. (I) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by full length DTX3L WT, H707A, Y719A, and E733A in the presence of E1, UBE2D2, Ub, Mg2+-ATP. Asterisks in (D) and (I) indicate contaminant band from ssDNA. Raw unedited and uncropped gel images of (D), (E) and (I) are shown in Figure 4—source data 1 and 2, respectively. Data points for (F) are shown in Figure 4—source data 3.
-
Figure 4—source data 1
Raw unedited gels for Figure 4.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig4-data1-v1.zip
-
Figure 4—source data 2
Uncropped and labelled gels for Figure 4.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig4-data2-v1.zip
-
Figure 4—source data 3
Kinetic data related to Figure 4F.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig4-data3-v1.xlsx
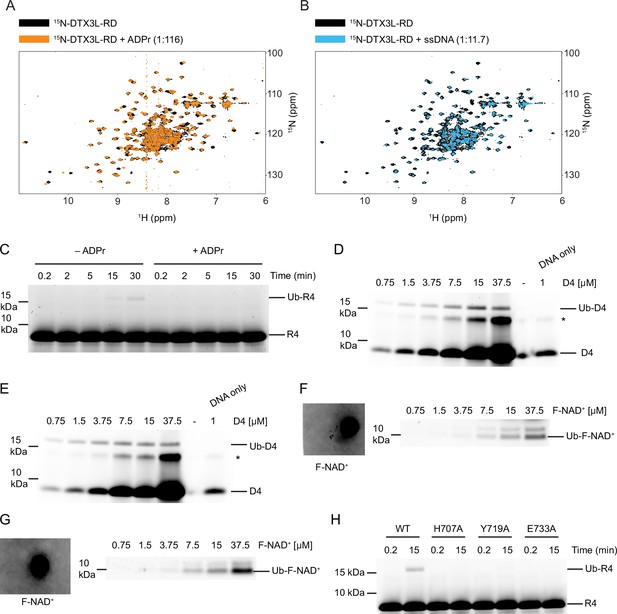
DTX3L-RD binds adenosine 5′-diphosphate (ADP)–ribose (ADPr) and single-stranded nucleic acids (ssNA).
(A) 1H-15N heteronuclear single-quantum coherence (HSQC) spectra of 15N-DTX3L-RD (black) and after the addition of ADPr (orange) (related to Figure 4A). (B) 1H-15N HSQC spectra of 15N-DTX3L-RD (black) and after the addition of single-stranded DNA (ssDNA) D30 (blue) (related to Figure 4A). (C) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssRNA R4 by DTX3L-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP and treated with excess ADPr. (D) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of increasing concentrations of 6-FAM-labelled ssDNA D4 by DTX3L-RD in the presence of E1, UBE2D2, Ub, and Mg2+-ATP. (E) Replicate of (D). (F) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of increasing concentrations of F-NAD+ by DTX3L-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP. A known volume of 100 µM F-NAD+ was pipetted onto Whatman filter paper and scanned alongside the gel for quantification. (G) Replicate of (F). (H) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssRNA R4 by FL DTX3L WT, H707A, Y719A, and E733A in the presence of E1, UBE2D2, Ub, and Mg2+-ATP. Asterisks in (D) and (E) indicate contaminant bands from ssDNA. Raw unedited and uncropped gel images are shown in Figure 4—figure supplement 1—source data 1 and 2, respectively.
-
Figure 4—figure supplement 1—source data 1
Raw unedited gels for Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig4-figsupp1-data1-v1.zip
-
Figure 4—figure supplement 1—source data 2
Uncropped and labelled gels for Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig4-figsupp1-data2-v1.zip
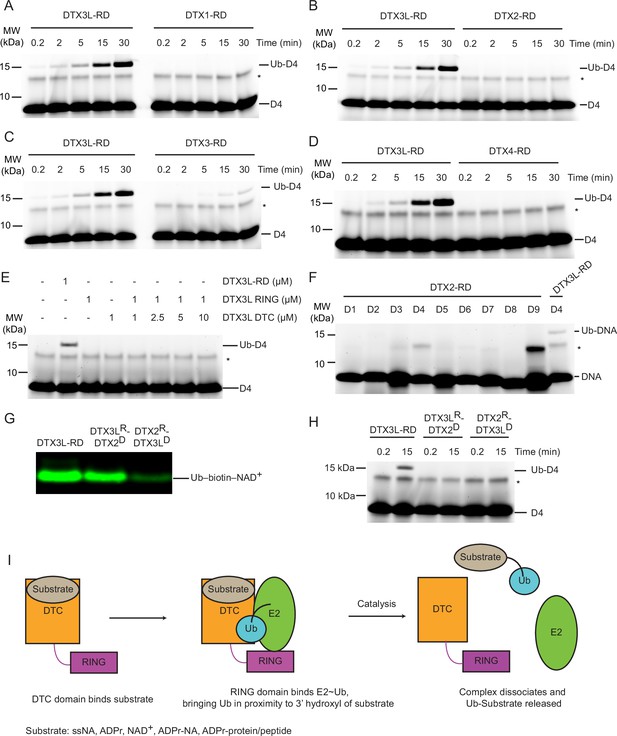
Select DTX RING-DTC domains catalyse ubiquitination of ssDNA.
(A) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L-RD or DTX1-RD in the presence of E1, UBE2D2, Ub, Mg2+-ATP. (B) As in (A) but with DTX2-RD. (C) As in (A) but with DTX3-RD. (D) As in (A) but with DTX4-RD. (E) As in (A) but with DTX3L-RD or DTX3L RING with increasing concentrations of DTX3L DTC. (F) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D1-9 by DTX2-RD. A reaction with DTX3L-RD and 6-FAM-labelled ssDNA D4 was included as a positive control. (G) Western blot of in vitro ubiquitination of biotin-NAD+ in the presence of E1, UBE2D2, Ub, Mg2+-ATP, NAD+, biotin-NAD+ with either DTX3L-RD, DTX3LR-DTX2D, or DTX2R-DTX3LD and separated by SDS-PAGE. (H) Fluorescently detected SDS-PAGE gel of in vitro ubiquitination of 6-FAM-labelled ssDNA D4 by DTX3L-RD, DTX3LR-DTX2D, or DTX2R-DTX3LD in the presence of E1, UBE2D2, Ub, Mg2+-ATP. (I) Schematic diagrams showing the proposed mechanism of ubiquitination of substrates by DTX3L-RD. Asterisks in (A–F) and (H) indicate contaminant bands from ssDNA. Raw unedited and uncropped gel images of (A–H) are shown in Figure 5—source data 1 and 2, respectively.
-
Figure 5—source data 1
Raw unedited gels for Figure 5.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig5-data1-v1.zip
-
Figure 5—source data 2
Uncropped and labelled gels for Figure 5.
- https://cdn.elifesciences.org/articles/98070/elife-98070-fig5-data2-v1.zip
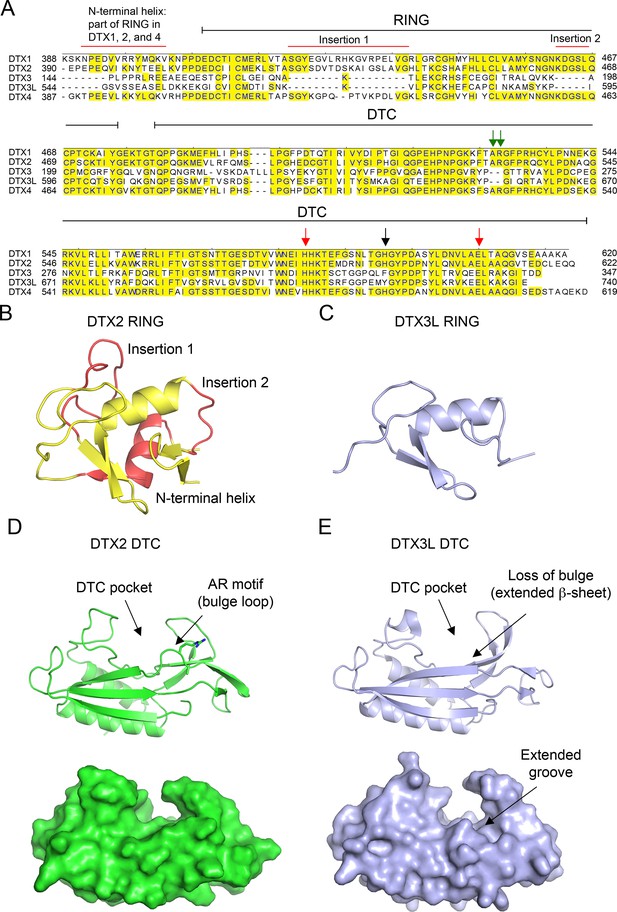
Properties of DELTEX (DTX) family DELTEX C-terminal (DTC) domains.
(A) Sequence alignment of human DTX family RING-DTC domains. Starting and ending residues of RING-DTC domains are indicated. Conserved residues are highlighted in yellow. A black arrow indicates residue that stacks with adenosine 5′-diphosphate (ADP)–ribose (ADPr), red arrows indicate conserved catalytic residues, and green arrows indicate AR motif present in DTX1, 2, and 4. Insertions are indicated. (B) Structure of DTX2 RING domain (from PDB: 6Y3J). The insertions and N-terminal helix are colored in red and the conserved RING domain region is colored yellow. (C) Alphafold2 model of DTX3L RING domain (light blue) displayed in the same orientation as in B. (D) Top: cartoon representation of the structure of DTX2 DTC domain (green; PDB: 6Y3J). Sidechains of the AR motif are shown as sticks. The DTC pocket that binds ADPr and the bulged AR motif are indicated by arrows. Bottom: as in the top panel but with surface representation. (E) Top: cartoon representation of the structure of DTX3L DTC domain (light blue; PDB: 3PG6). Bottom: as in the top panel but with surface representation. The absence of an AR motif in DTX3L causes a slight structural change near the ADPr-binding pocket, leading to the loss of the bulged loop, which results in a slightly extended groove.
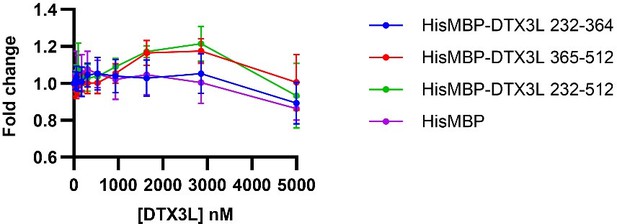
Fold change of fluorescence polarisation of 6-FAM-labelled ssDNA D4 upon titrating with DTX3L variants.
DTX3L KH domain fragments were expressed with a N-terminal His-MBP tag to increase the molecular weight to enhance the signal.
Tables
List of nucleotide sequences used in this study.
| Name | Type | Sequence | Modifications |
|---|---|---|---|
| D1 | ssDNA | 5’ TGTTTGTTTGTTTGTTTGTT 3’ | 5’ FAM |
| D2 | ssDNA | 5’ GCGCGCGCGCGCGCGCGCGC 3’ | 5’ FAM |
| D3 | ssDNA | 5’ AGTGAGTGAGTGAGTGAGTG 3’ | 5’ FAM |
| D4 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | 5’ FAM |
| D5 | ssDNA | 5’ AGAGAGAGAGAGAGAGAGAG 3’ | 5’ FAM |
| D6 | ssDNA | 5’ TCTCTCTCTCTCTCTCTCTC 3’ | 5’ FAM |
| D7 | ssDNA | 5’ TTTTTTTTTTTTTTTTTTTT 3’ | 5’ FAM |
| D8 | ssDNA | 5’ GTGCTGCGCTGCGCTGTGCT 3’ | 5’ FAM |
| D9 | ssDNA | 5’ AAAAAAAAAAAAAAAAAAAA 3’ | 5’ FAM |
| D10 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | 5’ IRDye 800 |
| D11 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | None |
| D12 | dsDNA | 5’ CAACAACAACAACAACAACA 3’ + 3’ TGTTGTTGTTGTTGTTGTTG 5’ | 5’ FAM None |
| D13 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | 3’ FAM |
| D14 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | 5’ IRD800 3’ 2FA |
| D15 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | 5’ IRDye 800 3’ Phos |
| D16 | ssDNA | 5’ CAACAACAACAACAACAACT 3’ | 5’ FAM |
| D17 | ssDNA | 5’ CAACAACAACAACAACAACG 3’ | 5’ FAM |
| D18 | ssDNA | 5’ CAACAACAACAACAACAACC 3’ | 5’ FAM |
| D19 | ssDNA | 5’ CAACAACAACAACAACAAAA 3’ | 5’ FAM |
| D20 | ssDNA | 5’ CAACAACAACAACAACAATA 3’ | 5’ FAM |
| D21 | ssDNA | 5’ CAACAACAACAACAACAAGA 3’ | 5’ FAM |
| D22 | ssDNA | 5’ CAACA 3’ | 5’ FAM |
| D23 | ssDNA | 5’ AACAACAACA 3’ | 5’ FAM |
| D24 | ssDNA | 5’ AACAACAACAACAACAACAACAACAACAACAACAACAACA 3’ | 5’ FAM |
| D25 | ssDNA | 5’ CAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACAACA 3’ | 5’ FAM |
| D26 | ssDNA | CGGCACATCACTCTTCAACA | 5’ FAM |
| D27 | dsDNA | 5’ CGGCACATCACTCTTCAACA 3’ + 5’ GTTGAAGAGTGATGTGCCG 3’ | 5’ FAM None |
| D28 | dsDNA | 5’ CGGCACATCACTCTTCAACA 3’ + 5’ TTGAAGAGTGATGTGCCG 3’ | 5’ FAM None |
| D29 | dsDNA | 5’ CGGCACATCACTCTTCAACA 3’ + 5’ TGAAGAGTGATGTGCCG 3’ | 5’ FAM None |
| D30 | ssDNA | CAACA | None |
| D31 | ssDNA | 5’ CAACAACAACAACAACAACA 3’ | 3’ Phos |
| R1 | ssRNA | 5’ UGUUUGUUUGUUUGUUUGUU 3’ | 5’ FAM |
| R2 | ssRNA | 5’ GCGCGCGCGCGCGCGCGCGC 3’ | 5’ FAM |
| R3 | ssRNA | 5’ AGUGAGUGAGUGAGUGAGUG 3’ | 5’ FAM |
| R4 | ssRNA | 5’ CAACAACAACAACAACAACA 3’ | 5’ FAM |
| R5 | ssRNA | 5’ AGAGAGAGAGAGAGAGAGAG 3’ | 5’ FAM |
| R6 | ssRNA | 5’ UCUCUCUCUCUCUCUCUCUC 3’ | 5’ FAM |
| R7 | ssRNA | 5’ UUUUUUUUUUUUUUUUUUUU 3’ | 5’ FAM |
| R8 | ssRNA | 5’ GUGCUGCGCUGCGCUGUGCU 3’ | 5’ FAM |
| R9 | ssRNA | 5’ AAAAAAAAAAAAAAAAAAAA 3’ | 5’ FAM |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Recombinant DNA reagent (Homo sapiens) | pGEX4T-3 TEV DTX3L FL | This paper | Uniprot: Q8TDB6-1 | Codon-optimised synthetic gene |
| Recombinant DNA reagent (Homo sapiens) | pGEX4T-3 TEV DTX3L FL mutants | This paper | Codon-optimised synthetic gene H707A Y719A E733A | |
| Recombinant DNA reagent (Homo sapiens) | pGEX4T-3 TEV DTX3L 232-C | This paper | Codon-optimised synthetic gene | |
| Recombinant DNA reagent (Homo sapiens) | pGEX4T-3 TEV DTX3L 607-C | This paper | Codon-optimised synthetic gene | |
| Recombinant DNA reagent (Homo sapiens) | pGEX4T-3 TEV DTX3L 544–606 | This paper | Codon-optimised synthetic gene | |
| Recombinant DNA reagent (Homo sapiens) | DTX1 388-C | Chatrin et al., 2020 | ||
| Recombinant DNA reagent (Homo sapiens) | DTX2 390-C | Chatrin et al., 2020 | ||
| Recombinant DNA reagent (Homo sapiens) | DTX3 148-C | Chatrin et al., 2020 | ||
| Recombinant DNA reagent (Homo sapiens) | DTX4 387-C | Chatrin et al., 2020 | ||
| Recombinant DNA reagent (Homo sapiens) | DTX3L 544-C | Chatrin et al., 2020 | ||
| Recombinant DNA reagent (Homo sapiens) | UBA1 | Nakasone et al., 2022 | ||
| Recombinant DNA reagent (Homo sapiens) | UBE2D2 | Dou et al., 2012 | ||
| Recombinant DNA reagent (Homo sapiens) | Ub | Gabrielsen et al., 2017; Volk et al., 2005 | ||
| Recombinant DNA reagent (Homo sapiens) | Fluorescent Ub | Magnussen et al., 2020 | ||
| Recombinant DNA reagent (Homo sapiens) | DTX3L(232-C)/PARP9(509-C) | This paper | Uniprot PARP9: Q8IXQ6-1 | Codon-optimised synthetic gene |
| Recombinant DNA reagent (Homo sapiens) | pRSFDuet 12 X His TEV PARG 448-C | Tucker et al., 2012 | Codon-optimised synthetic gene | |
| Recombinant DNA reagent (Homo sapiens) | USP2 260-C | Chatrin et al., 2020 | ||
| Strain, strain background (Escherichia coli) | BL21(DE3) Gold | Agilent | Cat# 230132 | Chemically competent |
| Strain, strain background (Escherichia coli) | Rosetta 2(DE3)pLysS | Merck Millipore Novagen | Cat# 71403 | Chemically competent |
| Peptide, recombinant protein | Neutravidin Protein, DyLight 800 | Invitrogen | Cat# 22853 | WB (1:10000) |
| Sequence-based reagent | Assorted oligos | Integrated DNA Technologies | Refer to Table 1 for sequences | |
| Peptide, recombinant protein | Benzonase Nuclease | Merck | Cat# E1014 | |
| Chemical compound, drug | NH2OH | Merck Millipore | Cat# 467804 | |
| Chemical compound, drug | ADPr | Merck | Cat# A0752 | |
| Chemical compound, drug | Biotin -NAD+ | Biolog | Cat# N 012 | |
| Chemical compound, drug | F-NAD+ | Biolog | Cat# N 023 | |
| Software, algorithm | GraphPad Prism | RRID:SCR_002798 | https://www.graphpad.com/features | |
| Software, algorithm | Quantity One 1-D analysis software | Bio-Rad | RRID:SCR_014280 | https://www.bio-rad.com/en-uk/product/quantity-one-1-d-analysis-software?ID=1de9eb3a-1eb5-4edb-82d2-68b91bf360fb |






