Synaptic deregulation of cholinergic projection neurons causes olfactory dysfunction across five fly Parkinsonism models
Figures
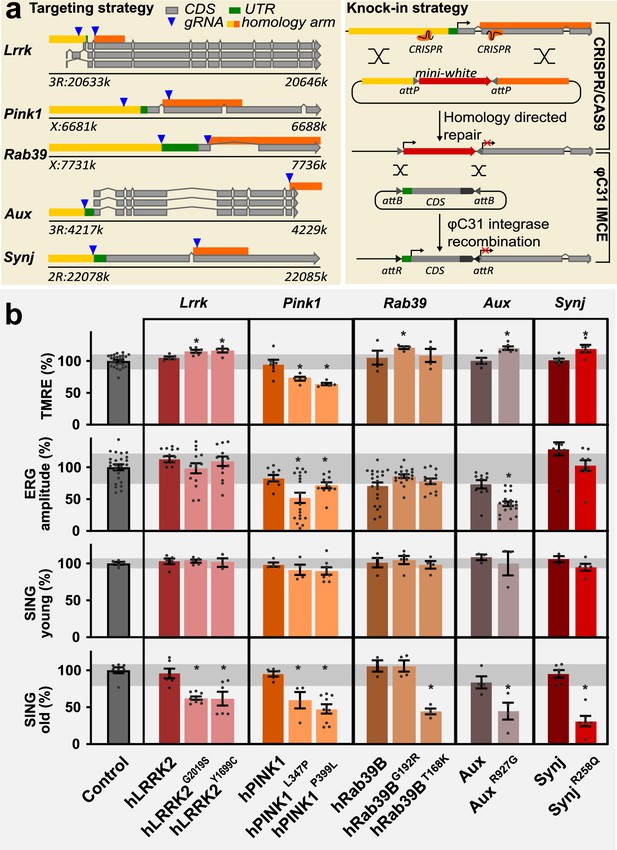
A collection of familial Parkinson’s disease (PD) knock-in models.
(a) Scheme of the knock-in strategy where the first common exon in all Drosophila transcripts (left) was replaced by an attP-flanked mini-white gene using CRISPR/Cas9 mediated homologous recombination, creating a null mutant, and then replaced by wild-type or pathogenic mutant human or fly coding DNA sequence (CDS) using PhiC31 mediated integration (right). The chromosomal positions of the genes are indicated. All knock-ins are in the same isogenic genetic background (cantonized w1118). (b) Phenotypic analysis of PD mutants: (1) Mitochondrial membrane potential measured by ratiometric tetramethyl rhodamine ethyl ester (TMRE) fluorescence at neuromuscular junction (NMJ) boutons in third-instar larvae. n≥4 animals per genotype and 10 boutons from ≥3 NMJs per animal. (2) Depolarization amplitude quantified from electroretinograms (ERGs), recorded after 7 days of light exposure. n≥20 animals per genotype. (3) Quantification of startle-induced negative geotaxis (SING) at 5±1 days after eclosion (young) and 25±1 days after eclosion (old). 95% of pink1 mutants died <25 days and were tested at 15 days. Values are normalized to control (see Materials and methods). Variance of control measurements are in gray. Bars are mean ± s.e.m.; *, p<0.05 ANOVA/Dunnett’s test.
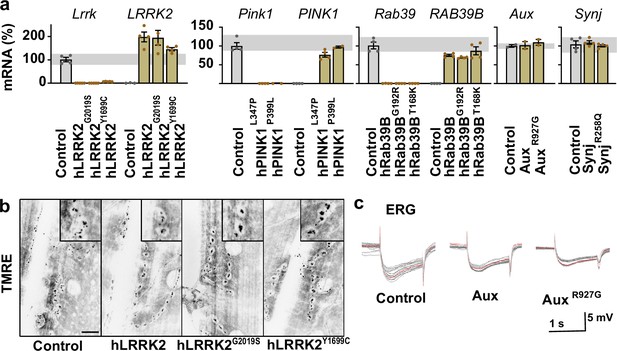
Characterization of Parkinson’s disease (PD) knock-in models.
(a) RNA expression levels, measured by RT-PCR with primers to the indicated coding DNA sequence (CDS) (above the graphs) in heads of controls and animals with wild-type or pathogenic mutant CDS knocked-in. Data are expressed as percent of endogenous Drosophila gene expression. Gray bars indicate the variance in controls. Bars are mean ± s.e.m. (b) Example images of mitochondrial tetramethyl rhodamine ethyl ester (TMRE) fluorescence at larval neuromuscular junctions (NMJs). Insets: magnified synaptic boutons (see also Figure 1). Scale bar: 10 μm. (c) Example electroretinogram (ERG) recordings (1 s light stimulus); see also Figure 1. Animals were kept for 7 days in constant light prior to the experiment. Red line: the average of the individual raw data traces that are shown.
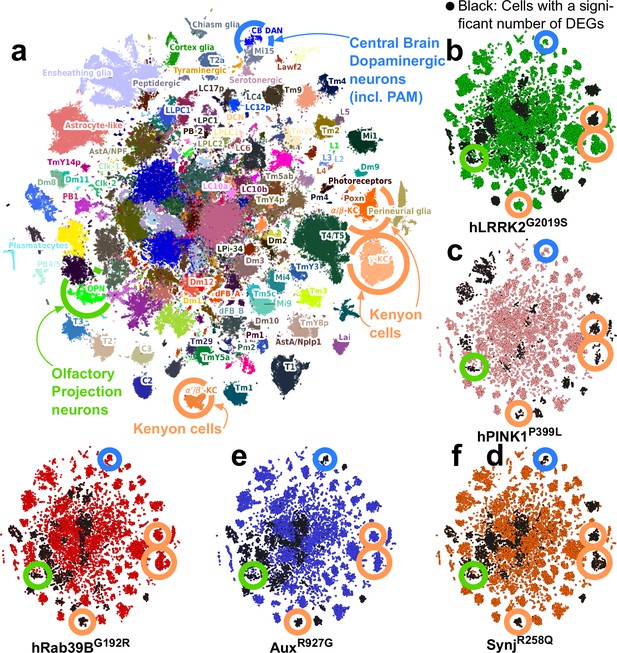
Single-cell RNA sequencing reveals that olfactory projection neurons (OPNs) are impaired across young Parkinson’s disease (PD) fly models.
(a) tSNE of 226k cells characterized by single-cell RNA sequencing of entire young (5-day-old) fly brains (controls and PD knock-in mutants). Colors indicate the cell types; 186 of which were identified (see Materials and methods). Key cell types are encircled, including dopaminergic neurons (DAN, blue), mushroom body Kenyon cells (orange), and OPN (green). (b–f) tSNE of the cells from five selective PD knock-in mutants (5 days of age). Black cells are those with a significant transcriptomic change. Key cell types labeled in (a) are indicated. Note that OPN are consistently affected across mutants, while DAN are not at this early stage.
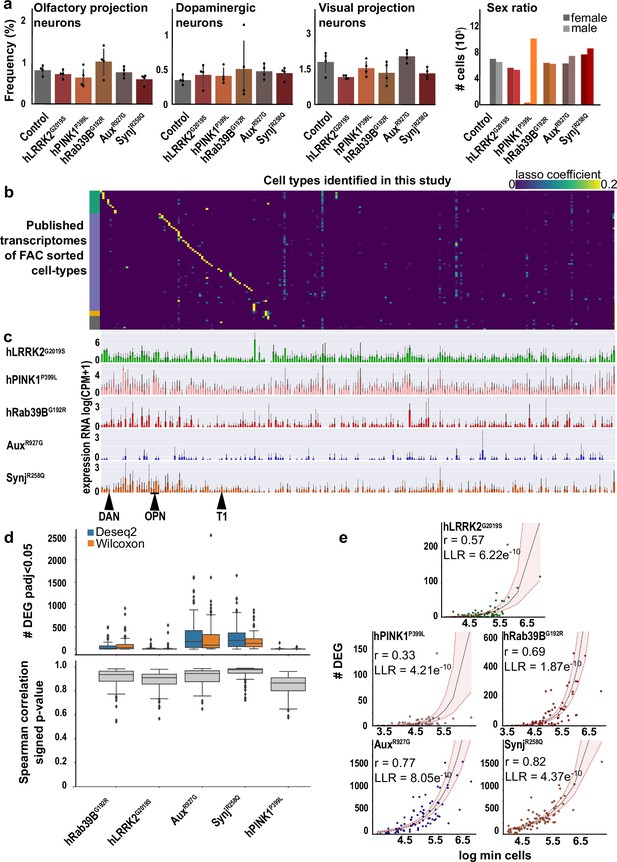
Validation and analysis of single-cell RNA sequencing data.
(a) Frequency of individual cell types (olfactory projection neurons [OPNs], dopaminergic neurons [DAN], visual projection neurons [T1]), and the sex ratios of the sequenced cells detected across the Parkinson’s disease (PD) knock-in models and controls. Note, we only sequenced males from hPINK1P399L mutants. Error bars represent mean±95% CI. (b) Alignment of published transcriptomic profiles defined by bulk sequencing of FAC sorted cell types (y-axis, see also Davie et al., 2018) and the cell types identified in this study (x-axis), validating the identification of cell types in the tSNE clusters. Green indicates central brain cell types; blue is optic lobe; yellow: glia; gray: others. (c) Expression level of knocked-in coding DNA sequence (CDS) across the cell types, indicating the knocked-in genes are broadly expressed. DAN - dopaminergic neurons (PAM cluster), OPNs - olfactory projection neurons, T1 - T1 visual interneurons. (d) Comparison of two different algorithms to detect cell-type-specific differentially expressed gene (DEG) (see Materials and methods) showing a similar number of DEG and high correlation of signed p-values. (e) Graphs showing the number of DEG versus the number of cells in a cluster (each dot is a cell type) for each PD fly model and the model of cell number dependency of p-values (dotted line, see Materials and methods). Confidence intervals in red. Cell types with residuals above confidence intervals are considered affected.
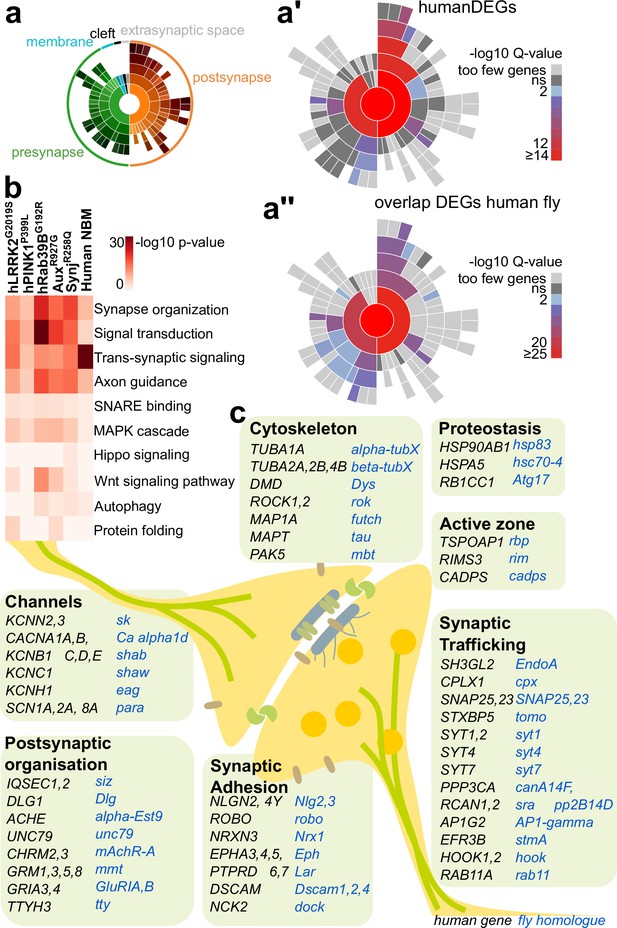
Common differentially expressed genes (DEGs) in cholinergic neuron-rich brain regions of human Parkinson’s disease (PD) patients and PD fly models.
(a–a”) Schematic of the sunburst plot indicating Gene Ontology (GO) terms for each sector (a) and the mapping of the DEGs in nucleus basalis of Meynert (NBM), nucleus accumbens, and putamen brain samples idiopathic PD patients (with LRRK2 risk mutations) and controls (a’) and mapping of the DEGs found commonly in fly and human samples (a”). Inner rings represent the different GO categories (indicated in a), with their subcategories in the outer rings, rings (in a’–a”) are color-coded according to enrichment Q-value. (b) GO analysis of DEG in cholinergic neurons of young PD fly models and NBM neurons of PD patients. Redundant terms were removed. Color: adjusted p-value. (c) Schematic of the DEGs found commonly in fly PD models (blue) and human PD samples (black) manually sorted according to their previously described synaptic functions. Supplementary file 7 contains the summarized results of the DEG analysis of fly brains and postmortem human brain samples and the SynGO analysis.
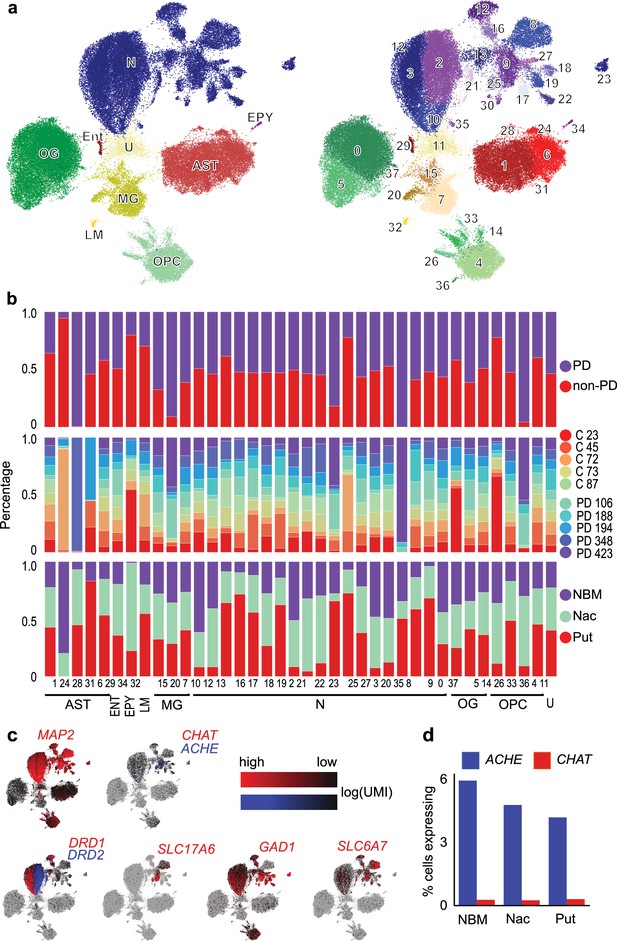
Single-cell RNA sequencing of cholinergic regions in postmortem human brain samples.
(a) tSNE of cells characterized in three cholinergic brain regions of 10 brains, annotated are the major cell types (left) and 36 individually identified cell types (numbered, right). N - neurons, OG - oligodendrocytes, ENT - endothelial cells, EPY - ependymal cells, LM - lymphocyte, MG - microglia, AST - astrocytes, OPC - oligodendrocyte precursor cells, U - unknown. (b) Frequency of the 36 cell types in pooled Parkinson’s disease (PD) versus non-PD samples, in individual brain samples, and in the three brain regions. (c) tSNE, as shown in (a), with different neuronal marker gene expression indicated (counts of unique molecular identifier [UMI]). (d) Frequency of cells expressing cholinergic marker genes ACHE and CHAT. NBM - nucleus basalis Meynert, Nac - nucleus accumbens, Put - putamen.
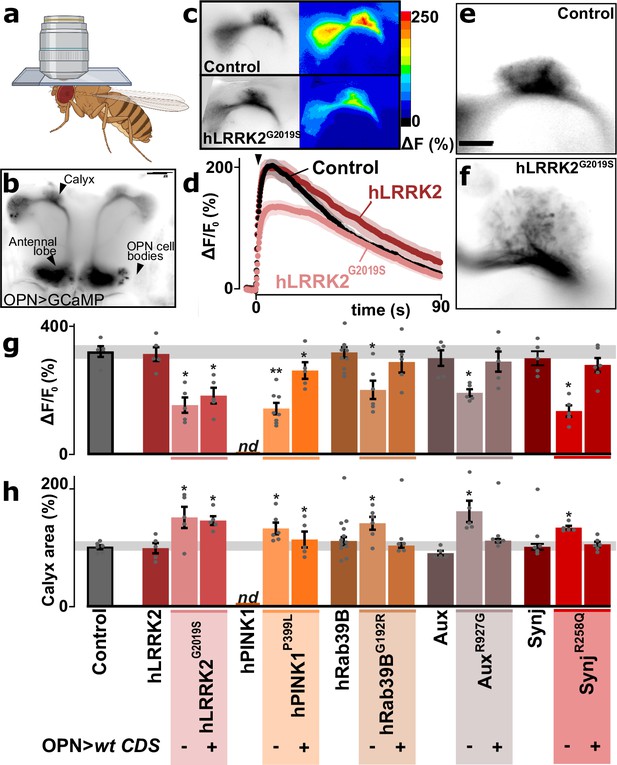
Synaptic defects in olfactory projection neuron (OPN) of Parkinson’s disease (PD) mutants.
(a) Schematic of a head-fixed awake fly for live Ca2+-imaging through a window in the head capsule. (b) Confocal image of GCaMP3-fluorescence expressed in OPN using GH146-Gal4, indicating the locations of the cell bodies, antennal lobe, and the calyx (scale bar: 50 µm). (c–d) Confocal image of the stimulus-induced change of fluorescence (peak amplitude) in the synaptic region of the calyx of a control and an hLRRK2G2019S knock-in animal (c) and quantification of fluorescence change (± SEM) over time (d). Arrowhead: time of stimulus application (10 mM nicotine, see Materials and methods). (e, f) Images of GFP fluorescence marking the synaptic area of OPN in the calyx of control (e) and hLRRK2G2019S knock-ins (f). Scale bar is 20 µm. (g) Quantification of GCaMP3 peak amplitude at OPN synapses in the calyx following stimulation (10 mM nicotine) in controls, wild-type coding DNA sequence (CDS) knock-ins, and in the PD knock-in mutants where the wild-type CDS is not (-) or is (+) expressed in OPN using GH146-Gal4 (OPN>wt CDS). Note that the hPINK1 control could not be determined as the combination of nSyb-Gal4>UAS-hPINK1 (expression in all neurons) in the Pink1 knock-out background interferes with the OPN-specific expression of Gal4 to drive UAS-GCaMP3 expression (g) (left bar; ‘nd’). In contrast, this issue is not present in hPINK1P399L mutant knock-in flies (nor the other flies used in the study) that could be rescued by OPN-specific expression of hPINK1 (g) (right bar); (‘OPN>wt CDS +’). (h) Quantification of the GFP fluorescence area of OPN synapses in the calyx (based on GCaMP3 signal) in controls, wild-type CDS knock-ins, and in the PD knock-in mutants where the wild-type CDS is not (-) or is (+) expressed in OPN (OPN>wt CDS). For (g, h) n≥5 animals per genotype. Bars are mean ± s.e.m. *, p<0.05 in ANOVA/Dunnett.
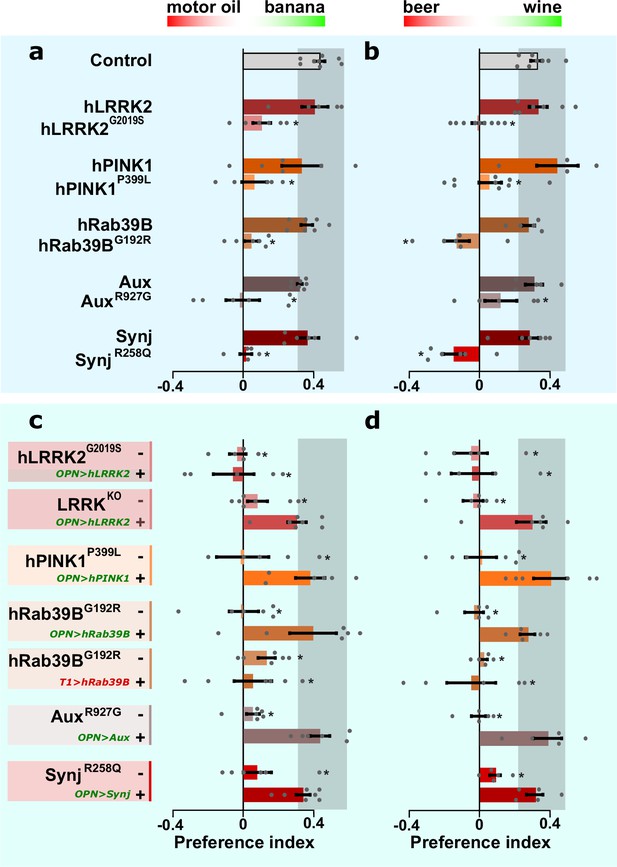
Olfactory projection neurons (OPN) dysfunction causes hyposmia in Parkinson’s disease (PD) mutants.
(a, b) Olfactory performance of PD knock-in flies and controls when given the choice between a blend of motor oil and banana odors (a) or a blend of beer and wine odors (b). (c, d) Olfactory performance of PD knock-in flies and LRRKKO when the relevant wild-type coding DNA sequence (CDS) is not (-) or is expressed selectively in OPN (‘OPN>wt CDS +’, green label) or T1 cholinergic interneurons (‘T1>hRab39B +’, red label), as a negative control. For (a–d) bars are mean ± s.e.m. n≥5 assays, ≥200 flies per genotype. *, p<0.05 in ANOVA/Dunnett.
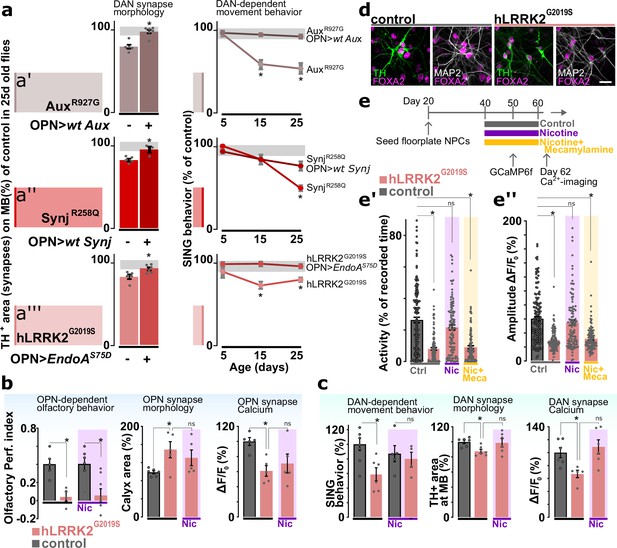
Cholinergic neuron activity rescues dopaminergic defects that occur at later stages in the life of Parkinson’s disease (PD) mutants.
(a, left) Synaptic area of dopaminergic neuron (DAN) innervating the mushroom body in aged PD models (AuxR927G, synjR258Q, or LRRK2G2019S) and with or without GH146-Gal4-driven expression of the wild-type PD gene (aux or synj) or EndoAS75D, respectively. Bars: mean ± SEM. n≥5, *p<0.05 in ANOVA, Dunnett’s test. (a, right) Startle-induced negative geotaxis (SING) of the PD models with or without GH146-Gal4-driven expression of wild-type gene or endoAS75D. Points: mean ± SEM. n≥5, *p<0.05 in two-way ANOVA. Gray zone: variance of controls. (b) Odor choice performance, stimulus-induced changes in synaptic Ca2+ signal and olfactory projection neuron (OPN) synapse area of young controls and hLRRK2G2019S flies with or without chronic nicotine (Nic) feeding (up to 1 day before testing). Bars: mean ± SEM. n≥5 assays, *p<0.05 in ANOVA, Dunnett’s test. (c) SING, stimulus-induced changes in synaptic Ca2+ and DAN synapse area of aged controls and hLRRKG2019S flies with or without chronic application of nicotine. Bars: mean ± SEM. n≥5 assays, *p<0.05 in ANOVA, Dunnett’s test. (d) Confocal images of differentiated (60 days) wild-type and LRRK2G2019S ventral midbrain DAN labeled with the ventral midbrain marker FOXA2, dopaminergic marker TH, and neuronal marker MAP2. Scale bar: 20 µm. (e) Scheme of the treatment protocol and spontaneous Ca2+ activity (e’) and amplitude (e’’) of human induced DAN, 2 days after 20 days of no treatment (Ctrl), nicotine (Nic) treatment or nicotine+mecamylamine (Nic+Meca) treatment. Bars: mean ± SEM. n≥60 DAN from three independent differentiations, *p<0.05 in ANOVA, Dunnett’s test.
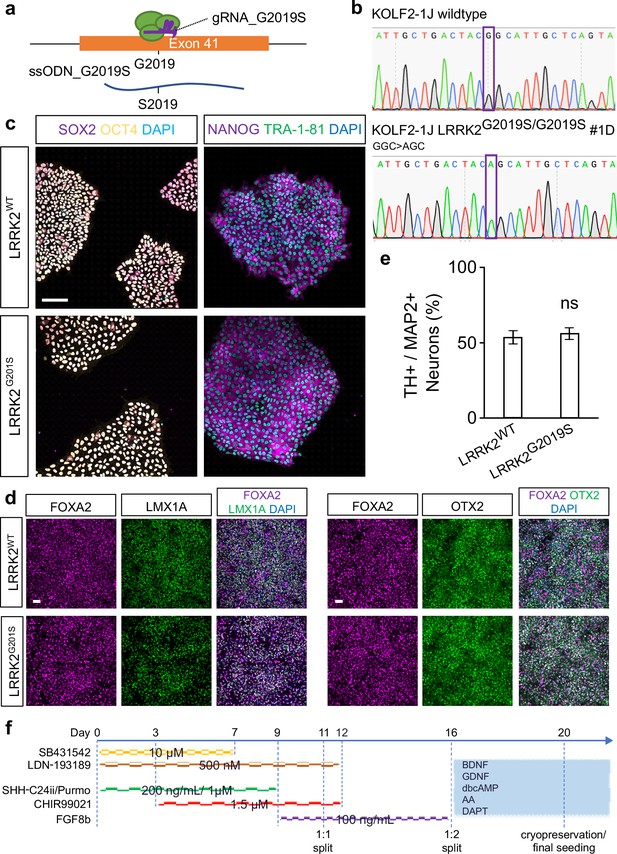
Generation and validation of human LRRK2G2019S ventral midbrain dopaminergic neuron (DAN).
(a) Schematic representation of gene editing strategy to knock-in the G2019S mutation in the LRRK2 locus. (b) Sanger sequencing of a single gene edited clone showing successful homozygous editing of the indicated nucleotide. (c) KOLF2-1J wild-type (LRRK2WT) control iPSCs and KOLF2-1J LRRK2G2019S/G2019S (LRRK2G2019S) iPSCs show normal expression of pluripotency markers OCT4 (yellow), SOX2 (purple), NANOG (purple), and TRA-1–81 (green). Nuclei are counterstained with DAPI (blue). Scale bar: 100 μm. (d) KOLF2-1J wild-type (LRRK2WT) control and KOLF2-1J LRRK2G2019S/G2019S (LRRK2G2019S) mutant ventral midbrain neural progenitor cells show normal expression of FOXA2 (magenta), LMX1A (green), and OTX2 (green), confirming that the neural progenitor cells are ventral midbrain-specific and capable of differentiating into DAN. Nuclei are counterstained with DAPI (blue). Scale bar: 100 μm. (e) Quantification of the frequency of TH+/MAP2+DAN. Statistical significance calculated with an ordinary t-test: ns, not significant. Error bars represent mean ± SEM. Data were collected in three independent vmDAN differentiations. (f) Differentiation protocol used.

Upregulation of protein homeostasis rescues hyposmia across familial models of PD.
Results of a behavioral screen for cell-specific rescue of olfactory preference defects of young PD fly models using up and downregulation of deregulated genes in affected cell types. Genes implicated in the indicated pathways are over expressed or knocked down using GH146-Gal4 (OPN>) and UAS-constructs (over expression or RNAi) . UAS-only (-) and OPN>UAS (+) were scored in parallel and are compared to each other. n.d. not determined; Bars represent mean ± s.e.m.; gray zone indicates the variance of controls; n≥5 independent experiments per genotype, with ~50 flies each; red bars: p<0.05 in ANOVA and Bonferroni-corrected comparison to UAS-only control.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (D. melanogaster) | Lrrk | FlyBase | FBgn0038816 | |
| Gene (D. melanogaster) | Pink1 | FlyBase | FBgn0029891 | |
| Gene (D. melanogaster) | Rab39 | FlyBase | FBgn0029959 | |
| Gene (D. melanogaster) | Aux | FlyBase | FBgn0037218 | |
| Gene (D. melanogaster) | Synj | FlyBase | FBgn0034691 | |
| Strain, strain background (Escherichia coli) | One Shot top10 cells | Thermo Fisher Scientific | C404010 | Competent cells |
| Genetic reagent (D. melanogaster) | GH146-Gal4 (OPN-Gal4) | Stocker et al., 1997; Bloomington Drosophila Stock Center | BDSC_30026; FlyBase: FBti0016783 | |
| Genetic reagent (D. melanogaster) | GMR31F10-Gal4 (T1-Gal4) | Jenett et al., 2012; Bloomington Drosophila Stock Center | BDSC_49685; FlyBase: FBti0134817 | |
| Genetic reagent (D. melanogaster) | GMR58E02-Gal4 (PAM-Gal4) | Jenett et al., 2012; Bloomington Drosophila Stock Center | BDSC_41347; FlyBase: FBti0137105 | |
| Genetic reagent (D. melanogaster) | UAS-GCaMP3 | Tian et al., 2009; Bloomington Drosophila Stock Center | BDSC_32236; BDSC_32116 | |
| Genetic reagent (D. melanogaster) | UAS-hLRRK2 | Venderova et al., 2009 | ||
| Genetic reagent (D. melanogaster) | UAS-hPINK1 | Yang et al., 2006 | ||
| Genetic reagent (D. melanogaster) | UAS-YFP-Rab39 | Zhang et al., 2007; Bloomington Drosophila Stock Center | BDSC_9825; FlyBase: FBti0100878 | |
| Genetic reagent (D. melanogaster) | UAS-Synj | Vanhauwaert et al., 2017 | ||
| Genetic reagent (D. melanogaster) | UAS-Aux | Jacquemyn et al., 2023 | ||
| Genetic reagent (D. melanogaster) | LRRKk.o. | This study (see Figure 1a and the Materials and methods section) | Knock-out fly line carrying an attP-flanked mini-white gene replacing the first common exon in all transcripts of lrrk | |
| Genetic reagent (D. melanogaster) | hLRRK2 | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the wild-type human LRRK2 CDS at the lrrk locus | |
| Genetic reagent (D. melanogaster) | hLRRK2G2019S | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic human LRRK2G2019S CDS at the lrrk locus | |
| Genetic reagent (D. melanogaster) | hLRRK2Y1699C | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic human LRRK2Y1699C CDS at the lrrk locus | |
| Genetic reagent (D. melanogaster) | PINK1k.o. | This study (see Figure 1a and the Materials and methods section) | Knock-out fly line carrying an attP-flanked mini-white gene replacing the first common exon in all transcripts of pink1 | |
| Genetic reagent (D. melanogaster) | hPINK1P399L | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic human PINK1P399L CDS at the pink1 locus | |
| Genetic reagent (D. melanogaster) | hPINK1L347P | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic human PINK1L347P CDS at the pink1 locus | |
| Genetic reagent (D. melanogaster) | Rab39k.o. | This study (see Figure 1a and the Materials and methods section) | Knock-out fly line carrying an attP-flanked mini-white gene replacing the first common exon in all transcripts of rab39 | |
| Genetic reagent (D. melanogaster) | hRab39B | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the wild-type human Rab39B CDS at the rab39 locus | |
| Genetic reagent (D. melanogaster) | hRab39BG192R | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic human Rab39BG192R CDS at the rab39 locus | |
| Genetic reagent (D. melanogaster) | hRab39BT168K | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic human Rab39BT168K CDS at the rab39 locus | |
| Genetic reagent (D. melanogaster) | Auxk.o. | This study (see Figure 1a and the Materials and methods section) | Knock-out fly line carrying an attP-flanked mini-white gene replacing the first common exon in all transcripts of aux | |
| Genetic reagent (D. melanogaster) | Aux | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the wild-type Drosophila Aux CDS at the aux locus | |
| Genetic reagent (D. melanogaster) | AuxR927G | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic Drosophila AuxR927G CDS at the aux locus | |
| Genetic reagent (D. melanogaster) | Synjk.o. | This study (see Figure 1a and the Materials and methods section) | Knock-out fly line carrying an attP-flanked mini-white gene replacing the first common exon in all transcripts of synj | |
| Genetic reagent (D. melanogaster) | Synj | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the wild-type Drosophila Synj CDS at the synj locus | |
| Genetic reagent (D. melanogaster) | SynjR258Q | This study (see Figure 1a and the Materials and methods section) | Knock-in fly line carrying the pathogenic Drosophila SynjR258Q CDS at the synj locus | |
| Cell line (H. sapiens) | KOLF2-1J wild-type (LRRK2WT) control iPSCs | Pantazis et al., 2022; from the Jackson Laboratory under the iPSC Neurodegenerative Disease Initiative | Product code: JIPSC001000 | |
| Cell line (H. sapiens) | KOLF2-1J LRRK2G2019S/G2019S (LRRK2G2019S) iPSCs | This study (see Figure 6—figure supplement 1 and the Materials and methods section) | Knock-in KOLF2-1J iPSC line carrying the pathogenic p.G2019S mutation in the LRRK2 locus | |
| Transfected construct (H. sapiens) | pLenti-hSyn1-mRuby2-T2A-GCaMP6f | This study (see the Materials and methods section) | RRID:Addgene_197595 | Lentiviral transfer vector for the neuronal expression of mRuby2-T2A-GCaMP6f (fluorescent reporter for calcium imaging) |
| Biological sample (H. sapiens) | Postmortem brain tissue | Parkinson's UK Brain Bank at the Imperial College London | EC reference NH019 2019-02-01 | |
| Antibody | Mouse monoclonal IgG1 anti-SOX2 | Santa Cruz | sc-365823 | IF (1:200) |
| Antibody | Rabbit polyclonal anti-OCT4 | Abcam | ab19857 | IF (1:50) |
| Antibody | Mouse monoclonal IgG1 anti-NANOG | Santa Cruz | sc-293121 | IF (1:50) |
| Antibody | Mouse monoclonal IgM anti-TRA-1–81 | Sigma | MAB4381 | IF (1:100) |
| Antibody | Rabbit polyclonal anti-LMX1A/B | Millipore | AB10533 | IF (1:1000) |
| Antibody | Mouse monoclonal IgG2a anti-FOXA2/HNF-3β | Santa Cruz | sc-101060 | IF (1:250) |
| Antibody | Mouse monoclonal IgG1k anti-OTX2 | Santa Cruz | sc-514195 | IF (1:100) |
| Antibody | Rabbit polyclonal anti-TH | Sigma | AB152 | IF (1:500) |
| Antibody | Mouse monoclonal IgG1 anti-MAP2 | Sigma | M1406 | IF (1:1000) |
| Antibody | Goat polyclonal Alexa Fluor-488 conjugated secondary antibody | Invitrogen | Anti-rabbit: # A-11034 for OCT4/TH Anti-mouse IgG1: # A-21121 for Nanog/OTX2 | IF (1:500) |
| Antibody | Goat polyclonal Alexa Fluor-555 conjugated secondary antibody | Invitrogen | Anti-mouse IgG1: # A-21127 for SOX2 Anti-mouse IgG2a: # A-21137 for FOXA2 | IF (1:500) |
| Antibody | Donkey/Goat polyclonal Alexa Fluor-647 conjugated secondary antibody | Invitrogen | Anti-rabbit: # A-31573 for LMX1A Anti-mouse IgM: # A-21238 for TRA-1–81 Anti-mouse IgG1: # A-21240 for MAP2 | IF (1:500) |
| Recombinant DNA reagent | pCFD4: U6:1-gRNA U6:3-gRNA | Port et al., 2014 | RRID:Addgene_49411 | |
| Recombinant DNA reagent | pUC19 | Norrander et al., 1983 | RRID:Addgene_50005 | |
| Recombinant DNA reagent | pUAST.attB | Bischof et al., 2007 | GenBank:EF362409.1 | |
| Recombinant DNA reagent | pFL44S{w+}-attB | Khuong et al., 2013 | ||
| Recombinant DNA reagent | pWhite-STAR | Choi et al., 2009 | ||
| Recombinant DNA reagent | 2XMyc-LRRK2-WT | Greggio et al., 2008 | RRID:Addgene_25361 | |
| Recombinant DNA reagent | pLenti6-DEST PINK1-V5 WT | Beilina et al., 2005 | RRID:Addgene_13320 | |
| Sequence-based reagent | Primers, gRNAs, oligos, gBlocks, ssODN | Integrated DNA Technologies (IDT) | See Supplementary file 1 | |
| Sequence-based reagent | iPSC gRNA | Synthego | See Supplementary file 1 | |
| Peptide, recombinant protein | Cas9 HiFi protein | Integrated DNA Technologies (IDT) | Alt-R S.p. HiFi Cas9 Nuclease V3, 500 µg; #1081061 | |
| Peptide, recombinant protein | Dispase | Sigma-Aldrich | D4818 | |
| Peptide, recombinant protein | Collagenase | Invitrogen, CA, USA | 17100-017 | |
| Chemical compound, drug | Ringer's solution | Estes et al., 1996 | ||
| Chemical compound, drug | TMRE | Abcam, Cambridge, UK | ab113852 | |
| Chemical compound, drug | Trypsin-EDTA | Invitrogen, CA, USA | 25300054 | |
| Chemical compound, drug | Nicotine | Sigma-Aldrich | N3876 | |
| Commercial assay, kit | NEBuilder HiFi DNA Assembly | New England Biolabs, Inc (NEB) | NEB #E2621 | |
| Commercial assay, kit | Maxwell RSC instrument and kit | Promega | AS8500, AS1340 | |
| Commercial assay, kit | Chromium Single Cell 3′ Library & Gel Bead Kit v2 | 10x Genomics | PN-120237 | |
| Commercial assay, kit | Chromium Single Cell A Chip Kit | 10x Genomics | PN-1000009 | |
| Commercial assay, kit | Chromium i7 Multiplex Kit | 10x Genomics | PN-120262 | |
| Commercial assay, kit | Chromium Next GEM Single Cell 3' Kit v3 | 10x Genomics | PN-1000075 | |
| Commercial assay, kit | Chromium Next GEM Chip B Single Cell Kit | 10x Genomics | PN-1000073 | |
| Commercial assay, kit | Single Index Kit T Set A | 10x Genomics | PN-1000213 | |
| Software, algorithm | Fiji | Schindelin et al., 2012 | RRID:SCR_002285 | https://imagej.net/Fiji |
| Software, algorithm | Cell Ranger (v 3.0.2) | 10x Genomics | RRID:SCR_017344 | https://www.10xgenomics.com/support/software/cell-ranger/latest |
| Software, algorithm | GOrilla | Eden et al., 2009 | RRID:SCR_006848 | http://cbl-gorilla.cs.technion.ac.il/ |
| Software, algorithm | GraphPad Prism | GraphPad software, CA, USA | RRID:SCR_002798 | https://www.graphpad.com/scientific-software/prism/ |
| Software, algorithm | Vib-singlecell-nf/vsn-pipelines (v 0.27.0) | Flerin et al., 2021 | https://github.com/vib-singlecell-nf/vsn-pipelines | |
| Software, algorithm | Python (v 3.7.3) | Python | RRID:SCR_008394 | http://www.python.org/ |
| Software, algorithm | pandas (v 0.25.1) | GitHub | RRID:SCR_018214 | https://github.com/pandas-dev/pandas |
| Software, algorithm | numpy (v 1.19.1) | Harris et al., 2020 | RRID:SCR_008633 | http://www.numpy.org/ |
| Software, algorithm | matplotlib (v 3.3.0) | GitHub | RRID:SCR_008624 | https://github.com/matplotlib/matplotlib |
| Software, algorithm | seaborn (v 0.9.0) | GitHub | RRID:SCR_018132 | https://github.com/mwaskom/seaborn |
| Software, algorithm | statsmodels (v 0.10.1) | Seabold and Perktold, 2010 | RRID:SCR_016074 | http://www.statsmodels.org/ |
| Software, algorithm | scipy (v 1.4.1) | Virtanen et al., 2020 | RRID:SCR_008058 | http://www.scipy.org/ |
| Software, algorithm | adjustText (v 0.7.3) | GitHub | RRID:SCR_022260 | https://github.com/Phlya/adjustText |
| Software, algorithm | loompy (v 3.0.6) | GitHub | RRID:SCR_016666 | https://github.com/linnarsson-lab/loompy |
| Software, algorithm | sklearn (v 0.20.1) | Pedregosa et al., 2011 | RRID:SCR_019053 | https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.NMF.html |
| Software, algorithm | patsy (v 0.5.1) | Smith, 2018 | https://github.com/pydata/patsy | |
| Software, algorithm | venn (v 0.1.3) | Python | https://pypi.org/project/venn/ | |
| Software, algorithm | R (v 3.6.1) | R Foundation | RRID:SCR_001905 | http://www.r-project.org/ |
| Software, algorithm | MAST (v 1.12.0) | GitHub | RRID:SCR_016340 | https://github.com/RGLab/MAST/ |
| Software, algorithm | SCopeLoomR (v 0.5.1) | De Waegeneer et al., 2022 | https://github.com/aertslab/SCopeLoomR | |
| Other | SCope Resource Website | Davie et al., 2018 | Visualization tool for large-scale sc-RNA-seq datasets; http://scope.aertslab.org |
Additional files
-
Supplementary file 1
List of gRNAs, gBlocks, and oligos used to generate and validate the fly lines and hiPSC line generated in this study.
List of full fly genotypes and hiPSC lines used in this study.
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp1-v1.xlsx
-
Supplementary file 2
Full results of the Cell Ranger summaries (10x Genomics).
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp2-v1.xlsx
-
Supplementary file 3
Summary of the number of cells per cell cluster for each mutant and controls, including the raw number of cells, the percentage of cells for each cluster per mutant, and the percentage of cells for each mutant and controls per cluster.
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp3-v1.xlsx
-
Supplementary file 4
Summary of the parameters used to model differentially expressed gene (DEG)-cell number correlation.
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp4-v1.xlsx
-
Supplementary file 5
List of cell types and respective z-residuals (deviation from model) to determine affected cell types and those that are outside of the 95% confidence interval.
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp5-v1.xlsx
-
Supplementary file 6
List of the clusters with significant changes (based on cell number-adjusted differentially expressed genes [DEGs]) for each model, black cell clusters (cells with significant number of DEG) are listed as TRUE.
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp6-v1.xlsx
-
Supplementary file 7
Summarized results of the differentially expressed gene (DEG) analysis in fly model brains and postmortem human samples and the summary of the SynGO analysis.
- https://cdn.elifesciences.org/articles/98348/elife-98348-supp7-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/98348/elife-98348-mdarchecklist1-v1.docx





