Sequential replacement of PSD95 subunits in postsynaptic supercomplexes is slowest in the cortex
Figures
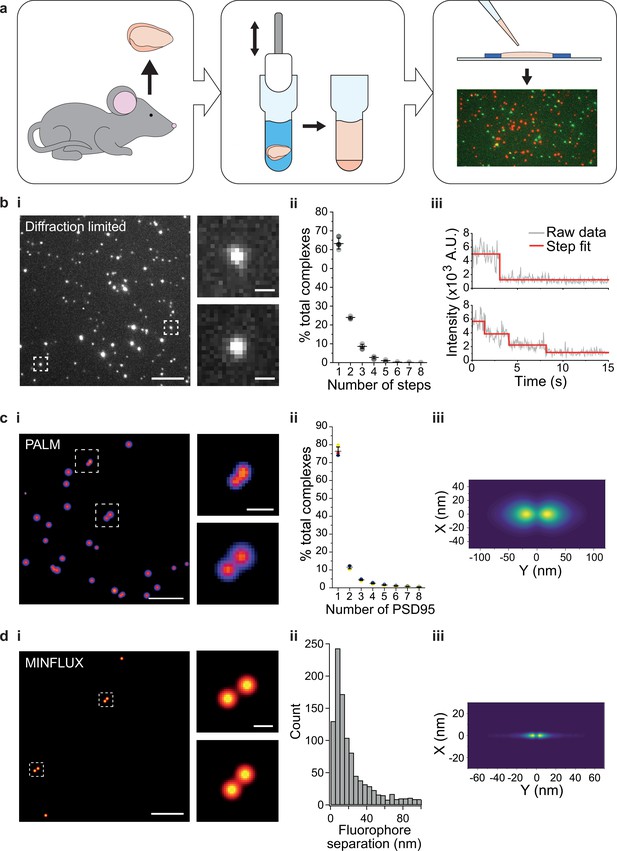
Stoichiometry and spatial arrangement of PSD95 within individual supercomplexes.
(a) Brain containing either endogenously tagged PSD95-eGFP or PSD95-mEoS2 was extracted from the genetically modified mouse, and the forebrain was homogenized to solubilize PSD95-containing supercomplexes. PSD95 supercomplexes were immobilized on glass coverslips and imaged using single-molecule and super-resolution approaches. (bi) Individual PSD95-eGFP supercomplexes (boxed) were imaged using total internal reflection fluorescence (TIRF) microscopy. Scale bar = 5 µm, inset scale bar = 500 nm. Photobleaching step counting revealed a distribution of PSD95 stoichiometries (bii) (10,178 PSD95-eGFP photobleaching steps were counted across 6461 supercomplexes). Representative intensity traces with fits shown in biii. (ci) Example photoactivated localization microscopy (PALM) images of individual PSD95 supercomplexes. Scale bar = 500 nm, inset scale bar = 50 nm. (cii) Subsequent analysis revealed that PSD95 exists at a range of stoichiometries within the supercomplexes (132,929 PSD95-mEoS2 molecules were detected in 82,501 individual supercomplexes). Plots show mean ± SD, n = 3 biological repeats. (ciii) Class averaging of the dimer population shows a distinct separation between the PSD95 proteins within the supercomplexes (class average of 9743 supercomplexes). (di) Example MINFLUX images of PSD95 supercomplexes. Scale bar = 100 nm, inset scale bar = 10 nm. (dii) Analysis of the supercomplexes containing two PSD95 molecules (1011 supercomplexes) showed a distribution of PSD95 separation distances. (diii) Class averaging of this population revealed two peaks separated by 12.7 nm.
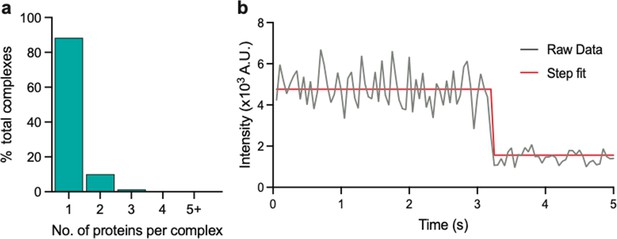
Stoichiometry of eGFP determined using photobleaching step count analysis.
(a) Stoichiometries of eGFP determined using the same approach as that for eGFP-tagged PSD95. (b) Example intensity trace with fitting.
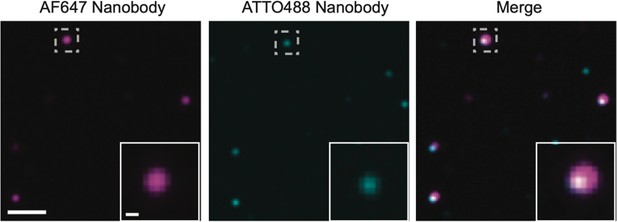
Multimers of PSD95 in wild-type supercomplexes.
Homogenate extracted from wild-type mice was processed using the same procedure as that for PSD95-eGFP mice. It was subsequently diluted 1:1000 and 100 µl of this solution was incubated on a plasma-cleaned coverslip for 30 min. The coverslip was washed three times with phosphate-buffered saline (PBS) and a mix of 2.5 pM AF647-tagged and 2.5 pM ATTO488-tagged FluoTag-X2 anti-PSD95 nanobody (NanoTag Biotechnologies) was added to the coverslip (total volume 100 µl) and incubated for 30 min. Following washing three times with PBS, the sample was imaged on the total internal reflection fluorescence (TIRF) microscope with 638 and 488 nm excitation. Coincident spots represent supercomplexes containing multiple PSD95 units. Scale bar = 2 µm, inset scale bar = 250 nm.
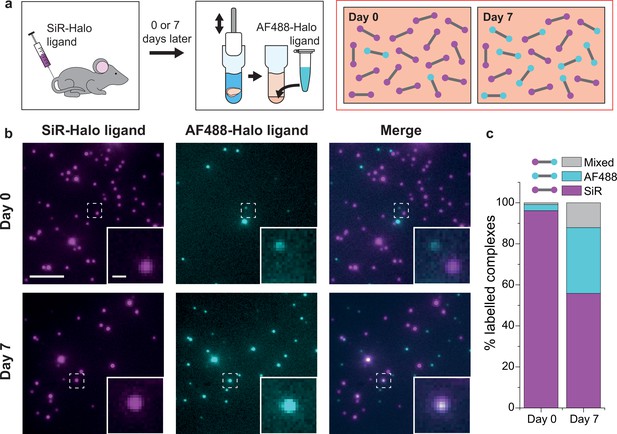
PSD95 turnover within supercomplexes in mouse brain homogenate.
(a) PSD95-HaloTag homozygous mice were injected with SiR-Halo ligand and culled 6 hr (day-0) or 7 days (day-7) post injection. The forebrains were homogenized and post hoc stained with AF488-Halo ligand to saturate remaining binding sites. At day-0, the vast majority of PSD95 is labeled with SiR-Halo only. After 7 days of protein turnover, three populations of supercomplex are possible: SiR-Halo only, AF488-Halo only, and both fluorophores. (b) Images of supercomplexes from homogenate at day-0 and day-7, showing increased AF488-Halo:SiR-Halo ratios at day-7, with increased coincidence. Scale bar is 5 μm and 500 nm in the zoom. (c) Quantified percentages of supercomplexes labeled only with SiR-Halo, AF488-Halo, or both. At day-0, 96% of supercomplexes were labeled with SiR-Halo only, indicating saturation of PSD95-HaloTag binding sites by injection. At day-7, this had decreased to 56%, with expansion of the AF488-Halo and co-labeled populations, indicating that PSD95 protein turnover had occurred over the 7 days.

Absolute number graphs for Figure 2c.
Number of labeled supercomplexes at day-0 and day-7 containing only SiR-labeled ‘old’ protein (SIR), AF488-labeled ‘new’ protein (AF488), or both ‘mixed’. In total, 13,710 supercomplexes were analyzed at day-0 and 15,391 supercomplexes were analyzed at day-7.
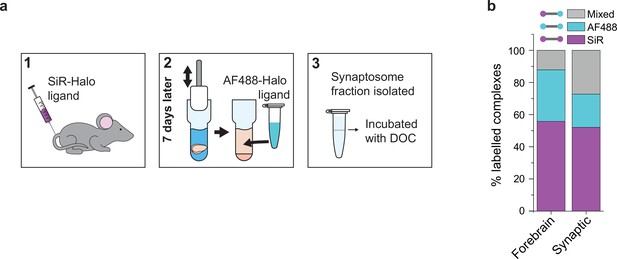
Protein turnover in synaptic and total forebrain PSD95 supercomplexes.
(a) SiR-Halo ligand was injected into the tail vein of three 3-month-old PSD95-HaloTag knock-in mice. Seven days later, the forebrain was extracted and synaptosomes isolated. AF488-HaloTag ligand was incorporated during the homogenization process. DOC, deoxycholate. (b) Comparison of populations of PSD95 supercomplexes in the total forebrain and synaptic fraction.
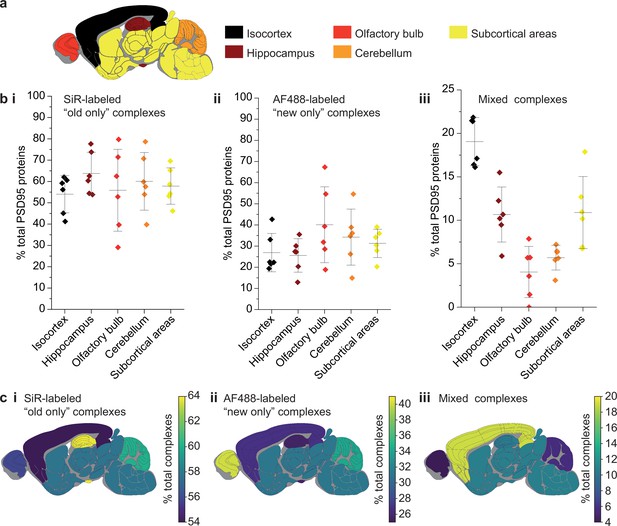
Differences in PSD95 turnover between mouse brain regions.
(a) Mouse brains were dissected into five broad regions. (b) The percentage of total PSD95 imaged contained in SiR-labeled (i), AF488-labeled (ii), and mixed (iii) supercomplexes. Error bars show the SD of 6 biological repeats. (c) The percentage of SiR-labeled (i), AF488-labeled (ii), and mixed (iii) PSD95 supercomplexes in each region of the brain. ANOVA tests for differences between the five brain region means was carried out for plots bi, bii, and biii. The ANOVA tests returned p-values of 0.71, 0.23, and 5.5 × 10−8, respectively. This indicates that there is no statistical significance in plots bi and bii, but there are statistical differences between some of the means in plot biii. Post hoc t-tests were carried out to identify the source of the significance. The p-values for comparisons between all regions are shown in Table 2.
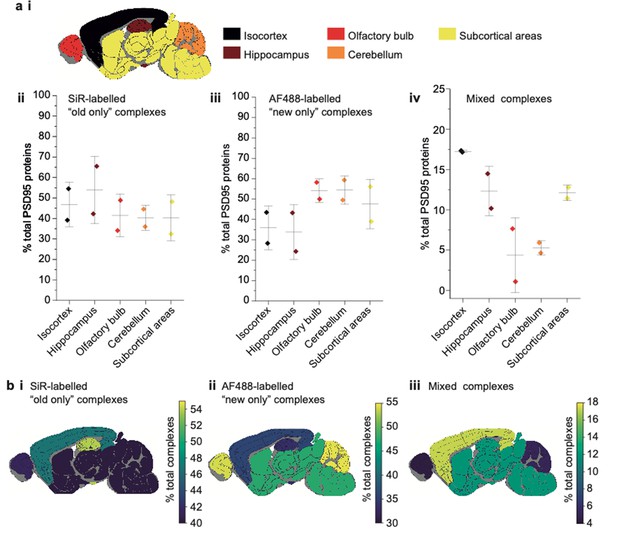
Characterization of PSD95 turnover in 3-week-old mice.
(a) Key showing the five dissected brain regions (i). Percentage of total PSD95 proteins in old-only (ii), new-only (iii), and mixed (iv) supercomplexes (mean ± SD, n = 3). (b) Mouse brain region heatmaps showing the same data as in a ii–iv.
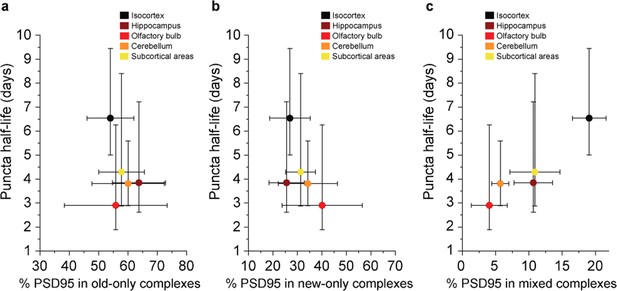
Comparison between the density-based PSD95 puncta half-life and old, new, and mixed proteins in supercomplexes.
The percentage of complexes that contain (a) old-only, (b) new-only, or (c) mixed old and new PSD95 (mean ± SD for % PSD95, and ± 95% CI for half-life, n = 3). There is no statistically significant correlation in a or b. However, R = 0.9536 and p = 0.01 from a Pearson’s correlation test, indicating a significant positive correlation between the percentage of PSD95 in mixed supercomplexes and PSD95 puncta half-life.
Tables
MINFLUX iterations.
| 1st | 2nd | 3rd | 4th | 5th | |
|---|---|---|---|---|---|
| L size [nm] | 300 | 300 | 150 | 75 | 40 |
| TCP – pattern | Hexagon | Hexagon | Hexagon | Hexagon | Hexagon |
| Minimum number of collected photons | 100 | 150 | 100 | 100 | 150 |
| Laser power factor | 1x | 1x | 2x | 4x | 6x |
| TCP dwell time [ms] | 1 | 1 | 1 | 1 | 1 |
| CFR | x | 0.5 | x | 0.8 | x |
| Background threshold [kHz] | 15 | 15 | 15 | 15 | 15 |
Post hoc t-tests to locate the source of significance between the means of the mixed supercomplexes in the five brain regions analyzed.
Only p-values less than 0.05 are shown. There are high levels of significance between the means of the isocortex and all other regions.
| Isocortex | Hippocampus | Olfactory bulb | Cerebellum | Subcortex | |
|---|---|---|---|---|---|
| Isocortex | 6.4 × 10−4 | 3.8 × 10−6 | 1.5 × 10−5 | 3.1 × 10−3 | |
| Hippocampus | 3.8 × 10−3 | 1.0 × 10−2 | N.S. | ||
| Olfactory bulb | N.S. | 9.3 × 10−3 | |||
| Cerebellum | N.S. | ||||
| Subcortex |





