By Emmy Tsang and Giuliano Maciocci
Since 2017, we have been working on the concept of computationally reproducible papers. The open-source suite of tools that started life as the Reproducible Document Stack is now live on eLife as ERA, the Executable Research Article, delivering a truly web-native format for taking published research to a new level of transparency, reproducibility and interactivity.
From today, authors with a published eLife paper can register their interest to enrich their published work with the addition of live code blocks, programmatically-generated interactive figures, and dynamically generated in-line values, using familiar tools like R Markdown and Jupyter in combination with Stencila Hub’s intuitive asset management and format conversion interface. The resulting new ERA publication will be presented as a complement to the original published paper. Very soon, a Google Docs plugin will also be made available to let authors insert executable code and data blocks into their documents using the cloud service.
Readers of ERA publications will be able to inspect the code, modify it, and re-execute it directly in the browser, enabling them to better understand how a figure is generated. They will be able to change a plot from one format to another, alter the data range of a specific analysis, and much more. All changes are limited to an individual’s browsing session and do not affect the published article, so anyone can experiment safely. Readers can also download the ERA publication – with all embedded code and data preserved – and use it as a basis for further study or derivative works.
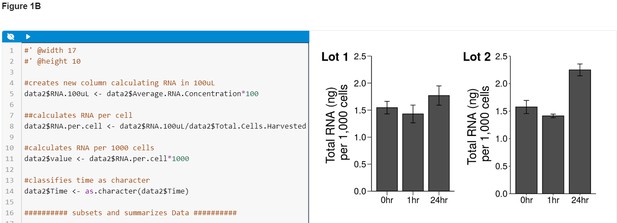
Figures in ERAs can be accompanied by editable code blocks, where the code can be edited and re-executed to immediately see the effects of those edits.
Started by eLife in partnership with Substance, and later developed by Stencila, ERA takes many of the important lessons learned from pioneering technologies like Jupyter and R Markdown, and folds them into a new open-source research publication format that treats code and data as first-class citizens alongside the traditional text and figures. The ERA technology stack is also designed specifically towards preprint and journal publishing, with strong XML standards, native schema.org compliance, and a versatile theming solution that makes it adaptable to the visual style of virtually any online publication.
Getting started with ERA
Explore some of the functionalities that ERA offers with two published examples:
- This ERA complement of Lewis et al., 2018 (original article) showcases executable figures, tables and in-line numbers.
- This ERA complement of “Inter- and intra-animal variation in the integrative properties of stellate cells in the medial entorhinal cortex” by Pastoll et al. (original article) demonstrates a variety of executable plots.
Start by clicking the Run Document button at the top of the article. This will start a compute session. Be patient, as it may take a moment. When you see the “Ready to run document” message, you’re good to go.
Then, look for the blue “eye” icons throughout the paper, which indicate code blocks. Clicking on those icons will reveal the underlying code, which can then be edited in place.
After making an edit, press the white “play” icon at the top of the code block to execute your new code and see its effects. Please note that it may take some time for the code to execute, depending on its complexity and on server load.
Publish an ERA with eLife
eLife authors can register their interest to enrich their published works on eLife by using the button below. At this initial stage of release, our aim is to work closely with the first few chosen ERA authors to further test and improve the ERA tools and workflow– your input will be crucial to helping us create a more seamless experience for more authors in the future.

Workflow for eLife authors to compose and publish an ERA complement.
Find out more about ERA
- If you are an eLife author and are interested in publishing an ERA complement to your published article, please get in touch.
- We hosted a webinar and demo on September 8– please view the recording.
- Please sign up for our approximately bi-monthly ERA community update.
Questions? Please check the Frequently Asked Questions section below, or email us at innovation [at] elifesciences [dot] org.
Frequently Asked Questions
Readers
Q: Can I share the changes I’ve made to the code?
A: No – at the moment the changes are not saved and will disappear once you close/refresh your browser. You can however download the ERA (containing the Markdown/notebook file with the associated code and data files), edit the files locally and share the results.
Q: I’d like to use the code that’s in this paper to analyse my own data – what’s the best way to do it?
A: You can download the ERA (see above) and replace the data file with your own (although you may need to change some of the variables/parameters). Please remember to cite the original paper if you do publish those results!
Q: Can I comment on the code?
A: Not at the moment, unfortunately – Hypothesis is yet to be made available on ERA articles.
Q: Can I download a particular code chunk/figure, or the underlying data file?
A: At the moment, you cannot download a particular code chunk/executable figure. You can download the entire ERA (which contains the underlying code, CSV data and HTML).
Authors
Q: What tools do I need to create an Executable Research Article?
A: The Stencila Hub interface allows for format conversion between various common tools such as RStudio, Jupyter, Google Docs, and even Word and PDF. Thanks to some embedding wizardry you can even convert to Word and PDF to share work with your colleagues for comment, and convert those back to R Markdown without losing any of your code and data!
Q: What kind of programming languages can ERA handle?
A: ERA can currently handle R Markdown and Python. Dependency compatibility for non-standard packages may vary, but if you’re experiencing issues or need new packages added for your article, you can contact Stencila to request this.
Q: How do I compose an executable companion to my paper?
A: Did you:
- publish with eLife,
- have figures that are generated using R/Python in that paper,
- have the raw data file for those figures?
If yes for all three, please express your interest.
Q: My figures are generated using SPSS, MATLAB or programming languages other than Python or R, can I make my paper executable?
A: Not at the moment – as we add compatibility for other languages, we will keep the community informed via our ERA community update.
Q: What if I’m already using Jupyter Notebook/R Markdown for data analysis/writing my articles?
A: That will make composing your ERA a lot easier – Using Stencila Hub, you can take a copy of your paper published on the eLife website and convert it to a Jupyter Notebook/R Markdown file. You can then download it and add the appropriate code chunks. Please read the tutorial for more information.
Q: What size/type of data can I upload to Stencila?
A: Theoretically there is no limit on size or type – but for this phase of development we ask you to please stick to tabular data (e.g. csv, tsv).
Q: Can I cite the executable complement?
A: You cannot at the moment – please cite the standard eLife article.
Q: Is my code/data going to be reviewed?
A: There will be some minimal checks to make sure that the code is not generating errors, but other than that, the code will be presented as-is. We are working with other code and data reviewing initiatives to improve this in the near future.
Q: Will this increase the amount of time it takes for my paper to be reviewed?
A: No, your paper would need to have been published already for you to be eligible for an ERA complement.
Q: How long will it take for me to compose an ERA?
A: It depends on the complexity of your code and paper – it could take anything between a few hours to 2-3 weeks.
Q: Does that mean people can do whatever they want with my paper?
A: Yes and no. Yes, because people can explore your data and change your code in their browsers; no, because none of these changes will be saved, nor will they affect anyone else’s views on your data and code.
Q: What happens when a package that my analysis is dependent on is updated/not maintained and my code doesn’t work anymore?
A: This should not affect the code that’s published, Stencila Hub will make sure that the correct (legacy) version of the package is installed when it is started.
Q: Do I have to convert all of my figures into executable versions? What if some (parts) of my figures cannot be made executable, e.g. images/diagrams?
A: You do not have to convert all of your figures into executable versions. When a single figure contains multiple panels/images, and some but not all of these could be made executable, it would be acceptable to re-capture these as separate Figures or figure supplements, as appropriate, so that those panels/images can be made executable.
Q: Can I only enrich figures? What about inline numbers/tables?
A: Inline numbers and tables can also be enriched with code.
Q: Can I style the executable version of my figures differently from those already published?
A: Please style your figures as closely as possible to the ones in your published eLife article – this will help speed up our quality control processes for the ERA. See above for figures which are only partially executable.
Q: Can I enrich figure supplements?
A: Yes
Q: How can I attach data – can I link to data published on other repositories, e.g. Zenodo, Figshare, etc?
A: Yes – you can upload tabular data and also add URLs of data sources to your Stencila Hub project.
Q: Can I preview my ERA before it’s published?
A: Yes, you can preview what your ERA will look like on Stencila Hub, before it is shared with the eLife Production Team.
Developers and publishers
Q: Is this an eLife-exclusive feature?
A: Quite the opposite. eLife and Stencila developed ERA as open-source software specifically to encourage other publishers to offer the format to their authors. Interested publishers should contact us at innovation [at] elifesciences [dot] org.
Q: Why not just use Jupyter or a commercial solution?
The ERA technology stack is designed specifically to be faster at loading, displaying and executing articles than Jupyter Notebooks, and to provide a reading experience that’s closer to that of a traditional online publication. ERA’s theming and format conversion capabilities also let publishers more easily integrate ERA content within their journal websites. Finally, unlike commercial solutions in this space, ERA’s open-source nature means there is no lock-in to a singler vendor – publishers can even run their own instance of the tool stack should they choose to, although they may want to choose an implementation hosted by Stencila as a more cost-effective option.
Q: How do I contribute to the development of this project?
A: You can sign up to our technology and innovation nesletter to receive ERA community update for the latest news and opportunities to contribute, and/or explore Stencila’s repositories on GitHub. If you see opportunities to collaborate, please don’t hesitate to email innovation [at] elifesciences [dot] org
Q: Who is paying for the hosting/compute?
A: eLife is paying at the moment – we hope to use this initial phase to assess the costs associated with these executable articles. Individual publishers wishing to adopt the technology will be able to negotiate their own hosted execution pricing directly with Stencila, or run their own instance of the technology stack on their own compute infrastructure.
#
We welcome comments, questions and feedback. Please annotate publicly on the article or contact us at innovation [at] elifesciences [dot] org.
Do you have an idea or innovation to share? Send a short outline for a Labs blogpost to innovation [at] elifesciences [dot] org.
For the latest in innovation, eLife Labs and new open-source tools, sign up for our technology and innovation newsletter. You can also follow @eLifeInnovation on Twitter.




