Author response:
The following is the authors’ response to the original reviews.
eLife Assessment
The study presents valuable findings of an optimized E. coli cell-free protein synthesis (eCFPS) system that has been simplified by reducing the number of core components from 35 to 7; furthermore, the findings communicate a simplified 'fast lysate' preparation that eliminates the need for traditional runoff and dialysis steps. This study is an advance towards simplifying protein expression workflows, and the evidence provided is solid, starting with nanoluc, a protein that expresses readily in many systems, to applications to more challenging proteins like the functional self-assembling vimentin and the active restriction endonuclease Bsal. Data on the underlying mechanisms and efficiency of the presented system in terms of protein yield relative to other known cell-free systems would greatly enhance the findings' significance and the strength of the evidence. The paper remains of interest to scientists in microbiology, biotechnology and protein synthesis.
We thank the editors for the positive assessment of our optimized E. coli cellfree protein synthesis (eCFPS) system and the "fast lysate" preparation.
As suggested, we have significantly strengthened the evidence by adding:
(1) Mechanism data: We have integrated a detailed analysis of the endogenous metabolic pathways (amino acids and nucleotides) into the Discussion section, supported by literature (Prinz et al. 1997; Yokoyama et al. 2010; Kigawa et al. 1999).
(2) Efficiency comparisons: We have added quantitative comparisons of absolute protein yields between our simplified 7-component system and the conventional 35-component system (now in Figure S3 E-F), demonstrating that our system matches or exceeds traditional titers.
Public Reviews:
Reviewer #1 (Public review):
The authors only provided the data for optimization, leaving the underlying mechanism that explains the phenomena unexplained.
We appreciate this feedback. To address the mechanism of how protein synthesis persists without exogenous additives, we have expanded the Discussion to explain how the "fast lysate" retains active endogenous enzymes. By omitting runoff and dialysis, our system preserves the metabolic capacity to synthesize amino acids (e.g., Cys and Trp from Ser) and nucleotides from residual precursors, as supported by the literature (Prinz et al. 1997; Yokoyama et al. 2010; Kigawa et al. 1999).
Reviewer #2 (Public review):
The production of the lysate requires special instrumentation, limiting accessibility. While the strengths of the study are well-emphasized, the limitations are not mentioned.
We thank the reviewer for this point. While a high-pressure homogenizer is common in many molecular biology labs, we acknowledge it may be a barrier for some. We have now included a dedicated Limitations paragraph in the Discussion addressing accessibility and the inherent challenges of prokaryotic systems in producing complex human proteins requiring post-translational modifications.
Reviewer #3 (Public review):
(1) Clarification on "highly efficient" and the lack of comparison with typical high-yield systems.
We have clarified "highly efficient" as a holistic balance of high yield, robustness, and simplified preparation. Crucially, we added absolute yield data (sfGFP standard curve) to Figure S3E-F demonstrating that our 7-component system performs comparably to or better than traditional high-yield protocols.
(2) How did the authors ensure chemical composition only affected translation and not transcription?
This is a key distinction. We performed new experiments using pretranscribed mRNA templates (Figure S3G) to isolate translational effects. While translation efficiency slightly decreased in the simplified buffer, the overall protein yield increased significantly due to a dramatic boost in transcription efficiency, confirming the system's net performance gain.
Recommendations for the authors:
Reviewer #1 (Recommendations for the authors):
There are specific concerns that need to be addressed:
(1) On page 4, lines 103-109, the authors speculate that protein synthesis persists even in the absence of amino acids like arginine, cysteine, and tryptophan. They suggest that this is likely due to residual amounts of these amino acids present in the cell lysate. Yokoyama et al. demonstrated that these amino acids are generated from other amino acids by endogenous amino acid metabolic enzymes in the cell lysate (J. Biomol. NMR 48, 193, (2010), doi: 10.1007/s10858-010-9455-3.). Cysteine and tryptophan can be derived from serine. In this context, asparagine and glutamine can be disregarded because they are synthesized from aspartate and glutamate, respectively. A more indepth analysis is required to interpret the results accurately.
We thank the reviewer for this insightful comment and for pointing us toward the relevant literature. We agree that the persistence of protein synthesis in the absence of exogenous amino acids like Arg, Cys, and Trp is driven by the robust metabolic capacity of our "fast lysate."
Unlike conventional protocols, our "fast lysate" procedure deliberately omits runoff and dialysis steps, ensuring the maximal retention of active endogenous metabolic enzymes and residual small-molecule pools. As demonstrated by Yokoyama et al. (2010), E. coli cell extracts retain functional enzymes capable of synthesizing acid-sensitive amino acids from precursors or more stable amino acids. We have integrated a detailed mechanistic analysis of these endogenous metabolic pathways into the Discussion section and have cited Yokoyama et al. (2010) to support this interpretation.
(2) On page 4, lines 111-115, the authors demonstrated that protein synthesis could occur even in the absence of CTP or UTP, provided ATP and GTP are present. This phenomenon can also be attributed to the analogous complementary actions of metabolic pathways.
We agree with the reviewer's assessment. The ability of the optimized eCFPS to function without exogenous CTP/UTP relies on the same principle of endogenous metabolic conversion mentioned above. The omission of dialysis ensures that the lysate retains not only residual nucleotide pools but also the full suite of nucleotide metabolic enzymes. Powered by our optimized energy regeneration system, these enzymes maintain sufficient levels of CTP and UTP to support transcription and translation. This explanation has been added to the Discussion section to clarify the robustness of our system.
(3) On Figure 3A, protein synthesis kinetics are presented in a stair plot instead of the commonly used scatterplot. Is there a specific reason for choosing the stair plot?
We chose the stair plot representation to more clearly visualize the cumulative process of protein synthesis and its stabilization over discrete time intervals. Given that sampling occurred every 10 minutes, a stair plot effectively highlights the "plateau" phases and the incremental nature of accumulation, which can sometimes be obscured by dense scatter plots.
(4) On Figure 3C. It is unclear which system is referred to as the "initial" system in Figure 3C. Which data point on Figures 3A and 3B corresponds to this "initial" system?
We apologize for the lack of clarity. In Figure 3C, "initial" refers to the traditional 35-component system prior to our streamlining process. Figures 3A and 3B characterize the performance of the final optimized system alone. To resolve this ambiguity, we have updated the legend for Figure 3 to explicitly define the "initial" system as the pre-optimization control.
(5) In Figure 5D, previously reported eCFPS and the system using "fast lysate" were compared. The only difference between the two systems seems to be the type of lysate used, according to the Supplementary table. Optimal concentrations for the components are the same for both lysates, or is there still room for optimization for "fast lysate"?
The "fast lysate" primarily differs from conventional lysates in its preparation speed and the retention of endogenous cofactors/enzymes. While the optimal salt and energy concentrations remained consistent across both lysates in our tests, the "fast lysate" provides a higher baseline signal due to the endogenous T7 RNA polymerase and metabolic factors. We believe this demonstrates the robustness of the optimized reaction buffer across varying lysate preparation qualities.
(6) The study suggests that the removal of DTT didn't negatively affect protein expression. However, based on my experience, certain proteins, especially those with cysteine residues on their surface, tend to aggregate without DTT. Did the authors attempt to express such proteins, or did they draw this conclusion based on the limited number of proteins tested?
This is a valid concern. We based our conclusion on the functional expression of Bsal and vimentin—two proteins that are inherently prone to aggregation and misfolding. Their successful synthesis suggests that the intrinsic reducing capacity of the lysate (e.g., glutathione and thioredoxin systems) is sufficient for many targets (Prinz et al. 1997). However, we acknowledge that specialized cysteine-rich proteins may still require exogenous DTT. We have addressed this in the Discussion.
Reviewer #2 (Recommendations for the authors):
(1) Line 77-78 "we iteratively evaluated the contribution of individual constituents through luciferase reporter assays" - where is all the data? Please use an appropriate figure citation. Figure 1 cherry picks some components, but I think all data should be included.
We have structured the data presentation to show dispensable components in Figure 1 (where removal does not inhibit reaction) and essential components in Figure 2 (where 0-concentration results in zero activity). This ensures a logical flow of the "streamlining" narrative. All raw data for these screenings have been included in the Source Data files.
(2) Line 127 typo "concentrations".
We thank the reviewer for pointing out this error. The typo "concentrations" has been corrected.
(3) Figure 2: "protein expression levels" measured how?/what is the unit of the vertical bar on the right? I'm assuming that this experiment was conducted for discrete concentrations and thus generated discrete data points. However, the graph makes it seem as if this is continuous data. Kindly change the type of graphing to indicate that this is discrete data, showing each data point.
We appreciate the reviewer's suggestion. Protein expression levels were measured using the Nanoluciferase (NLuc) reporter gene assay. We utilized heatmaps/contour plots because our data are bivariate, representing the simultaneous optimization of two concentrations (e.g., Mg2+ and K+ in Figure 2A). For such matrix-based screenings, heatmaps are significantly more effective than scatter plots at conveying synergistic trends and identifying optimal reaction landscapes. Notably, this visualization approach for discrete biochemical optimization data was successfully employed by Ban lab in their recent study on translation system optimization (Bothe and Ban 2024). The vertical color bar on the right represents the relative expression ratio, normalized to the maximum yield. Although we have provided a scatter plot of this discrete data for reference (see Author response image 1), we believe it appears visually cluttered due to the high density of data points, making it difficult to discern overarching trends. Heatmaps, by contrast, offer a much clearer representation of the optimal reaction landscape. To maintain transparency, the discrete concentration points tested are clearly reflected by the axis ticks, and all raw discrete data are available in the Source Data files.
Author response image 1.
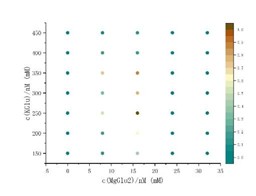
(4) Also, for all figures: the way the units are presented (DTT/mM) is confusing to me; it could just be something like [DTT] (mM).
We have revised all figures and tables to follow the standard format (e.g., [Component] (unit)) as suggested.
(5) Do the sucrose gradient sedimentation data have replicates? If so, please indicate statistics.
The sucrose gradient data provided (Figure 5C) is intended as qualitative evidence that the "fast lysate" method preserves intact 70S ribosomes across different preparation batches. This experiment has been performed independently multiple times with consistent results, demonstrating the high reproducibility of our preparation method. While we did not perform a quantitative comparative analysis of ribosome concentration, the consistency of the peaks confirms the integrity of the translational machinery.
(6) Line 457: fix the red line.
We thank the reviewer for pointing this out. The formatting issue has been resolved in the revised manuscript.
(7) Please mention the limitations of this study in the discussion.
We thank the reviewer for this suggestion. We have added a paragraph to the Discussion addressing the limitations of prokaryotic systems regarding complex eukaryotic post-translational modifications and chaperone requirements.
(8) Please include all uncropped gels in the source data, alongside the raw data, as you have already done.
As requested, we have provided all original, uncropped gel images in the Source Data files, alongside the raw data, to ensure full transparency and compliance with the journal's data sharing policies.
Reviewer #3 (Recommendations for the authors):
(1) The study lacks a comparison of protein levels with a typical cell-free protein synthesis system.
We have performed new quantitative experiments (now included in Figure S3 E-F) to measure absolute protein yields. Our optimized system achieves yields comparable to, or exceeding, several widely recognized highyield protocols while utilizing significantly fewer components. We have also clarified in the text that "highly efficient" refers to the synergistic balance of high yield, low cost, and simplified preparation time.
(2) What do the authors mean by "highly efficient", often used in the manuscript?
We thank the reviewer for the opportunity to clarify our terminology. We have performed new quantitative experiments (now included in Figure S3) to measure absolute protein yields, demonstrating that our optimized system achieves yields comparable to, or exceeding, several widely recognized highyield protocols while utilizing significantly fewer components.
In the context of this manuscript, we use the term "highly efficient" as a holistic descriptor that encapsulates three key dimensions of the system:
(1) Performance Superiority: Achieving higher expression levels and faster kinetics compared to conventional 35-component systems.
(2) Functional Robustness: The ability to efficiently synthesize challenging targets, such as cytotoxic proteins (BsaI) and aggregation-prone proteins (vimentin), which often fail in simplified systems.
(3) Practical Utility: A drastic reduction in preparation time and cost through the "fast lysate" protocol and the removal of 28 auxiliary components, thereby lowering the barrier to adoption.
This definition aligns with the study's core objective: developing a system where efficiency is measured not only by final yield but by the synergy between high performance and extreme ease of use.
(3) In this article, the term 'optimisation' is used as a synonym for 'simplification'. In biochemistry, optimisation commonly refers to an increase in yield, or the same yield achieved more easily or at a lower cost. In this case, however, we have no idea how this new system compares to a conventional expression system in terms of yield.
We thank the reviewer for this conceptual clarification. We agree that in biochemistry, "optimization" typically implies an improvement in yield or cost-effectiveness. In our study, we use the term to describe the process of achieving a superior balance between system simplicity and protein production. To address the reviewer's concern regarding the lack of a direct yield comparison, we have added new data in Figure S3. This figure provides a sideby-side comparison of protein yields between our simplified 7-component system and the conventional 35-component system. The results demonstrate that our system not only matches the performance of the traditional setup but frequently exceeds it in terms of final protein titer, while significantly reducing the reagent cost and preparation complexity. Thus, the simplification achieved in this work represents a true biochemical optimization of the cell-free synthesis process.
(4) The levels of transcripts of the proteins studied were not determined in any of the experiments performed. Therefore, it is unknown whether the effects of different experimental conditions on NLuc, GFP or other protein expression are due to an effect on transcription, translation, or both.
This is an excellent point. We performed a new set of experiments using mRNA templates instead of DNA to isolate the effects on translation (Figure S3G). Our results indicate that while the system's overall boost in NLuc expression is partially attributable to enhanced transcription efficiency, the translation machinery remains highly robust. We have updated the Results and Discussion to reflect this distinction.
References
Bothe, Adrian, and Nenad Ban. 2024. “A Highly Optimized Human in Vitro Translation System.” Cell Reports Methods 4 (4): 100755.
Kigawa, T., T. Yabuki, Y. Yoshida, M. Tsutsui, Y. Ito, T. Shibata, and S. Yokoyama. 1999. “Cell-Free Production and Stable-Isotope Labeling of Milligram Quantities of Proteins.” FEBS Letters 442 (1): 15–19.
Prinz, W. A., F. Aslund, A. Holmgren, and J. Beckwith. 1997. “The Role of the Thioredoxin and Glutaredoxin Pathways in Reducing Protein Disulfide Bonds in the Escherichia Coli Cytoplasm.” The Journal of Biological Chemistry 272 (25): 15661–67.
Yokoyama, Jun, Takayoshi Matsuda, Seizo Koshiba, and Takanori Kigawa. 2010. “An Economical Method for Producing Stable-Isotope Labeled Proteins by the E. Coli Cell-Free System.” Journal of Biomolecular NMR 48 (4): 193–201.




