Author response:
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public review):
Giordano et al. demonstrate that yeast cells expressing separated N- and C-terminal regions of Tfb3 are viable and grow well. Using this creative and powerful tool, the authors effectively uncouple CTD Ser5 phosphorylation at promoters and assess its impact on transcription. This strategy is complementary to previous approaches, such as Kin28 depletion or the use of CDK7 inhibitors. The results are largely consistent with earlier studies, reinforcing the importance of the Tfb3 linkage in mediating CTD Ser5 phosphorylation at promoters and subsequent transcription.
Notably, the authors also observe effects attributable to the Tfb3 linker itself, beyond its role as a simple physical connection between the N- and C-terminal domains. These findings provide functional insight into the Tfb3 linker, which had previously been observed in structural studies but lacked clear functional relevance. Overall, I am very positive about this manuscript and offer a few minor comments below that may help to further strengthen the study.
We appreciate the reviewer’s positive assessment of our work and suggestions for improvement.
(1) Page 4
PIC structures show the linker emerging from the N-terminal domain as a long alpha-helix running along the interface between the two ATPase subunits, followed by a turn and a short stretch of helix just N-terminal to a disordered region that connects to the C-terminal region (see schematic in Figure 1A).
The linker helix was only observed in the poised PIC (Abril-Garrido et al., 2023), not in other fully-engaged PIC structures.
Thanks for clarifying. We note that some structures of TFIIH alone also see the long helix. Accordingly, we modified this section to read:
“In many TFIIH and PIC structures the linker is not visible, presumably due to flexibility. However, when it is seen (Abril-Garrido et al., 2023; Greber et al., 2019), the linker emerges from the N-terminal domain as a long alpha-helix running along the interface between the two ATPase subunits…”
(2) Page 8
Recent structures (reviewed in (Yu et al., 2023)) show that the Kinase Module would block interactions between the Core Module and other NER factors. Therefore, TFIIH either enters into the NER complex as the free Core Module, or the Kinase Module must dissociate soon after.
To my knowledge, this is still controversial in the NER field. I note the potential function of the kinase module is likely attributed to the N-terminal region of Tfb3 through its binding to Rad3.
We are not experts on NER, but in reviews of the field this appears to be a widely held assumption. A 2008 paper from the Egly lab (Coin et al., DOI 10.1016/j.molcel.2008.04.024) is usually cited, which shows that the interaction between XPD (metazoan Rad3) and XPA is likely incompatible with XPD-MAT1 interaction. In addition to the Yu 2023 review, we now also cite a more recent publication that more extensively reviews the models for core TFIIH interactions (van Sluis et al, 2025). We looked at the multiple recently published structures of various TCR-NER and GG-NER intermediate complexes, and none of them show the CAK module or even the Tfb3/Mat1 N-term, even though those proteins were typically included during assembly. We also consulted with our colleagues Johannes Walter and Lucas Farnung, who are studying various TC-NER intermediates biochemically and structurally. Although the CAK module is included in their assembly reactions, it is not visible in their cryoEM structures. They tell me that the presence of CAK would be compatible with early TC-NER intermediates, but is predicted to overlap with later interactions of XPD with the TC-NER factor STK19 (see Mevissen et al., Cell 2024). To be conservative, we modified the sentence to say “Recent structures … suggest” rather than “show”.
Because the yeast strains used in Figure 6 retain the N-terminal region of Tfb3, the UV sensitivity assay presented here is unlikely to directly address the contribution of the kinase module to NER.
We agree that our experiment only shows that the connection between Tfb3 N- and C-term domains is not necessary for NER. The individual domains might still be able to function independently. Accordingly, we changed the heading of that section from “Disconnected core TFIIH does not cause an NER defect” to “Split Tfb3 does not cause an NER defect.” This more closely matches the figure legend title.
(3) Page 11
Notably, release of the Tfb3 Linker contact also results in the long alpha-helix becoming disordered (Abril-Garrido et al., 2023), which could allow the kinase access to a far larger radius of area. This flexibility could help the kinase reach both proximal and distal repeats within the CTD, which can theoretically extend quite far from the RNApII body.
Although the kinase module was resolved at low resolution in all PIC-Mediator structures, these structural studies consistently reveal the same overall positioning of the kinase module on Mediator, indicating that its localization is constrained rather than variable. This observation suggests that the linker region may help position the kinase module at this specific site, likely through direct interactions with the PIC or Mediator. This idea is further supported by numerous cross-links between the linker region and Mediator (Robinson et al., 2016).
That is true. But please note that this sentence was meant to describe movement of the kinase module AFTER release from Mediator (see previous sentence). Re-reading the passage, we realized the confusion is because we propose multiple possible pathways in that paragraph. In the first half, we suggest the capture of the kinase module by Mediator might trigger the conformation changes in the linker. In the second half (where it says “Alternatively….”) we suggest the Mediator-CAK interaction could instead come first, and the release of this contact could free the CAK module to move around. We have modified the paragraph to make it clear these are two different distinct models.
Reviewer #2 (Public review):
Summary:
This work advances our understanding of how TFIIH coordinates DNA melting and CTD phosphorylation during transcription initiation. The finding that untethered kinase activity becomes "unfocused," phosphorylating the CTD at ser5 throughout the coding sequence rather than being promoter-restricted, suggests that the TFIIH Core-Kinase linkage not only targets the kinase to promoters but also constrains its activity in a spatial and temporal manner.
Strengths:
The experiments presented are straightforward, and the models for coupling initiation and CTD phosphorylation and for the evolution of these linked processes are interesting and novel. The results have important implications for the regulation of initiation and CTD phosphorylation.
Weaknesses:
Additional data that should be easily obtainable and analysis of existing data would enable an additional test of the models presented and extract additional mechanistic insights.
We thank the reviewer for the positive assessment and address their specific suggestions below.
Reviewer #3 (Public review):
Summary:
Eukaryotic gene transcription requires a large assemblage of protein complexes that govern the molecular events required for RNA Polymerase II to produce mRNAs. One of these complexes, TFIIH, comprises two modules, one of which promotes DNA unwinding at promoters, while the other contains a kinase (Kin28 in yeast) that phosphorylates the repeated motif at the C-terminal domain (CTD) of the largest subunit of Pol II. Kin28 phosphorylation of Ser5 in the YSPTSPS motif of the CTD is normally highly localized at promoter regions, and marks the beginning of a cycle of phosphorylation events and accompanying protein association with the CTD during the transition from initiation to elongation.
The two modules of TFIIH are linked by Tfb3. Tfb3 consists of two globular regions, an N-terminal domain that contacts the Core module of TFIIH and a C-terminal domain that contacts the kinase module, connected by a linker. In this paper, Giordano et al. test the role of Tfb3 as a connector between the two modules of TFIIH in yeast. They show that while no or very slow growth occurs if only the C-terminal or N-terminal region of Tfb3 is present, near normal growth is observed when the two unlinked regions are expressed. Consistent with this result, the separate domains are shown to interact with the two distinct TFIIH modules. ChIP experiments show that the Core module of TFIIH maintains its localization at gene promoters when the Tfb3 domains are separated, while localization of the kinase module and of Ser5 phosphorylation on the CTD of Pol II is disrupted. Finally, the authors examine the effect of separating the Tfb3 domains on another function of TFIIH, namely nucleotide excision repair, and find little or no effect when only the N-terminal region of Tfb3 or the two unlinked domains are present.
Strengths:
Experiments involving expression of Tfb3 domains in yeast are well-controlled, and the data regarding viability, interaction of the separate Tfb3 domains with TFIIH modules, genome-wide localization of the TFIIH modules and of phosphorylated Ser5 CTDs, and of effects on NER, are convincing. The experiments are consistent with current models of TFIIH structure and function and support a model in which Tfb3 tethers the kinase module of TFIIH close to initiation sites to prevent its promiscuous action on elongating Pol II.
We appreciate that the reviewer finds that our main conclusions are convincing.
Weaknesses:
(1) The work is limited in scope and does not provide any major insights into the mechanism of transcription. One indication of this limitation is that in the Discussion, published structural and functional results on transcription are used to support the interpretations of the results here more than current results inform previous models or findings.
The story we present here is pretty simple, so in that sense we agree it is limited. However, we believe the findings do have mechanistic implications. That the Tfb3/Mat1 tether not only targets kinase activity to the 5’ end, but also somehow limits it from acting downstream seems significant. As for the Discussion, in our papers we always attempt to tie in our results and models with as much of the relevant published literature as possible. We believe this is more interesting, useful, and convincing than simply summarizing the Results section.
(2) The first described experiment, which purports to show that three kinases cannot function in place of Kin28 when tethered (by fusion) to Tfb3, is missing the crucial control of showing that Kin28 can support viability in the same context. This result also does not connect with the rest of the manuscript.
Our original motivation for the experiment in Figure 1 was to develop a system where we could plug different kinases into the CTD-proximal position. This didn’t work, so it is true that this negative result is somewhat unconnected to the rest of the paper. We choose to include it because it produced the unexpected observation that the Tfb3 C-term domain was not essential for viability, contradicting an earlier report. As for the suggested control of fusing Kin28, please see our reply to the editor’s comments below.
(3) Finally, the authors present the interesting and reasonable speculation that the TFIIH complex and connecting Tfb3 found in mammals and yeast may have evolved from an earlier state in which the two TFIIH subdomains were present as unconnected, distinct enzymes. This idea is supported by a single example from the literature (T. brucei). A more thorough evolutionary analysis could have tested this idea more rigorously.
Please see our full reply to Point 5 in the editor’s comments. In short, T. brucei was the only primitive eukaryote for which h we found an actual biochemical analysis of TFIIH. However, we now cite some papers reporting protein sequence comparisons for organisms not having a consensus CTD, which lend further support to our idea of fusion of a CDK to TFIIH co-evolved with the CTD during very early in eukaryotic evolution.
Recommendations for the authors:
Reviewing Editor Comments:
Suggestions for Improvement:
(1) Analyze existing Pol II ChIP-seq data to determine whether reduced TSS-proximal vs. gene-body occupancy observed with the split Tfb3 alleles reflects initiation defects, and whether different gene classes (high vs. low expression, stress-induced genes) show differential effects of splitting Tfb3.
Thanks for the suggestion. The new analysis is included as Supplemental Figure S6. Several factors indicate an initiation defect rather than an elongation defect (either elongation processivity or elongation rate). First, the shape of the RNApII occupancy trace is flat in all mutants, arguing against a processivity defect, which would have led to a downward slope due to RNApII progressively dropping off from the gene. Because this effect is best seen on long genes (more than 2kb), we generated metagene profiles on long, well-expressed genes only, which led to the same conclusion (see Sup Fig 6A). Second, the mutants lead to decreased RNApII occupancy, arguing against a strong decrease in elongation rate, which -if anything- would have led to an increase in RNApII during early transcription. While we cannot completely exclude the possibility of a mild decrease in elongation rate, such an effect doesn’t fit the patterns we observe. The overall decrease of RNApII occupancy is rather a strong indication of a decrease in early steps (PIC assembly or initiation).
As requested, we looked at potential differences between gene classes two ways. First, we generated RNApII metagenes on RNApII occupancy quintiles (Q1-Q5). As shown in Sup Fig 6B, RNApII occupancy is similarly decreased in all mutants for all quintiles, demonstrating that the effect of Tfb3 splitting on transcription is not linked to expression level. Second, we generated RNApII occupancy metagenes for TFIID-regulated genes and coactivator redundant (CR) genes. This classification from the Hahn lab (doi:10.7554/eLife.50109) is very similar to the one developed by the Pugh lab (doi:10.1016/s1097-2765(04)00087-5). TFIID-regulated genes are enriched for housekeeping genes and are typically devoid of a TATA box, while the CR genes tend to be highly regulated and to contain a TATA box. As shown in Sup Fig 6C, the effect of the Tfb3 split mutants is similar on both gene classes.
(2) Determine whether Kin28 abundance in whole cell extracts is reduced by splitting Tfb3, as a factor in reducing its occupancies at gene promoters.
We actually did test for Kin28 and Ccl1 levels in the extracts when we did the IP experiment shown in Fig 3. We ran the extracts next to the precipitated factors. Unfortunately, as can be seen on the bottom blot, our antibodies were not strong enough to detect either Kin28 or Ccl1 in extracts, even with WT Tfb3. Although we don’t include this inconclusive result in the final paper, we show it in Author response image 1 (note that extracts are labeled as “IgG input”).
Author response image 1.
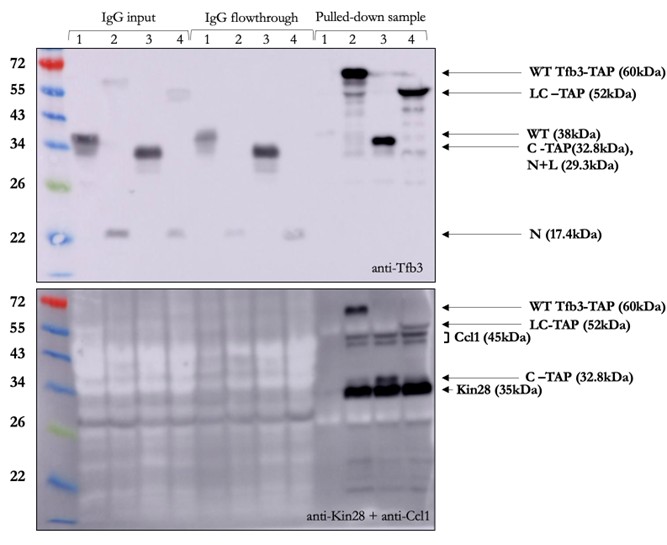
(3) Include the key positive control construct of replacing the C-term of Tfb3 with Kin28 in the experiments of Figure 1.
We elected not to do this experiment for several reasons. As reviewer 3 points out, this kinase fusion experiment turned out to be somewhat disconnected from the rest of the paper. Even though it didn’t work, we included it in the paper because the results led us to the realization that the Tfb3 C-term was actually not fully essential for viability as reported, which in turn led us to the idea of splitting Tfb3. Structural studies (https://doi.org/10.1126/sciadv.abd4420, https://doi.org/10.1073/pnas.2009627117, https://doi.org/10.7554/eLife.44771) show that, in addition to providing linkage to the core module, the C-term of Tfb3 induces a conformation change in Kin28/Cdk7 necessary for full kinase activity (which is likely why the strains without C-term are just barely viable). If we were to pursue why the fusions didn’t work, we could tether Kin28 directly to the Tfb3 linker (and may try this in the future), but then would need to also express the C-term separately for its activating function. Even then, this would be an imperfect control for the fusion experiments in Figure 1. Because were trying to best mimic Kin28 being tethered via the accessory subunit Tfb3/Mat1, in the Figure 1 experiment we did not directly attach the kinases to Tfb3. For Ctk1/Cdk12, we fused the Tfb3 linker to the Ctk3 accessory subunit (analogous to Tfb3), and for Bur1/Cdk9, we fused to the cyclin subunit Bur2 (there is no known third subunit in this complex). The one exception was Mpk1, which has no partner subunits and is not a CDK. There are many reasons why this high-risk protein fusion experiment may not have worked, but we feel it’s not that useful to pursue it in this paper.
(4) Provide direct evidence for the claimed dominant negative effect of the N-term-Linker construct by extending results in Figure 2C to compare growth of WT TFB3 cells expressing this construct vs. vector alone.
We thank the reviewers for this suggestion. We tested this by transforming high copy plasmids expressing the different Tfb3 truncations into cells expressing the WT Tfb3. We did not see a clear dominant negative effect (some colonies were small, but many looked normal). Accordingly, in the absence of a reproducible effect, we removed this claim from the paper. In Fig 2C, the WT plasmid was transformed into cells already expressing the truncation on a high copy plasmid (the opposite order of our new experiment). It’s possible that phenotypes vary depending on which plasmid was there first (2 micron plasmids have variable copy number and can compete with each other for replication and passage during cell division). In any case, in the face of ambiguous results we no longer claim a dominant negative effect of the N-term-Linker protein. This was a minor side-point of the paper and does not affect any of our other conclusions.
(5) Expand the evolutionary analysis to provide evidence beyond the case of T. brucei that the Tfb3-mediated connection between core and kinase modules is an evolutionary addition to the ancestral state.
We note that the two papers we cited for the lack of a CAK module in T. brucei reached that conclusion based on purification of its TFIIH complex. We were unable to find similar biochemical studies in other primitive eukaryotes. Another way to expand the evolutionary comparison would be through sequence homology searches. We attempted to do this using various tools available at NCBI and EMBL. These show that Tfb3/Mat1 is found extensively throughout eukaryotes. Unfortunately, because the NTD of Tfb3 is a RING domain, homology searches in primitive eukaryotes yield a number of weak matches in the zinc binding motif, but no way of knowing if any of these are related to TFIIH. Similarly, searches with Cdk7/Kin28 or Cyclin H/Ccl1 pulls up all CDKs and cyclins, with roughly equal statistical similarity to the yeast kinase/cyclin. Someone with more experience with evolutionary analysis would likely have better luck, but our efforts were inconclusive. However, we did find two papers from Guo and Stiller (2004 and 2005) that analyzed genome sequences available at the time and reached the conclusion that both concensus CTD and the CAK module are absent in the evolutionary branch of primitive eukaryotes that contains T. brucei and Giardia lamblia. We also found papers identifying a putative Mat1/Tfb3 in Plasmodium falciparum, although this protein was not yet shown to be associated with TFIIH. We now cite these papers in the discussion of our evolutionary hypothesis.
(6) Include Western blot analysis of the Tfb3 chimeras and truncations analyzed in Figures 1-2 to determine if poor expression contributes to any of the poor-growth phenotypes.
The western blot of the Tfb3 fusions used in Figure 1 is shown in Sup Fig 1. The Tfb3 truncations are shown in the Input panel of Fig 3A (although some of these are TAP fusions, the growth phenotypes did not change with TAP-tagging). In general, all the fusions and truncations are detectable but possibly reduced relative to WT Tfb3. Note that the anti-Tfb3 antibody is a polyclonal made against recombinant Tfb3, and we don’t know that the reactive epitopes are distributed equally throughout the protein, so it’s difficult to be confident about relative quantitation with partial Tfb3 proteins.
(7) Provide direct evidence that the N-terminal Tfb3 segment interacts exclusively with the core TFIIH module and not Kin28, analogous to the opposite results shown in Figure 3B and 4A-B for the C-terminal domain.
This could be interesting, but we elected not do this experiment due to time and manpower limitations. Since the N-term is unambiguously essential for viability, we can assume it retains at least some interactions with core TFIIH (unless the N-term has some other essential function that hasn’t been discovered).
(8) Confirm that the Ser5P phosphorylation levels given by the different Tfb3-TAP immune complexes are all much higher than the background level observed with control complexes prepared with extracts expressing WT, untagged Tfb3.
We should have done this control in Sup Fig 2B, especially since we did pull down the beads from the untagged strain as shown in panel A. We haven’t seen appreciable kinase activity when we’ve done this control in the past, so we feel confident the signals seen are not background. Therefore, we elected not to repeat this experiment.
(9) Conduct an in vitro reconstitution comparing the activity of free kinase module and intact TFIIH on elongating RNA polymerase II in directing promoter-localized vs. downstream Ser5P accumulation.
This would be a nice experiment, but would require a substantial amount of work that is beyond our resources at the time.
(10) Revise the text to better emphasize any novel mechanistic insights afforded by the work and address all other minor comments/criticisms.
Done, as addressed in all the other comment replies.
Reviewer #2 (Recommendations for the authors):
(1) The authors suggest that their results support model 3, in which intact TFIIH restrains kinase activity outside the PIC. Directly testing this model would be a significant addition and would strengthen the proposed mechanism. An in vitro reconstitution comparing the activity of the free kinase module and intact TFIIH on elongating RNA polymerase II (or, at a minimum, purified Pol II) would directly test the mechanism underlying downstream Ser5P accumulation.
Sup Fig 2 addresses this point to some extent, since we the TAP pull-down of full-length Tfb3 precipitates at least some intact TFIIH, whereas the split C-term TAP constructs do not (as shown in Fig 4). However, this is not a very quantitative assay and we agree with the reviewer that a careful reconstitution, especially in the context of real transcription, would be far better. Unfortunately, this is currently beyond our capabilities. However, in the Discussion we do cite some published data arguing that association of the core TFIIH does have some inhibitory effect on the kinase module. First, in our 2002 MCB paper (Keogh et al., see Fig 7) using a GST-CTD kinase assay, we found that free kinase module (called TFIIK there) was strongly active even with a non-phosphorylatable mutation in the activating T-loop. In contrast, the same mutation inactivated CTD kinase activity in the intact TFIIH. Similarly, the Taatjes lab (Rimel et al., Genes Dev. 2020) found that free CAK was active on multiple substrates that were not phosphorylated by the full TFIIH complex.
(2) Experiments from Carl Wu's laboratory (Nguyen et al., 2021) showed that there is a significant amount of apparently free Kin28 as well as free TFIIH in cells. Please reference and comment on this when discussing the model, suggesting that TFIIH is mostly sequestered at promoters.
Good point. We added this to the discussion where we discuss the arguments against a sequestering model.
(3) The existing ChIP-seq data could be analyzed more thoroughly to extract additional mechanistic insights. Specifically: (i) quantify TSS-proximal vs. gene body Pol II to determine if reduced occupancy reflects initiation defects (ii) analyze whether gene classes (high vs. low expression, stress-induced genes) show differential effects.
Thanks for the suggestion. We did this and show the results as a new Supplemental Figure 6. No differences were found. Please see our response to the Editor’s comment #1 for a fuller description.
(4) The complete loss of Kin28 ChIP signal in mutant strains (Figure 5B) could reflect kinase mislocalization or reduced protein abundance. Figure 3B examines TAP-purified material but does not address total cellular protein levels. Examining whole-cell extracts for Kin28 and Ccl1 in all strains would strengthen the interpretation of the ChIP results.
As described in our response to Point 2 in the Editor’s comments section, we did do this control. Unfortunately, the Kin28 and Ccl1 antibodies were not strong enough to detect these proteins in extracts before precipitation.
Reviewer #3 (Recommendations for the authors):
(1) The experiment of Figure 1 should be repeated with a Tfb3-Kin28 positive control or dropped from the manuscript.
This could be an interesting experiment, but please see our response to Editor comment #3 for why we decided to keep the figure as is.
(2) Figure 2C legend doesn't mention linker C-term low copy construct.
Thanks for catching that error. It is now fixed.
(3) The claim that the N-term linker has a dominant negative effect (Figure 2C) requires direct comparison (growth on the same plate) of TFB3+ cells with and without expression of the N-term linker.
As detailed in the response to the Editor’s comment #4, we did this test. The results did not support a dominant negative phenotype, so we removed this claim. Thanks for helping us avoid a mistake.
(4) Page 7, "Supplementary Fig. S4A, B, promoters in green boxes" should read "Supplementary Fig. S5A, B, promoters in green boxes".
Thanks for catching that error. It is now fixed.
(5) Readers might be concerned that the ChIP-seq signal observed in Figure 5 and S5 could reflect an artifactual signal over highly transcribed regions. The different distributions of Rpb1, Ser5p, and Ser2p argue against this. This might be worth mentioning in the text.
Thanks for raising this issue. “Hyper-ChIPpable” genes can be a problem in metagene analysis. We now include the analysis suggested by Reviewer 2 where we separately look at genes with different transcription frequencies. Seeing the same relative patterns regardless of expression level makes us confident that the results are not artifactual.
(6) p. 12, "the Tfb3 the linker"; "In contrast, The N-term linker"; "suggest" should be "suggests"
We appreciate the reviewer’s careful reading of the manuscript and have corrected these typos.




