Author response:
The following is the authors’ response to the previous reviews.
Reviewer #2 (Public Review):
Weaknesses:
The comparison of affinity predictions derived from AlphaFold2 and H3-opt models, based on molecular dynamics simulations, should have been discussed in depth. In some cases, there are huge differences between the estimations from H3-opt models and those from experimental structures. It seems that the authors obtained average differences of the real delta, instead of average differences of the absolute value of the delta. This can be misleading, because high negative differences might be compensated by high positive differences when computing the mean value. Moreover, it would have been good for the authors to disclose the trajectories from the MD simulations.
Thanks for your careful checks. We fully understand your concerns about the large differences when calculating affinity. To understand the source of these huge differences, we carefully analyzed the trajectories of the input structures during MD simulations. We found that the antigen-antibody complex shifted as it transited from NVT to NPT during pre-equilibrium, even when restraints are used to determine the protein structure. To address this issue, we consulted the solution provided on Amber's mailing list (http://archive.ambermd.org/202102/0298.html) and modified the top file ATOMS_MOLECULE item of the simulation system to merge the antigen-antibody complexes into one molecule. As a result, the number of SOLVENT_POINTERS was also adjusted. Finally, we performed all MD simulations and calculated affinities of all complexes.
We have corrected the “Afterwards, a 25000-step NVT simulation with a time step of 1 fs was performed to gradually heat the system from 0 K to 100 K. A 250000-step NPT simulation with a time step of 2 fs was carried out to further heat the system from 100 K to 298 K.” into “Afterwards, a 400-ps NVT simulation with a time step of 2 fs was performed to gradually heat the system from 0 K to 298 K (0–100 K: 100 ps; 100-298 K: 200 ps; hold 298 K: 100 ps), and a 100-ps NPT simulation with a time step of 2 fs was performed to equilibrate the density of the system. During heating and density equilibration, we constrained the antigen-antibody structure with a restraint value of 10 kcal×mol-1×Å-2.” and added the following sentence in the Method section of our revised manuscript: “The first 50 ns restrains the non-hydrogen atoms of the antigen-antibody complex, and the last 50 ns restrains the non-hydrogen atoms of the antigen, with a constraint value of 10 kcal×mol-1×Å-2”
In addition, we have corrected the calculation of mean deltas using absolute values and have demonstrated that the average affinities of structures predicted by H3-OPT were closer to those of experimentally determined structures than values obtained through AF2. These results have been updated in the revised manuscript. However, significant differences still exist between the estimations of H3-OPT models and those derived from experimental structures in few cases. We found that antibodies moved away from antigens both in AF2 and H3-OPT predicted complexes during simulations, resulting in RMSDbackbone (RMSD of antibody backbone) exceeding 20 Å. These deviations led to significant structural changes in the complexes and consequently resulted in notable differences in affinity calculations. Thus, we removed three samples (PDBID: 4qhu, 6flc, 6plk) from benchmark because these predicted structures moved away from the antigen structure during MD simulations, resulting in huge energy differences from the native structures.
Author response table 1.
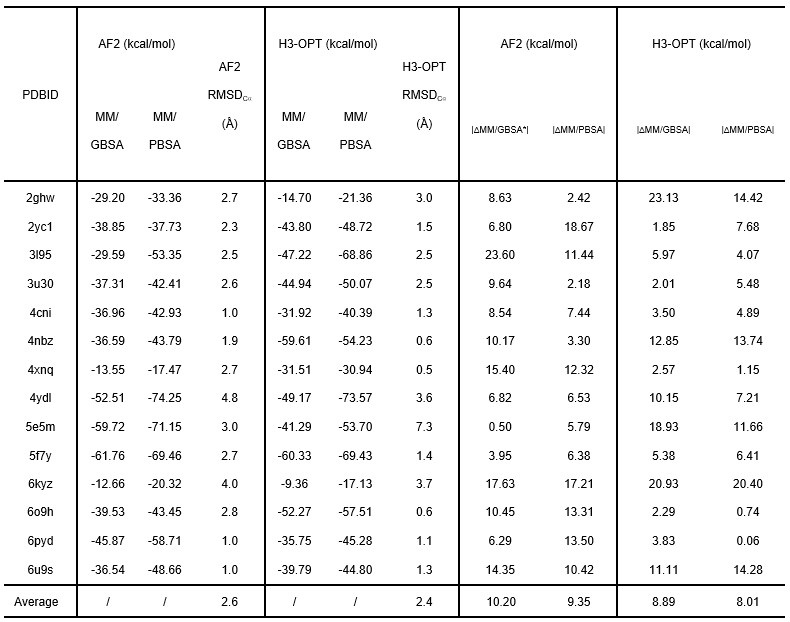
We also appreciate your reminder, and we have calculated all RMSDbackbone during production runs (SI Fig. 5).
Author response image 1.
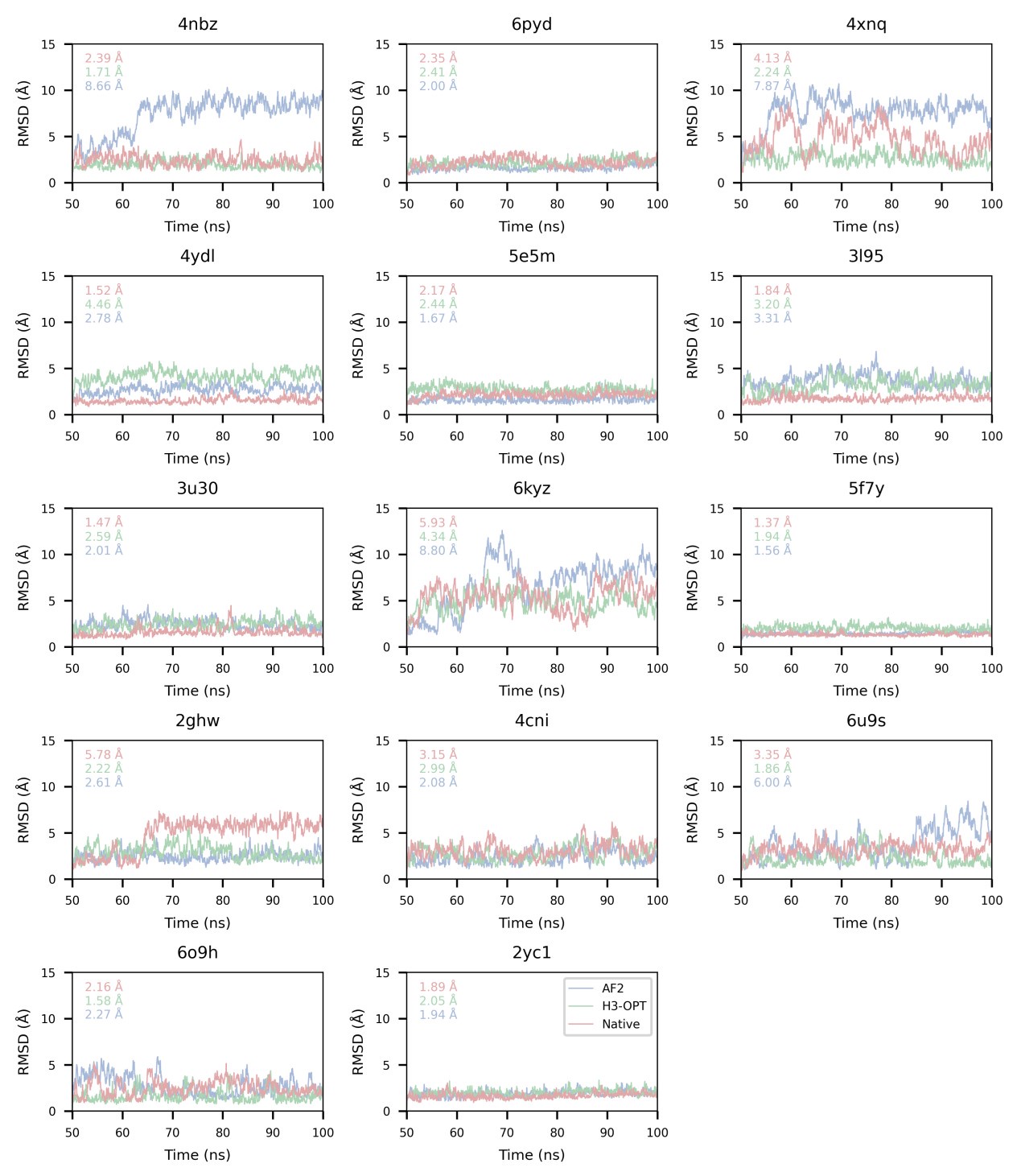
Reviewer #3 (Public Review):
Weaknesses:
The proposed method lacks of a confidence score or a warning to help guiding the users in moderate to challenging cases.
We were sorry for our mistakes. We have updated our GitHub code and added following sentences to clarify how we train this confidence score module in Method Section: “Confidence score prediction module
We apply an MSE loss for confidence prediction, label error was calculated as the Cα deviation of each residue after alignment. The inputs of this module are the same as those used for H3-OPT, and it generates a confidence score ranging from 0 to 100. The dropout rates of H3-OPT were set to 0.25. The learning rate and weight decay of Adam optimizer are set to 1 × 10−5 and 1 × 10−4, respectively.”
Reviewer #2 (Recommendations For The Authors):
I would strongly suggest that the authors deepen their discussion on the affinity prediction based on Molecular Dynamics. In particular, why do the authors think that some structures exhibit huge differences between the predictions from the experimental structure and the predicted by H3-opt? Also, please compute the mean deltas using the absolute value and not the real value; the letter can be extremely misleading and hidden very high differences in different directions that are compensating when averaging.
I would also advice to include graphical results of the MD trajectories, at least as Supp. Material.
We gratefully thank you for your feedback and fully understand your concerns. We found the source of these huge differences and solved this problem by changing method of MD simulations. Then, we calculated all affinities and corrected the mean deltas calculation using the absolute value. The RMSDbackbone values were also measured to enable accurate affinity predictions during production runs (SI Fig. 5). There are still big differences between the estimations of H3-OPT models and those from experimental structures in some cases. We found that antibodies moved away from antigens both in AF2 and H3-OPT predicted complexes during simulations, resulting in RMSDbackbone exceeding 20 Å. These deviations led to significant structural changes in the complexes and consequently resulted in notable differences in affinity calculations. Thus, we removed three samples (PDBID: 4qhu, 6flc, 6plk) from benchmark.
Thanks again for your professional advice.
Reviewer #3 (Recommendations For The Authors):
(1) I am pleased with the most of the answers provided by the authors to the first review. In my humble opinion, the new manuscript has greatly improved. However, I think some answers to the reviewers are worth to be included in the main text or supporting information for the benefit of general readers. In particular, the requested statistics (i.e. p-values for Cα-RMSD values across the modeling approaches, p-values and error bars in Fig 5a and 5b, etc.) should be introduced in the manuscript.
We sincerely appreciate your advice. We have added the statistics values to Fig. 4 and Fig. 5 to our manuscript.
Author response image 2.
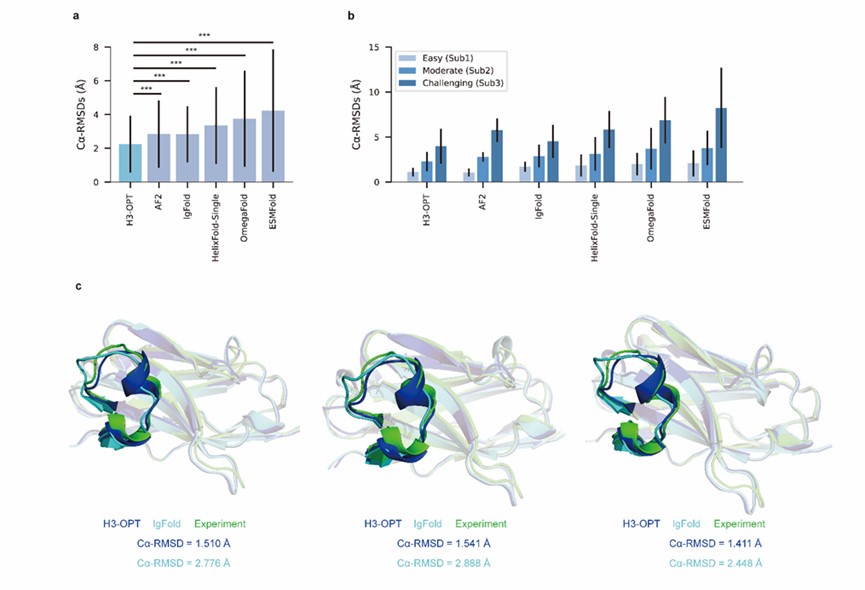
Author response image 3.
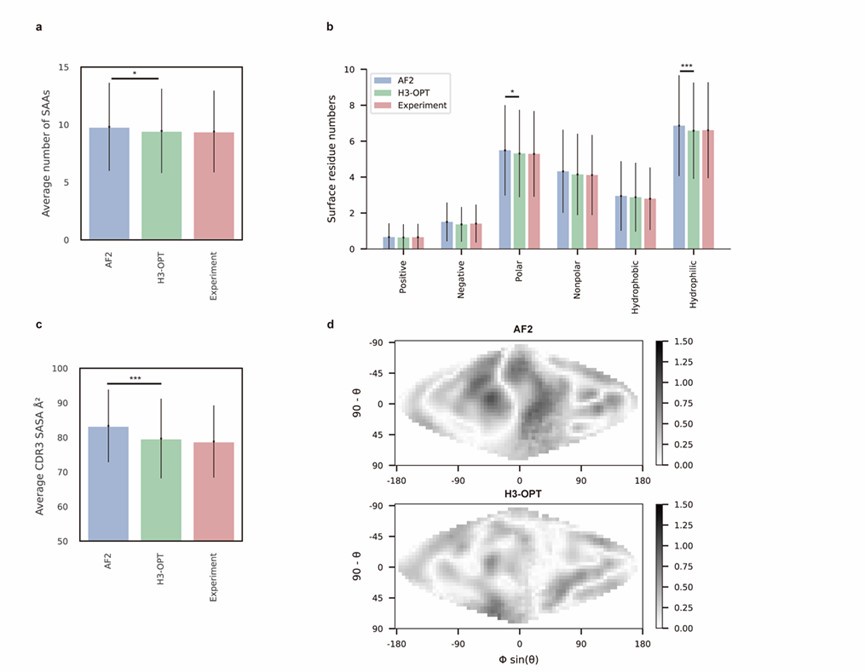
(2) Similarly, authors state in the answers that "we have trained a separate module to predict the confidence score of the optimized CDR-H3 loops". That sounds a great improvement to H3-OPT! However, I couldn't find any reference of that new module in the reviewed version of the manuscript, nor in the available GitHub code. That is the reason for me to hold the weakness "The proposed method lacks of a confidence score".
We were really sorry for our careless mistakes. Thank you for your reminding. We have updated our GitHub code and added following sentences to clarify how we train this confidence score module in Method Section:
“Confidence score prediction module
We apply an MSE loss for confidence prediction, label error was calculated as the Cα deviation of each residue after alignment. The inputs of this module are the same as those used for H3-OPT, and it generates a confidence score ranging from 0 to 100. The dropout rates of H3-OPT were set to 0.25. The learning rate and weight decay of Adam optimizer are set to 1 × 10−5 and 1 × 10−4, respectively.”
(3) I acknowledge all the efforts made for solving new mutant/designed nanobody structures. Judging from the solved structures, mutants Y95F and Q118N seems critical to either crystallographic or dimerization contacts stabilizing the CDR-H3 loop, hence preventing the formation of crystals. Clearly, solving a molecular structure is a challenge, hence including the following comment in the manuscript is relevant for readers to correctly asset the magnitude of the validation: "The sequence identities of the VH domain and H3 loop are 0.816 and 0.647, respectively, comparing with the best template. The CDR-H3 lengths of these nanobodies are both 17. According to our classification strategy, these nanobodies belong to Sub1. The confidence scores of these AlphaFold2 predicted loops were all higher than 0.8, and these loops were accepted as the outputs of H3-OPT by CBM."
We appreciate your kind recommendations and have revised “Although Mut1 (E45A) and Mut2 (Q14N) shared the same CDR-H3 sequences as WT, only minor variations were observed in the CDR-H3. H3-OPT generated accurate predictions with Cα-RMSDs of 1.510 Å, 1.541 Å and 1.411 Å for the WT, Mut1, and Mut2, respectively.” into “Although Mut1 (E45A) and Mut2 (Q14N) shared the same CDR-H3 sequences as WT (LengthCDR-H3 = 17), only minor variations were observed in the CDR-H3. H3-OPT generated accurate predictions with Cα-RMSDs of 1.510 Å, 1.541 Å and 1.411 Å for the WT, Mut1, and Mut2, respectively (The confidence scores of these AlphaFold2 predicted loops were all higher than 0.8, and these loops were accepted as the outputs of H3-OPT by CBM). ”. In addition, we have added following sentence in the legend of Figure 4 to ensure that readers can appropriately evaluate the significance and reliability of our validations: “The sequence identities of the VH domain and H3 loop are 0.816 and 0.647, respectively, comparing with the best template.”.
(4) As pointed out in the first review, I think the work https://doi.org/10.1021/acs.jctc.1c00341 is worth acknowledging in section "2.2 Molecular dynamics (MD) simulations could not provide accurate CDR-H3 loop conformations" of supplementary material, as it constitutes a clear reference (and probably one of the few) to the MD simulations that authors pretend to perform. Similarly, the work https://doi.org/10.3390/molecules28103991 introduces a former benchmark on AI algorithms for predicting antibody and nanobody structures that readers may find interest to contrast with the present work. Indeed, this later reference is used by authors to answer a reviewer comment.
Thanks a lot for your valuable comments. We have added these references in the proper positions in our manuscript.







