Peer review process
Not revised: This Reviewed Preprint includes the authors’ original preprint (without revision), an eLife assessment, public reviews, and a provisional response from the authors.
Read more about eLife’s peer review process.Editors
- Reviewing EditorAnne-Florence BitbolEcole Polytechnique Federale de Lausanne (EPFL), Lausanne, Switzerland
- Senior EditorAleksandra WalczakCNRS, Paris, France
Reviewer #1 (Public Review):
Summary:
Given knowledge of the amino acid sequence and of some version of the 3D structure of two monomers that are expected to form a complex, the authors investigate whether it is possible to accurately predict which residues will be in contact in the 3D structure of the expected complex. To this effect, they train a deep learning model that takes as inputs the geometric structures of the individual monomers, per-residue features (PSSMs) extracted from MSAs for each monomer, and rich representations of the amino acid sequences computed with the pre-trained protein language models ESM-1b, MSA Transformer, and ESM-IF. Predicting inter-protein contacts in complexes is an important problem. Multimer variants of AlphaFold, such as AlphaFold-Multimer, are the current state of the art for full protein complex structure prediction, and if the three-dimensional structure of a complex can be accurately predicted then the inter-protein contacts can also be accurately determined. By contrast, the method presented here seeks state-of-the-art performance among models that have been trained end-to-end for inter-protein contact prediction.
Strengths:
The paper is carefully written and the method is very well detailed. The model works both for homodimers and heterodimers. The ablation studies convincingly demonstrate that the chosen model architecture is appropriate for the task. Various comparisons suggest that PLMGraph-Inter performs substantially better, given the same input than DeepHomo, GLINTER, CDPred, DeepHomo2, and DRN-1D2D_Inter. As a byproduct of the analysis, a potentially useful heuristic criterion for acceptable contact prediction quality is found by the authors: namely, to have at least 50% precision in the prediction of the top 50 contacts.
Weaknesses:
My biggest issue with this work is the evaluations made using *bound* monomer structures as inputs, coming from the very complexes to be predicted. Conformational changes in protein-protein association are the key element of the binding mechanism and are challenging to predict. While the GLINTER paper (Xie & Xu, 2022) is guilty of the same sin, the authors of CDPred (Guo et al., 2022) correctly only report test results obtained using predicted unbound tertiary structures as inputs to their model. Test results using experimental monomer structures in bound states can hide important limitations in the model, and thus say very little about the realistic use cases in which only the unbound structures (experimental or predicted) are available. I therefore strongly suggest reducing the importance given to the results obtained using bound structures and emphasizing instead those obtained using predicted monomer structures as inputs.
In particular, the most relevant comparison with AlphaFold-Multimer (AFM) is given in Figure S2, *not* Figure 6. Unfortunately, it substantially shrinks the proportion of structures for which AFM fails while PLMGraph-Inter performs decently. Still, it would be interesting to investigate why this occurs. One possibility would be that the predicted monomer structures are of bad quality there, and PLMGraph-Inter may be able to rely on a signal from its language model features instead. Finally, AFM multimer confidence values ("iptm + ptm") should be provided, especially in the cases in which AFM struggles.
Besides, in cases where *any* experimental structures - bound or unbound - are available and given to PLMGraph-Inter as inputs, they should also be provided to AlphaFold-Multimer (AFM) as templates. Withholding these from AFM only makes the comparison artificially unfair. Hence, a new test should be run using AFM templates, and a new version of Figure 6 should be produced. Additionally, AFM's mean precision, at least for top-50 contact prediction, should be reported so it can be compared with PLMGraph-Inter's.
It's a shame that many of the structures used in the comparison with AFM are actually in the AFM v2 training set. If there are any outside the AFM v2 training set and, ideally, not sequence- or structure-homologous to anything in the AFM v2 training set, they should be discussed and reported on separately. In addition, why not test on structures from the "Benchmark 2" or "Recent-PDB-Multimers" datasets used in the AFM paper?
It is also worth noting that the AFM v2 weights have now been outdated for a while, and better v3 weights now exist, with a training cutoff of 2021-09-30.
Another weakness in the evaluation framework: because PLMGraph-Inter uses structural inputs, it is not sufficient to make its test set non-redundant in sequence to its training set. It must also be non-redundant in structure. The Benchmark 2 dataset mentioned above is an example of a test set constructed by removing structures with homologous templates in the AF2 training set. Something similar should be done here.
Finally, the performance of DRN-1D2D for top-50 precision reported in Table 1 suggests to me that, in an ablation study, language model features alone would yield better performance than geometric features alone. So, I am puzzled why model "a" in the ablation is a "geometry-only" model and not a "LM-only" one.
Reviewer #2 (Public Review):
This work introduces PLMGraph-Inter, a new deep-learning approach for predicting inter-protein contacts, which is crucial for understanding protein-protein interactions. Despite advancements in this field, especially driven by AlphaFold, prediction accuracy and efficiency in terms of computational cost) still remains an area for improvement. PLMGraph-Inter utilizes invariant geometric graphs to integrate the features from multiple protein language models into the structural information of each subunit. When compared against other inter-protein contact prediction methods, PLMGraph-Inter shows better performance which indicates that utilizing both sequence embeddings and structural embeddings is important to achieve high-accuracy predictions with relatively smaller computational costs for the model training.
The conclusions of this paper are mostly well supported by data, but test examples should be revisited with a more strict sequence identity cutoff to avoid any potential information leakage from the training data. The main figures should be improved to make them easier to understand.
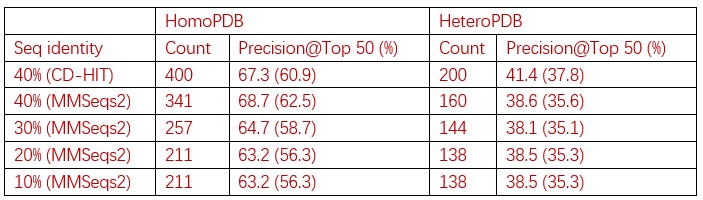
1) The sequence identity cutoff to remove redundancies between training and test set was set to 40%, which is a bit high to remove test examples having homology to training examples. For example, CDPred uses a sequence identity cutoff of 30% to strictly remove redundancies between training and test set examples. To make their results more solid, the authors should have curated test examples with lower sequence identity cutoffs, or have provided the performance changes against sequence identities to the closest training examples.
2) Figures with head-to-head comparison scatter plots are hard to understand as scatter plots because too many different methods are abstracted into a single plot with multiple colors. It would be better to provide individual head-to-head scatter plots as supplementary figures, not in the main figure.
3) The authors claim that PLMGraph-Inter is complementary to AlphaFold-multimer as it shows better precision for the cases where AlphaFold-multimer fails. To strengthen the point, the qualities of predicted complex structures via protein-protein docking with predicted contacts as restraints should have been compared to those of AlphaFold-multimer structures.
4) It would be interesting to further analyze whether there is a difference in prediction performance depending on the depth of multiple sequence alignment or the type of complex (antigen-antibody, enzyme-substrates, single species PPI, multiple species PPI, etc).