Peer review process
Revised: This Reviewed Preprint has been revised by the authors in response to the previous round of peer review; the eLife assessment and the public reviews have been updated where necessary by the editors and peer reviewers.
Read more about eLife’s peer review process.Editors
- Reviewing EditorNicolas LehrbachFred Hutchinson Cancer Research Center, United States of America
- Senior EditorJonathan CooperFred Hutch Cancer Center, Seattle, United States of America
Reviewer #2 (Public Review):
Summary
In this work, Bartolome and colleagues develop a new approach to identify proteasome interacting proteins and substrates. The approach is based on fusing proteasome subunits with a biotin ligase that will label proteins that come in close physical distance of the ligase. These biotin-labeled proteins (or their resulting tryptic peptides) can be affinity purified using streptavidin and identified by mass spectrometry.
This elegant solution was able to identify a large proportion of known proteasome interactors, as well as multiple potential new interactors. Combining this approach with a proteasome inhibitor allowed also for the enrichment of substrates, due to increased contact time between substrates and the proteasome. Again, the authors were able to identify novel substrates. Finally, the authors implemented this strategy in vivo, providing the hints for potential tissue-specific proteasome interactors.
This novel strategy provides an additional approach to identify new proteasome substrates, which can be particularly powerful for low abundant proteins, e.g., transcription factors. The possibility to implement it in vivo in specific cell types opens the possibility for identifying proteasome interactors in small cell subpopulations or in subpopulations involved in disease.
Strengths
The authors carefully characterized their genetically engineered proteasome-biotin ligase fusions to ensure that proteasome structure and activity was not altered. This is key to ensure that the proteins identified to interact with the proteasome reflect interactions that occur under physiological conditions.
The authors implemented an algorithm that controls the false positive rate of the identified interactors of the proteasome. This is an important aspect to avoid spending time on the characterization of potential interactors that are just an artifact of the experimental setup.
The addition of a proteasome inhibitor allowed the authors to identify substrates of the proteasome. Although there are other strategies to do this (e.g., affinity purification of Gly-Gly modified peptides, which is a marker for ubiquitination), this additional approach can highlight currently unknown substrates. One example are low abundance proteins, such as transcription factors.
The overall strategy developed by the authors can be implemented in vivo, which opens for the possibility of determining cell type-specific proteasome interactors (and perhaps substrates).
Weaknesses
There is a proportion (approximately 38%) of the PSMA4-biotin ligase fusion that remains unassembled (i.e., not part of the functional proteasome) and that can contribute to a small proportion of false positive interactions.
Reviewer #3 (Public Review):
Summary:
Bartolome et al. present ProteasomeID, a novel method to identify components, interactors, and (potentially) substrates of the proteasome in cell lines and mouse models. As a major protein degradation machine that is highly conserved across eukaryotes, the proteasome has historically been assumed to be relatively homogeneous across biological scales (with few notable exceptions, e.g., immunoproteasomes and thymoproteasomes). However, a growing body of evidence suggests that there is some degree of heterogeneity in the composition of proteasomes across cell tissues, and can be highly dynamic in response to physiologic and pathologic stimuli. This work provides a methodological framework for investigating such sources of variation. The authors start by adapting the increasingly popular biotin ligation strategy for labelling proteins coming into close proximity with one of three different subunits of the proteasome, before proceeding with PSMA4 for further development and analysis based on their preliminary labelling data. In a series of well-constructed and convincing validation experiments, the authors go on to show that the tagged PSMA4 construct can be incorporated into functional proteasomes, and is able to label a broad set of known proteasome components and interacting proteins in HEK293T cells. They also attempt to identify novel proteasomal degradation substrates with ProteasomeID; while this was convincing for known substrates with particularly short half-lives, the results for substrates with longer half-lives were less clear. One of the most compelling results was from a similar experiment to confirm proteasomal degradation induced by a BRD-targeting PROTAC, which I think is likely to be of keen interest to the targeted degradation community. Finally, the authors establish a ProteasomeID mouse model, and demonstrate its utility across several tissues.
Strengths:
(1) ProteasomeID itself is an important step forward for researchers with an interest in protein turnover across biological scales (e.g., in sub-cellular compartments, in cells, in tissues, and whole organisms). I especially see interest from two communities: those studying fundamental proteostasis in physiological and pathologic processes (e.g., ageing; tissue-specific protein aggregation diseases), and those developing targeted protein degradation modalities (e.g., PROTACs; molecular glues). All the datasets generated and deposited here are likely to provide a rich resource to both. The HEK293T cell line data are a valuable proof-of-concept to allow expansion into more biologically-relevant cell culture settings; however, I envision the greatest innovation here to be the mouse model. For example, in the targeted protein degradation space, two major hurdles in early-stage pre-clinical development are (i) evaluation of degradation efficacy across disease-relevant tissues, and (ii) toxicity and safety implications caused by off-target degradation, e.g., of newly-identified molecular glues and/or in particularly-sensitive tissues. The ProteasomeID mouse allows early in vivo assessment of both these questions. The results of the BRD PROTAC experiment in 293T cells provides an excellent in vitro proof-of-concept for this approach.
(2) The mass-spectrometry-based proteomics workflows used and presented throughout the manuscript are robust, rigorous, and convincing. For example, the algorithm the authors use for defining enrichment score cut-offs are logical and based on rational models, rather than on arbitrary cut-offs that are common for similar proteomics studies. The construction (and subsequent validation) of both BirA*- and miniTurbo- tagged PSMA4 variants also increases the utility of the method, allowing researchers to choose the variant with the labelling time-scale required for their particular research question.
(3) The optimised BioID and TurboID protocol the authors develop (summarised in Fig. S2A) and validate (Fig. S2B-D) is likely to be of broad interest to cell and molecular biologists beyond the protein degradation field, given that proximity labelling is a current gold-standard in global protein:protein interaction profiling.
Limitations:
I think the authors do an excellent job in highlighting the limitations of ProteasomeID throughout the Results and Discussion. I do have some specific comments that might provide additional context for the reader.
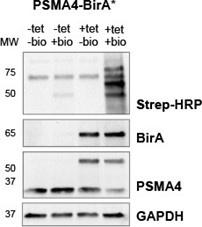
(1) The authors do a good job in showing that a substantial proportion of PSMA4-BirA* is incorporated into functional proteasome particles; however, it is not immediately clear to me how much background (false-positive IDs) might be contributed by the ~40 % of PSMA4-BirA* that is not incorporated into the mature core particle (based on the BirA* SEC-MS traces in Fig. 2b and S3b, i.e., the large peak ~ fraction 20). Are there any bands lower down in the native gel shown in Fig. 2c, i.e., corresponding to lower molecular weight complexes or monomeric PSMA4-BirA*? The enrichment of proteasome assembly factors in all the ProteasomeID experiments might suggest the presence of assembly intermediates, which might themselves become substrates for proteasomal degradation (as has been shown for other incompletely-assembled protein complexes, e.g., the ribosome, TRiC/CCT).
(2) Although the authors attempt to show that BirA* tagging of PSMA4 does not interfere with proteasome activity (Fig. 2e-f), I think the experimental evidence for this is incomplete. They show that the overall chymotrypsin-like activity (attributable to PSMB5) in cells expressing PSMA4-BirA* is not markedly reduced compared with control BirA*-expressing cells. However, they do not show that the activity of the specific proteasome sub-population that contains PSMA4-BirA* is unaffected (e.g., by purifying this sub-population via the Flag tag). The proteasome activity of the sub-population of wild-type proteasome complexes that do not contain the PSMA4-BirA* (~50%, based on the earlier immunoblots) could account for the entire chymotrypsin-like activity-especially in the context of HEK293T cells, where steady-state proteasome levels are unlikely to be limiting. It would also be useful to assess any changes in tryspin- and caspase- like activities, especially as tagging of PSMA4 could conceivably interfere with the activity of some PSMB subunits, but not others.
(3) I was left slightly unsure as to the general utility of ProteasomeID for identifying novel proteasomal substrates in homeostatic conditions--especially for proteins with longer half-lives. The cycloheximide chases in Fig. 4g/S4j are clear for MYC and TIGD5 (which have short half-lives), but are not so clear for ARMC6 and BRAT1: the reduction in the bands are modest, and might have been clearer with longer "chase" time-points. Furthermore, classifying candidates based on enrichment following proteasome inhibition with MG-132 have the potential to lead to a high number of false positives. ProteasomeID's utility in identifying potential substrates in more targeted settings (e.g., molecular glues, off-target PROTAC substrates) is far more apparent.