Peer review process
Revised: This Reviewed Preprint has been revised by the authors in response to the previous round of peer review; the eLife assessment and the public reviews have been updated where necessary by the editors and peer reviewers.
Read more about eLife’s peer review process.Editors
- Reviewing EditorGenevieve KonopkaDavid Geffen School of Medicine at UCLA, Los Angeles, United States of America
- Senior EditorSacha NelsonBrandeis University, Waltham, United States of America
Reviewer #1 (Public Review):
Summary:
This is an interesting study by Xu et al showing the effects of infection with the Treponema pallidum virus (which causes syphilis disease) on neuronal development using iPSC-derived human brain organoids as a model and single-cell RNA sequencing. This work provides an important insight into the impact of the virus on human development, bridging the gap between the phenomena observed in studies using animal models as well as non-invasive human studies showing developmental abnormalities in fetuses infected with the virus in utero through maternal vertical transmission.
Using single-cell RNAseq in combination with qPCR and immunofluorescence techniques, the authors show that T. pallidum infected organoids are smaller in size, in particular during later growth stages, contain a larger number of undifferentiated neuronal lineage cells, and exhibit decreased numbers of specific neuronal subcluster, which the authors have identified as undifferentiated hindbrain neurons.
The study is an important first step in understanding how T. pallidum affects human neuronal development and provides important insight into the potential mechanisms that underlie the neurodevelopmental abnormalities observed in infected human fetuses.
Strengths:
(1) The study is well written, and the data quality is good for the most part.
(2) The study provides an important first step in utilizing human brain organoids to study the impact of T. pallidum infection on neuronal development.
(3) The study's conclusions may provide important insight to other researchers focused on studying how viral infections impact neuronal development.
Reviewer #3 (Public Review):
This article is the first report to study the effects of T. pallidum on the neural development of an iSPC-derived brain organoid model. The study indicates that T. pallidum inhibits the differentiation of subNPC1B neurons into hindbrain neurons, hence affecting brain organoid neurodevelopment. Additionally, the TCF3 and notch signaling pathways may be involved in the inhibition of the subNPC1B-hindbrain neuron differentiation axis. While the majority of the data in this study support the conclusions, there are still some questions that need to be addressed and data quality needs to be improved. The study provides valuable insights for future investigations into the mechanisms underlying congenital neurodevelopment disability.
Author response:
The following is the authors’ response to the original reviews.
eLife assessment
This is a valuable study that describes the effects of T. pallidum on neural development by applying single-cell RNA sequencing to an iPSC-derived brain organoid model. The evidence supporting the claims of the authors is solid, although further evidence to understand the differences in infection rates would strengthen the conclusions of the study. In particular, the conclusions would be strengthened by validating infection efficiency as this can impact the interpretation of single-cell sequencing results, and how these metrics affect organoid size as well as comparison with additional infectious agents. Furthermore, additional validations of downstream effectors are not adequate and could be improved.
Thank you very much for your valuable comments. Since we used the organoid model for the first time to investigate the effects of T. pallidum on brain development, the study design is not perfect. As you have accurately mentioned, the results of the paper do not have more in-depth details, especially to verify the infection rate of T. pallidum. Your valuable comments will be very useful for us for carrying out further research. In addition, the downstream effector validation is inadequate, so we performed an analysis of single-cell sequencing data to strengthen our view in the revised manuscript (See Figure 5F for a description in current manuscript).
Public Reviews:
Reviewer #1 (Public Review):
Summary:
This is an interesting study by Xu et al showing the effects of infection with the Treponema pallidum virus (which causes syphilis disease) on neuronal development using iPSC-derived human brain organoids as a model and single-cell RNA sequencing. This work provides an important insight into the impact of the virus on human development, bridging the gap between the phenomena observed in studies using animal models as well as non-invasive human studies showing developmental abnormalities in fetuses infected with the virus in utero through maternal vertical transmission.
Using single-cell RNAseq in combination with qPCR and immunofluorescence techniques, the authors show that T. pallidum infected organoids are smaller in size, in particular during later growth stages, contain a larger number of undifferentiated neuronal lineage cells, and exhibit decreased numbers of specific neuronal subcluster, which the authors have identified as undifferentiated hindbrain neurons.
The study is an important first step in understanding how T. pallidum affects human neuronal development and provides important insight into the potential mechanisms that underlie the neurodevelopmental abnormalities observed in infected human fetuses. Several important weaknesses have also been noted, which need to be addressed to strengthen the study's conclusions.
Strengths:
(1) The study is well written, and the data quality is good for the most part.
(2) The study provides an important first step in utilizing human brain organoids to study the impact of T. pallidum infection on neuronal development.
(3) The study's conclusions may provide important insight to other researchers focused on studying how viral infections impact neuronal development.
Thank you very much for your positive feedback. Below, you will find our detailed responses to your concerns, addressed point-by-point. I once again sincerely appreciate your time and effort in reviewing our manuscript.
Weaknesses:
(1) It is unclear how T. pallidum infection was validated in the organoids. If not all cells are infected, this could have important implications for the study's conclusions, in particular the single-cell RNAseq experiments. Were only cells showing the presence of the virus selected for sequencing? A detailed description of how infection was validated and the process of selection of cells for RNAseq would strongly support the study's conclusions.
Thank you for your valuable comment. We completely agree with your point. Exploring the infection rate of T. pallidum to brain organoids is a key factor that must be considered. We selected pluripotent stem cell-derived brain organoids to simulate the process of foetal brain neurodevelopment and cultured them mixed with T. pallidum to mimic T. pallidum invading brain tissue. Since brain organoids are three-dimensional structures formed by nerve cell aggregation, T. pallidum invades organoids from the periphery to the center of the organoids gradually. T. pallidum acts on organoids long enough to increase the infection rates; however, the pathogen is selective in invading human cells. If we only select cells present in T. pallidum for sequencing, the authenticity of simulating "real world" infections is somewhat weakened. To better carry out this study, selecting cells from intact organoids for sequencing, without eliminating cells without T. pallidum, can better simulate the effect of T. pallidum infection on the nervous system. Of course, we should also set up a blank control group.
(2) The authors show that T. pallidum infection results in impaired development of hindbrain neurons. How does this finding compare to what has already been shown in animal studies? Is a similar deficit in this brain region observed with this specific virus? It would be useful to strengthen the study's conclusions if the authors added a discussion about the observed deficits in hindbrain neuronal development, and prior literature on similar studies conducted in animal models or human patients. Does T. pallidum preferentially target these neurons, or is this a limitation of the current organoid model system?
Thank you for your valuable comments. The finding that T. pallidum infection results in impaired development of hindbrain neurons has not been verified in animal experiments. Of course, it is better to further validate the findings in organoid studies through animal experiments. Unfortunately, due to the technical challenges, mature animal models have not been developed for the study of congenital syphilis. Although our team has been working on the development of animal models of congenital neurosyphilis, the current progress is still not satisfactory. After struggling hard in this field for many years, we decided to attempt to utilize human brain organoids instead of animal models to study the impact of T. pallidum infection on neuronal development.
We also checked prior literature on similar studies that have referred to the content in human patients. Dan Doherty et al. reported that patients with pontocerebellar hypoplasia develop microcephaly at birth or over time after birth (PMID: 23518331). Based on your constructive suggestions, we have added some content related to hindbrain to the “Discussion” section.
Our study found that T. pallidum could inhibit the differentiation of subNPC1B in brain organoids, thereby reducing the differentiation from subNPC1B to hindbrain neurons, and ultimately affecting the development and maturation of hindbrain neurons during pregnancy. Based on our results, T. pallidum does not preferentially target hindbrain neurons. Of course, there are limitations to the current organoid model system, see the "Limitations" section.
PMID: 23518331- Dan Doherty et al, Midbrain and hindbrain malformations: advances in clinical diagnosis, imaging, and genetics.
Revision in the “Discussion” section, line 343-352:
“The vertebrate hindbrain contains a complex network of dedicated neural circuits that play an essential role in controlling many physiological processes and behaviors, including those related to the cerebellum, pons, and medulla oblongata (Shoja et al., 2018). Patients with pontocerebellar hypoplasia represent the less severe end of the spectrum with early hyperreflexia, developmental delay, and feeding problems, eventually developing spasticity and involuntary movements in childhood, while some patients represent the severe end of the spectrum characterised by polyhydramnios, severe hyperreflexia, contracture, and early death from central respiratory failure. Patients with pontocerebellar hypoplasia develop microcephaly at birth or over time after birth (Doherty et al., 2013).”
(3) The authors show that T. pallidum-infected organoids are smaller in size by measuring organoid diameter during later stages of organoid growth, with no change during early stages. Does that represent insufficient infection at the early stages? Is this due to increased cell death or lack of cell division in the infected organoids? Experiments using IHC to quantify levels of cleaved caspase and/or protein markers for cell proliferation would be able to address these questions.
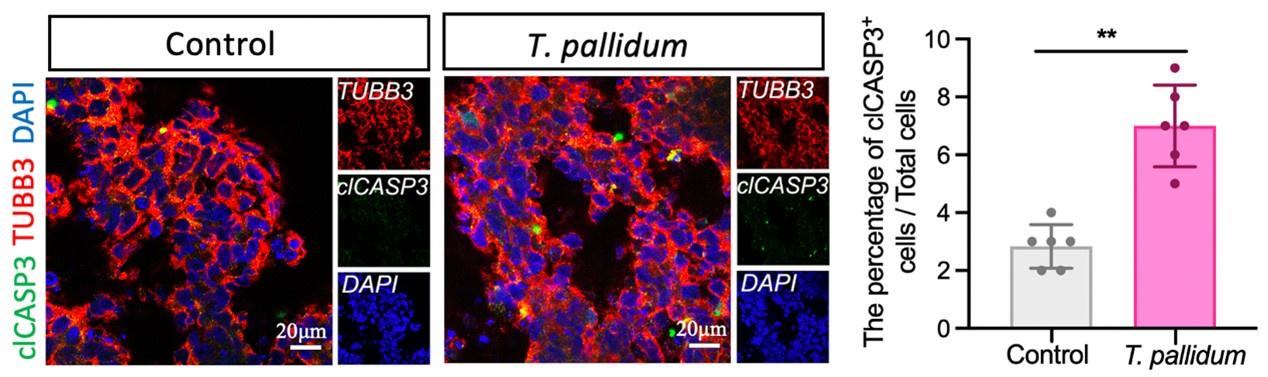
Thank you for your valuable suggestion. The concentration of T. pallidum in patients with syphilis was generally very low (PMID: 21752804, 35315702, 33099614). In this study, a low concentration of T. pallidum was applied to brain organoids to simulate early foetal transmission of syphilis. Nerve cells mainly establish intercellular connections to form brain organoids in the way of adhesion, which can easily cause organoids to divide and die if treated with a high concentration of T. pallidum. Furthermore, based on your suggestions, we performed additional immunostaining analyses to verify the apoptosis of brain organoids infected by T. pallidum. Cleaved caspase 3 (clCASP3) staining showed that the number of apoptotic cells increased following T. pallidum infection; however, the proportion of apoptotic cells in both groups of brain organoids was very low (Figure supplement 2) (N=12 organoids, each group from three independent bioreactors), which would be not enough to affect the results of the experiment, thereby suggesting that neural differentiation and development of brain organoids were mainly inhibited following T. pallidum infection (rather than promoting organoid apoptosis).
PMID: 21752804-- Craig Tipple et al, Getting the measure of syphilis: qPCR to better understand early infection.
PMID: 35315702-- Cuini Wang et al, Quantified Detection of Treponema pallidum DNA by PCR Assays in Urine and Plasma of Syphilis Patients.
PMID: 33099614—Cuini Wang et al, A New Specimen for Syphilis Diagnosis: Evidence by High Loads of Treponema pallidum DNA in Saliva.
Revision in the “Results” section, line 105-108:
“… cleaved caspase 3 (clCASP3) staining showed that the number of apoptotic cells increased significantly following T. pallidum infection, but the proportion of apoptotic cells in both groups of brain organoids was very low (Figure supplement 2) (N=12 organoids, each group from three independent bioreactors) …”
Revision in the “Materials and methods” section, line 446-447:
“…anti-cleaved caspase 3 (rabbit, 1:100, Cell Signaling Technology, 9661S),”
Revision in the “Supplementary File” section, line 78-81:
Author response image 1.
The number of clCASP3+ cells in the microscopic field of brain organoids. A nonparametric t-test was used to evaluate the statistical differences between the two groups. (**: P < 0.01).
(4) In Figure 1D authors show differences in rosette-like structure in the infected organoids. The representative images do not appear to be different in any of the discussed components (e.g., the sox2 signal looks fairly similar between the two conditions). No quantification of these structures was presented. Authors should provide quantification or a more representative image to support their statement.
Thank you for your valuable suggestion. I have quantified the neural rosette structure and compared the number of intact rosette-like structures between the two groups (See Figure 1D for a description in current manuscript).
(5) The IHC images shown in Figures 3E, G, and Figure 4E look very similar between the two conditions despite the discussed decrease in the text. A more suitable representative image should be presented, or the analysis should be amended to reflect the observed results.
Thank you for your valuable suggestion. I have replaced more representative images in Figure 3E, G, and Figure 4E in the manuscript.
Reviewer #2 (Public Review):
Summary:
This study provides an important overview of infectious etiology for neurodevelopment delay.
Strengths:
Strong RNA evaluation.
Weaknesses:
The study lacks an overview of other infectious agents. The study should address the epigenetic contributors (PMID: 36507115) and the role of supplements in improving outcomes (PMID: 27705610).
Addressing the above - with references included - is recommended.
Thank you for your valuable comment. Our research is mainly inspired by other infectious agents, such as Zika virus; there are many descriptions of Zika virus in the “Discussion” section of the manuscript to better describe and demonstrate our point of view (See pages 12–13). I was unable to retrieve the article (PMID: 36507115), kindly help in confirming the PMID number. I will be very grateful if you can provide the full text. Secondly, I have carefully read the article (PMID: 27705610), which is a very rich and comprehensive review, and summarised and cited it in appropriate places in our manuscript.
Revision in the “Discussion- limitation” section, line 375-379:
“First, although several recent protocols have made use of growth factors to promote further neuronal maturation and survival (Lucke-Wold et al., 2018), the organoid culture scheme needs to be further improved owing to the lower percentage of mature neurons and the challenge of cell necrosis within the organoids at this stage in day 55 organoids.”
Reviewer #3 (Public Review):
This article is the first report to study the effects of T. pallidum on the neural development of an iPSC-derived brain organoid model. The study indicates that T. pallidum inhibits the differentiation of subNPC1B neurons into hindbrain neurons, hence affecting brain organoid neurodevelopment. Additionally, the TCF3 and notch signaling pathways may be involved in the inhibition of the subNPC1B-hindbrain neuron differentiation axis. While the majority of the data in this study support the conclusions, there are still some questions that need to be addressed and data quality needs to be improved. The study provides valuable insights for future investigations into the mechanisms underlying congenital neurodevelopment disability.
I sincerely appreciate your comments on our paper. The comments have helped us greatly improve the quality of our paper. Thank you for your time and constructive critique.
Recommendations for the authors:
Reviewer #1 (Recommendations For The Authors):
Paired t-test analysis is not appropriate if two distinct groups are compared.
I sincerely apologize for our presentation. We used a nonparametric t-test to compare the two groups. I have confirmed and corrected the statistical method description of this manuscript (Revision in the “Materials and methods” section (line 553-555) and “Figures-legend” section (line 789-790, 817-818, 829-830) in current manuscript).
Reviewer #3 (Recommendations For The Authors):
(1) Can the authors explain why the mean size of organoids infected with T. pallidum is smaller?
Thank you for your valuable comment. In our study, T. pallidum infection resulted in brain organisational changes in neural rosette-like structures resembling the proliferative regions of the human ventricular zone and caused fewer and incomplete rosette-like structures. Next, the ventricular zone is also the main area where neural progenitor cells (NPCs) reside (PMID: 33838105); our results showed that the proportion of neural progenitor cells (NPC)1 was reduced after T. pallidum infection. Rosette-like structure size changes owing to NPC depletion. Therefore, the mean size of organoids infected with T. pallidum is smaller.
Revision in the “Results” section, line 101-104:
“T. pallidum infection resulted in brain organisational changes in neural rosette-like structures resembling the proliferative regions of the human ventricular zone where NPC reside (Krenn et al., 2021), and caused fewer and incomplete rosette-like structures (P < 0.01) (Figure 1D)”
(2) Why was the target gene for qRT-PCR validation selected to be HOXA5、HOXC5、HOXA4?
Thank you for your valuable comment. The qRT-PCR experiment was selected here to verify the analysis results of the scRNA-seq. HOX family genes are key factors controlling early hindbrain development, which are expressed in the hindbrain region during the gastrulation stage of early embryonic development and persist into the nerve cell stage, and are essential for the correct induction of hindbrain development and segmentation (PMID: 2571936, 1983472, 1673098, 15930115). Therefore, we selected the HOX family gene for verification.
PMID: 2571936-WILKINSON D G, et al. Segmental expression of Hox-2 homoeobox- containing genes in the developing mouse hindbrain.
PMID: 1983472-- FROHMAN M A, et al. Isolation of the mouse Hox-2.9 gene; analysis of embryonic expression suggests that positional information along the anterior-posterior axis is specified by mesoderm.
PMID: 1673098--MURPHY P, et al. Expression of the mouse labial-like homeobox-containing genes, Hox 2.9 and Hox 1.6, during segmentation of the hindbrain.
PMID: 15930115-- MCNULTY C L, et al. Knockdown of the complete Hox paralogous group 1 leads to dramatic hindbrain and neural crest defects.
(3) Why was qRT-PCR not employed in other experimental validations, but solely to validate early neural-specific transcription factor changes?
Thank you for your valuable comment. The qRT-PCR experiment was selected to validate early neural-specific transcription factor changes, indicating the reliability of the scRNA-seq. Then, validated scRNA-seq data were used to analyze for other neuro-specific gene differences, such as violin plots and heatmap showing differentially expressed genes (Figure 4D and Figure 5B, C). Of course, we also tested it with other experiments, such as immunohistochemistry and flow cytometric screening.
(4) The authors found that T. pallidum might reduce the differentiation from subNPC1B to hindbrain neurons by inhibiting subNPC1B differentiation in brain organoids. Why were the subNPC1B-specific markers declining?
Thank you for your valuable comment. scRNA-seq is aimed at complete brain organoids. Cluster analysis of cell types of organoids is performed according to specific marker genes of different cells. The decrease in the expression of marker genes of certain cell groups indicates that the cell proportion of such cell groups in the whole organoids is reduced. We analysed organoids following T. pallidum infection, uniform manifold approximation and projection (UMAP), and clustering of the NPC1 population demonstrated that T. pallidum downregulated the number of subNPC1B population. Therefore, the results demonstrated a decrease in the subNPC1B -specific markers.
(5) In comparison to the other figures, Figure 5E letter size is excessively small and ambiguous.
Thanks for your valuable comments, I have adjusted Figure 5E letter size.
(6) Figure 5E shows that TCF3, more than one gene, is specifically enriched in subNPC1B of the T. pallidum group. It is best to confirm the impact of the other gene.
Thank you for raising this key issue that we had not addressed properly in our previous version of the manuscript; we have added further analytical data. The SCENIC analysis found that the transcriptional activity of 52 genes has significantly changed after T. pallidum infection. Furthermore, GO analyses demonstrated that 27 transcription factors were significantly enriched in four key pathways of neural differentiation and development. TCF3 is the sole transcription factor present in all four terms simultaneously, speculating that TCF3 is the key transcription factor for the inhibition of subNPC1B-hindbrain neuron differentiation caused by T. pallidum.
Revision in the “Results” section, line 261-273:
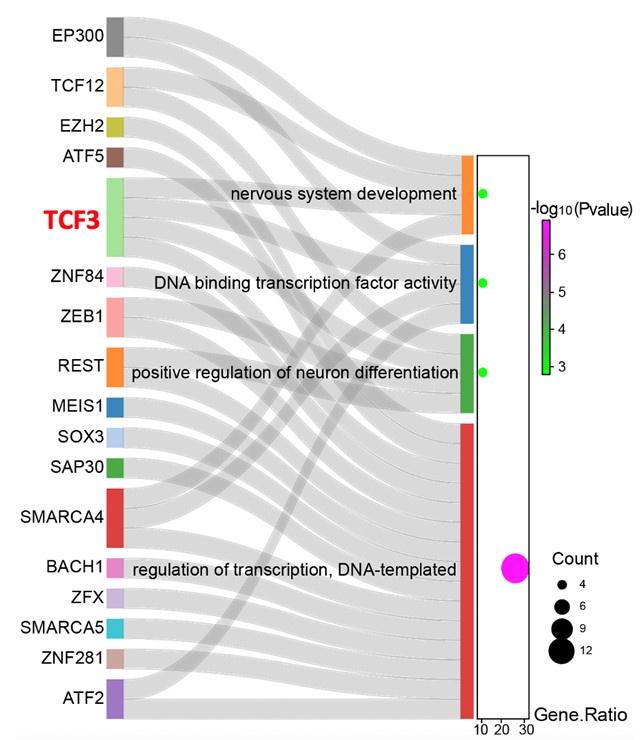
“Next, the single-cell regulatory network inference and clustering (SCENIC) analysis for the subNPC1B subcluster was performed to assess the differences in the transcriptional activity of the transcription factors between the two groups and found that the transcriptional activity of 52 genes significantly changed after T. pallidum infection (Figure 5E). Furthermore, GO analyses demonstrated that 27 transcription factors were significantly enriched in key pathways of neural differentiation and development in response to nervous system development, positive regulation of sequence-specific DNA-binding transcription factor activity, positive regulation of neuronal differentiation, and DNA templated transcription regulation. Remarkably, transcription factor 3 (TCF3) is the sole transcription factor present in all four terms simultaneously (Figure 5F), speculating that TCF3 is the key transcription factor for the inhibition of subNPC1B-hindbrain neuron differentiation caused by T. pallidum.”
Revision in the “Materials and methods” section, line 540-543:
“The Sankey diagram was created using SankeyMATIC (https://sankeymatic.com/) (Zhang et al., 2023), which was used to characterize the interactions between differential transcription factors and neural differentiation and development.”
Revision in the “Figure and Figure Legend” section, line 832, 842-844:
Author response image 2.
Sankey diagram showing the correspondence between differential transcription factors and neural differentiation and development.
(7) Are there other experiments demonstrating that TCF3 is a key transcription factor for the inhibition of subNPC1B-hindbrain neuron differentiation caused by T. pallidum?
Thank you for your valuable comment. In the previous experiment, we attempted to select a subNPC1B subcluster by flow sorting to verify the relevant molecular mechanism. Due to the small proportion of subNPC1B subcluster in the whole organoids, the selected cells were in a poor state and could not reach the number of cells required for the experiment. However, we used scRNA-seq data to further identify TCF3 as a key transcription factor that inhibits subNPC1B - hindbrain neuron differentiation induced by T. pallidum. The relevant results and descriptions of the analysis are detailed in the revised manuscript, please see our response to point (6) above.