DNA-PK is a DNA sensor for IRF-3-dependent innate immunity
Figures
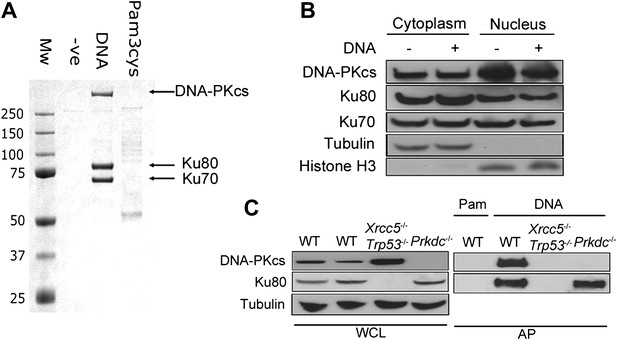
DNA-PK binds DNA in the cytoplasm.
(A) Biotinylated DNA or biotinylated pam-3-cys were transfected into HEK 293T cells and 30 min later DNA was affinity purified from the cytoplasmic fraction before analysis of DNA/protein complexes by SDS-PAGE. The three major protein bands visible by coomassie blue staining were excised from the gel and identified as Ku70, Ku80 and DNA-PKcs by mass spectrometry. (B) Cells were untreated or transfected with DNA and 2 hr later proteins were extracted from the nucleus and cytoplasm. 50 μg of protein from each fraction (representing 10% of total cytoplasmic protein by volume and 5% of the total nuclear protein) was analysed by immunoblotting for DNA-PKcs, Ku70 and Ku80. β-tubulin and histone-H3 were used as controls to indicate successful fractionation. (C) Ku80 is required for efficient binding to DNA. MEFs of indicated genotypes were transfected with DNA biotinylated at the 3′ end or biotinylated pam-3-cys (Pam) and lysed 1 hr later. Following affinity purification (AP) of biotinylated ligands with streptavidin agarose, proteins were analysed by SDS-PAGE and immunoblotting for DNA-PKcs and Ku80. WCL; whole cell lysate.
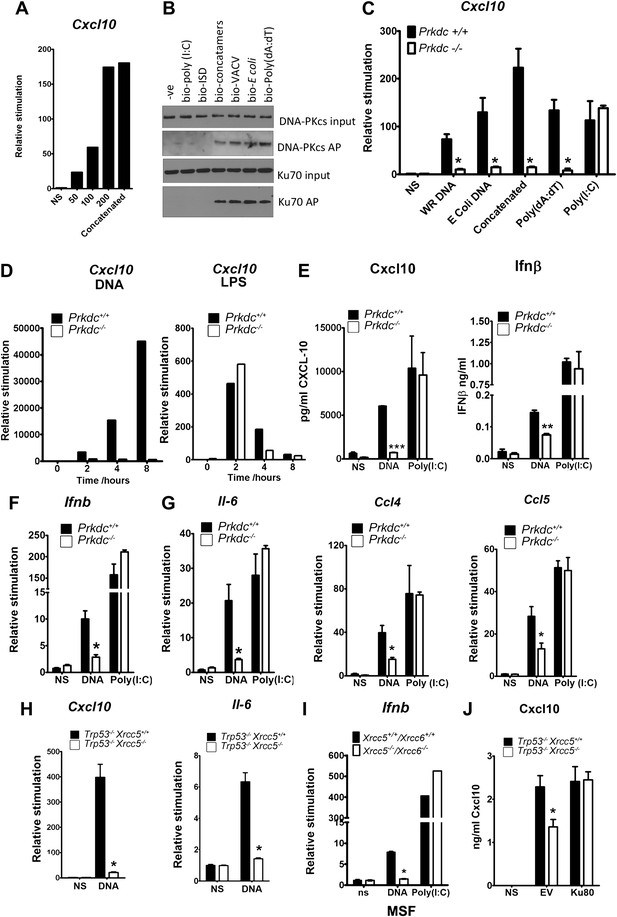
The innate immune response to DNA requires DNA-PK in fibroblasts.
(A) ISD DNA of different lengths was transfected into MEFs and the transcription of Cxcl10 was assayed by qPCR 6 hr later. (B) Double stranded oligonucleotides (bio-ISD), concatenated ISD DNA (bio-concatamers), genomic vaccinia virus DNA (bio-VACV), genomic E. coli DNA (bio-E. coli), poly (dA:dT) or the RNA analogue poly (I:C) were biotinylated and transfected into HEK293 cells. Following affinity purification of proteins from cytoplasmic extracts using streptavidin agarose, the bound proteins were analysed by SDS-PAGE and immunoblotting. AP; affinity purification. (C) Primary MEFs of the indicated genotype were transfected with 10 μg/ml of the same (non-biotinylated) nucleic acids as in (A) followed by qRT-PCR analysis measuring induction of Cxcl10 mRNA 6 hr later. (D) Wild type and Prkdc−/− transformed MEFs were transfected with DNA (10 μg/ml, left panel) or stimulated with LPS (100 ng/ml, right panel) and the level of transcription of cxcl10 was measured at the indicated times post stimulation. (E) Levels of Cxcl10 and Ifnβ were measured by ELISA from the supernatants of primary wild type and Prkdc−/− MEFs at passage 1, 24 hr after transfection with DNA or poly (I:C). (F),(G) Primary wild type and Prkdc−/− MEFs at passage 1 were transfected with DNA or poly (I:C) and the level of induction of (F) Ifnb and Il-6 and (G) ccl4 and ccl5 mRNA was measured by qRT-PCR 6 hr later. (H) MEFs expressing Ku80 or lacking Ku80 were transfected with DNA and the transcription of Cxcl10 or Il6 was measured by qPCR 6 hr later. (I) Primary murine skin fibroblasts (MSF) from wild type adult mice or those lacking both Ku genes were transfected with DNA or poly (I:C) and the level of Ifnb induction was measured 6 hr later by qRT-PCR. (J) Xrcc5+/+/Trp53−/− and Xrcc5−/−/Trp53−/− MEFs were transfected with an expression plasmid encoding Ku80 or an empty vector (EV) control and Cxcl10 production was measured 24 hr later by ELISA. *** p<0.001, ** p<0.01, * p<0.05, n ≥ 3, error bars ± SEM, ns; non-stimulated.
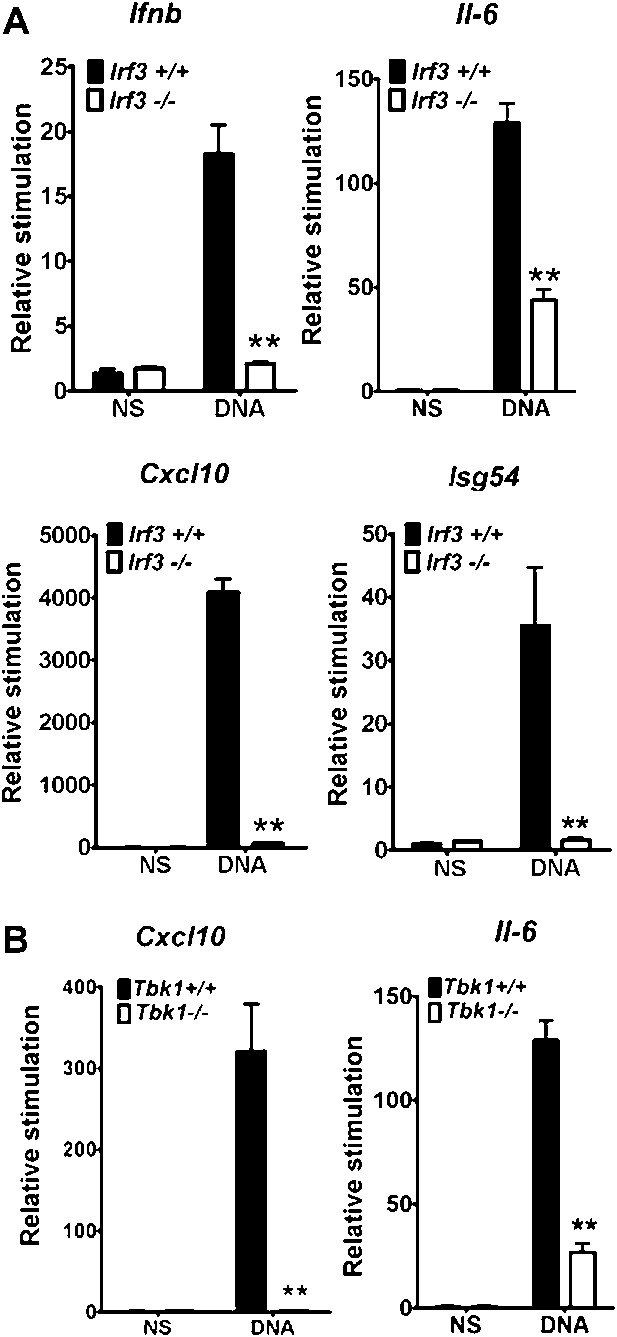
IRF-3 and TBK1 are required for the stimulation of multiple genes in response to DNA.
(A) Primary wild type or Irf3−/− MEFs were transfected with DNA and the level of induction of Ifnb, Il6, Cxcl10 and Isg54 mRNAs were measured by qRT-PCR 6 hr later. (B) Immortalised wild type or Tbk1−/− MEFs were transfected with DNA or poly(I:C) and the level of induction of Il-6 and Cxcl10 was measured by qRT-PCR 6 hr later. (B) *** p<0.001, ** p<0.01, n = 3, error bars ± SEM, ns; non-stimulated.
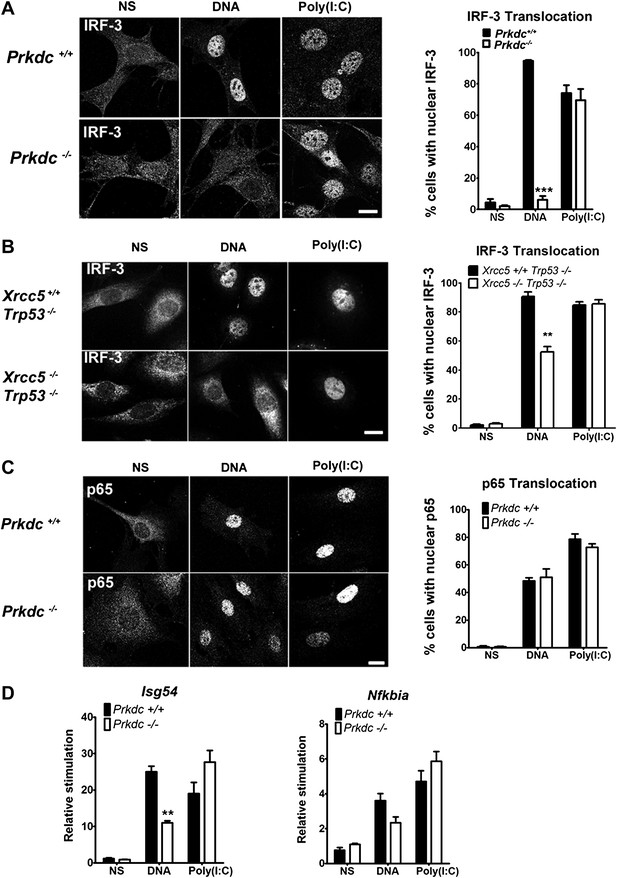
DNA-PK activates IRF-3-dependent, NF-κB–independent signalling.
(A) The localisation of endogenous IRF-3 was analysed by immunofluorescence 1 hr after transfection of primary wild type or Prkdc−/− MEFs with DNA or poly (I:C) (left panels) and quantified by scoring cells with nuclear staining (right panels, n = 3, counts of at least 50 nuclei per slide in randomised fields of view). (B) As (A) but with Xrcc5+/+/Trp53−/− and Xrcc5−/−/Trp53−/− MEFs. (C) Analysis of p65 translocation in wild type and Prkdc−/− MEFs carried out as in (A). (D) Primary wild type or Prkdc−/− MEFs were transfected with DNA or poly (I:C) and the level of induction of Isg54 and Nfkbia were measured by qRT-PCR 6 hr later. ** p<0.01, * p<0.05, n = 3, error bars ± SEM. ns; non-stimulated. Scale bar; 10 μm.
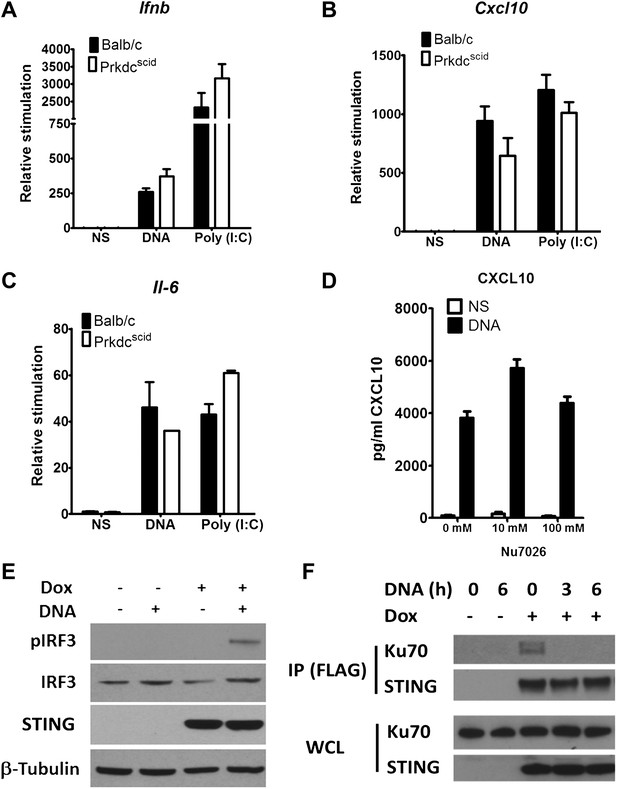
DNA-PKcs kinase activity is dispensable for the innate immune response to DNA.
Primary fibroblasts from Balb/c or PrkdcSCID mice were transfected with DNA or poly(I:C) or infected with MVA or NDV and the level of induction of (A) Ifnb, (B) Cxcl10, or (C) Il-6 was measured 6 hr later by qRT-PCR. (D) Fibroblasts were incubated with the indicated dose of DNA-PKcs kinase inhibitor, Nu7026, or carrier control and then stimulated with 10 μg/ml DNA. Cxcl10 was measured by ELISA in the supernatants 24 hr following stimulation. n = 3, error bars ± SEM. (E). Hek293 Trex cells were stably transfected with FLAG-tagged STING under the control of a doxycycline-inducible promoter. STING expression was induced by addition of doxycycline (Dox, 2 μg/ml) for 24 hr and cells were stimulated by transfection with 5 μg/ml DNA for 6 hr. Protein lysates were then immunoblotted with the indicated antibodies. (F) STING-293Trex cells were induced to express STING by addition of doxycycline (Dox, 2 μg/ml) for 24 hr and stimulated with 5 μg/ml DNA for the indicated times. STING was then immunoprecipitated and whole cell lystes (WCL) or precipitated proteins (IP) were immunoblotted using the indicated antibodies.
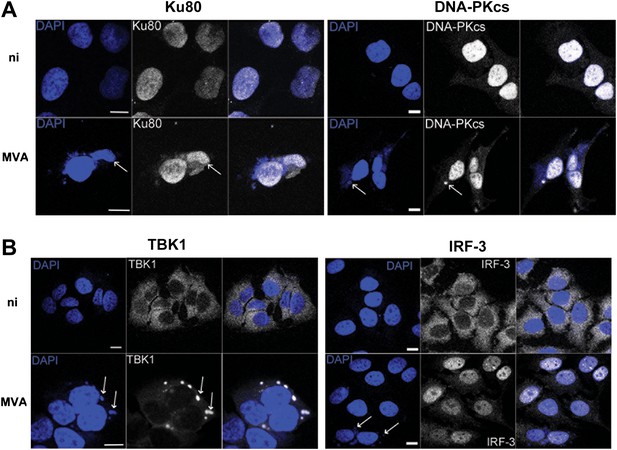
DNA-PK and TBK1 localise to sites of VACV DNA replication in infected cells.
HeLa cells were untreated or infected with MVA (m.o.i. = 5) for 6 hr. Cells were then fixed and stained with antibodies against (A) Ku80 or DNA-PKcs, and (B) TBK1 or IRF-3. Cytoplasmic viral factories formed after MVA infection are visualised with DAPI (blue). Sites of co-localisation of DNA-PKcs or Ku80 with viral factories are indicated by white arrows. Scale bars; 10 μm. ni; non-infected.
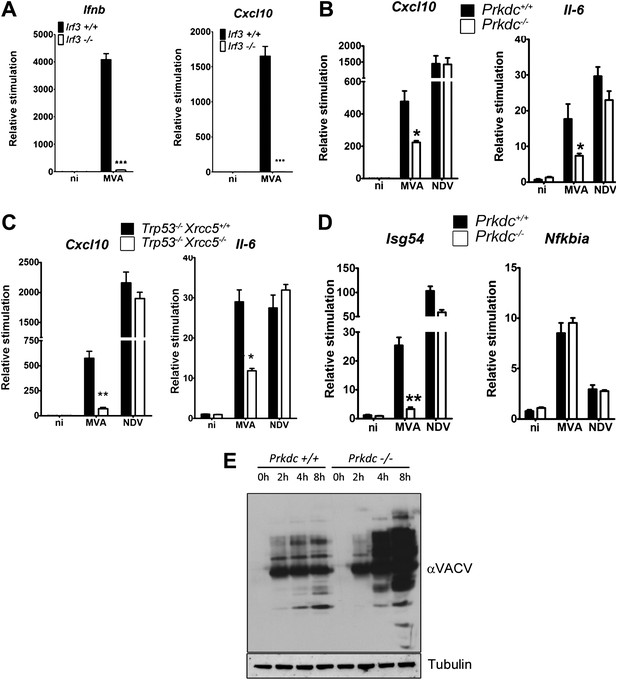
DNA-PK contributes to the IRF-3-dependent innate immune response to MVA.
(A) Ifnb and Cxcl10 transcription was measured 6 hr following MVA infection of primary WT and Irf3−/− fibroblasts at an m.o.i. of 5. (B) Wild type and Prkdc−/− MEFs were infected with MVA or NDV and the level of induction of Cxcl10 and Il-6 were measured 6 hr later by qRT-PCR. (C) As (A) but with immortalised Xrcc5+/+/Trp53−/− and Xrcc5−/−/Trp53−/− MEFs. (D) The induction of Isg54 and Nfkbia mRNA was measured by qRT-PCR 6 hr after MVA or NDV infection of wild type and Prkdc−/− cells. (E) Expression of VACV proteins, analysed by immunoblotting with a rabbit polyclonal anti-VACV serum, at the indicated times following infection of primary Prkdc+/+ and Prkdc−/− MEFs with MVA (m.o.i. = 5). *** p<0.001, ** p<0.01, * p<0.05, n = 3, error bars ± SEM, ni; non-infected.
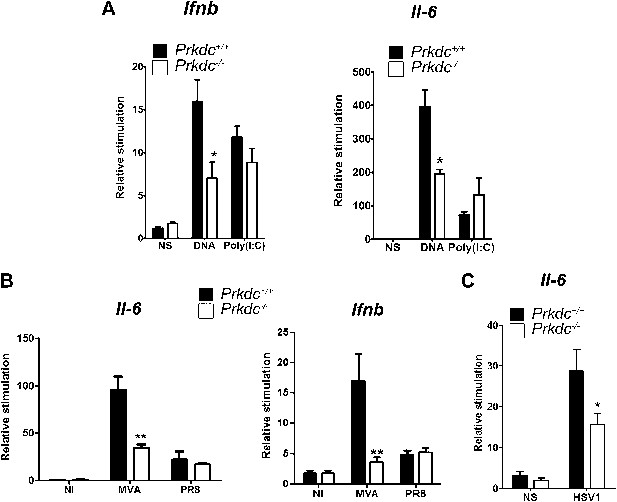
DNA-PKcs contributes to the innate immune response to MVA, HSV-1 and DNA in vivo.
(A) Groups of five Prkdc+/+ and Prkdc−/− mice were injected intradermally into the ear pinna with cationic lipids complexed with 1 μg DNA or poly(I:C). RNA was extracted from the tissue 12 hr later and Ifnb and Il6 transcription was measured by qPCR. (B) Groups of five Prkdc+/+ and Prkdc−/− mice were infected intradermally with 106 pfu of MVA or influenza virus strain A/PR/8/34 (PR8) and 12 hr later Ifnb and Il6 transcription was measured by qPCR from RNA extracted from the local site of infection. (C) As (B) but with HSV-1 strain S17. Note that levels of ifnb could not be measured above background in this experiment. *p<0.1, ** p<0.01, n = 5, error bars ± SEM, ni; non-infected.





