Non-canonical TAF complexes regulate active promoters in human embryonic stem cells
Figures
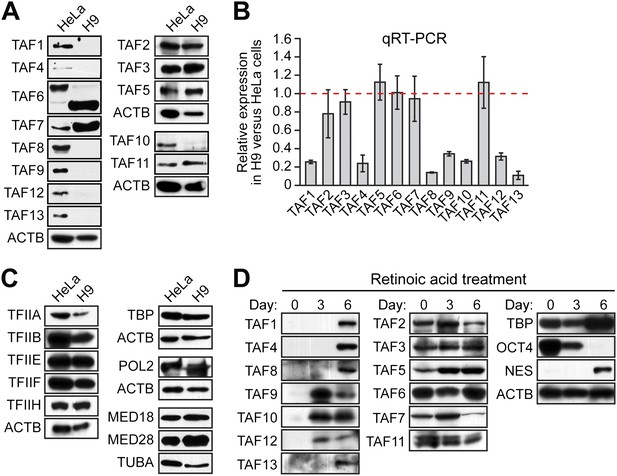
Undifferentiated hESCs express only a subset of TFIID TAFs.
(A) Immunoblot analysis showing TAF levels in HeLa cells and H9 hESCs. β-actin (ACTB) was monitored as a loading control. (B) qRT-PCR analysis monitoring TAF expression in H9 cells relative to HeLa cells. A ratio of 1 (indicated by the red dotted line) indicates no difference in expression. Data are represented as mean ± SEM. (C) Immunoblot analysis showing levels of GTFs in HeLa cells and H9 hESCs. α-tubulin (TUBA) was monitored as a loading control. (D) Immunoblots showing TAF and TBP levels in H9 hESCs following induction of differentiation by retinoic acid treatment for 0, 3 or 6 days. OCT4 and NES were monitored as controls.
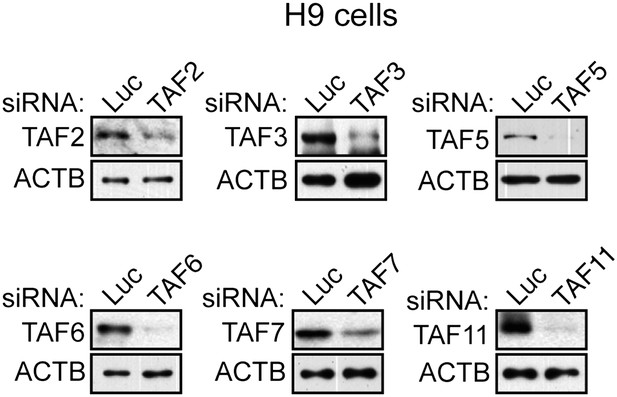
Confirmation of specificity of TAF antibodies by RNAi-mediated knockdown in H9 hESCs.
Immunoblot analysis showing TAF levels in H9 hESCs 48 hr after transfection with a control luciferase (Luc) or TAF siRNA. β-actin (ACTB) was monitored as a loading control.
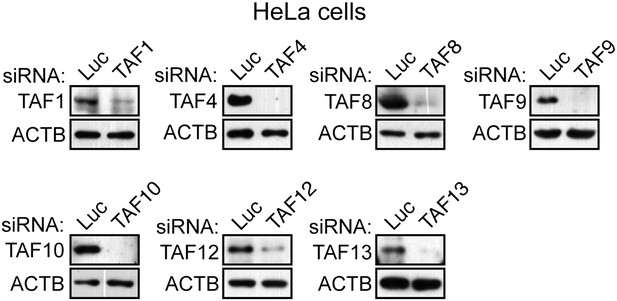
Confirmation of specificity of TAF antibodies by RNAi-mediated knockdown in HeLa cells.
Immunoblot analysis showing TAF levels in HeLa cells 48 hr after transfection with a control luciferase (Luc) or TAF siRNA. β-actin (ACTB) was monitored as a loading control.
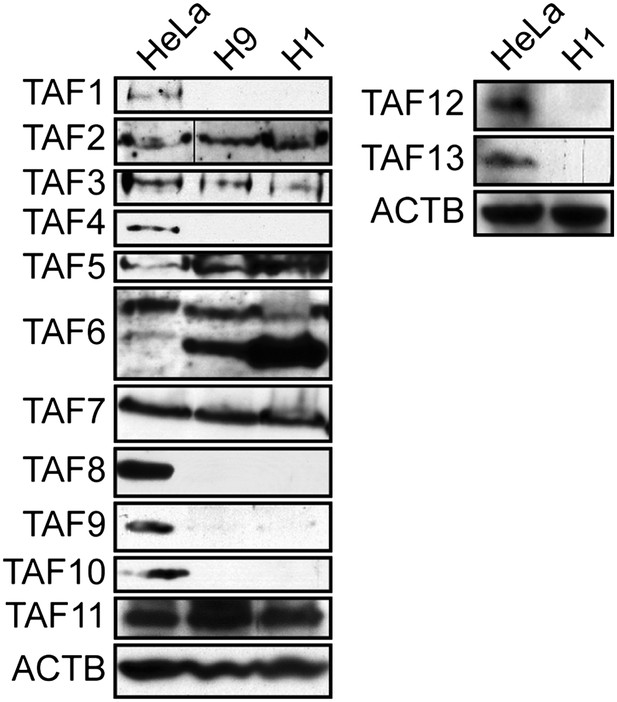
TAF expression levels in H1 hESCs.
Immunoblot analysis showing TAF levels in H1 hESCs and, as a comparison, H9 hESCs and HeLa cells. β-actin (ACTB) was monitored as a loading control. The results demonstrate an identical pattern of TAF expression in both H9 and H1 hESCs.
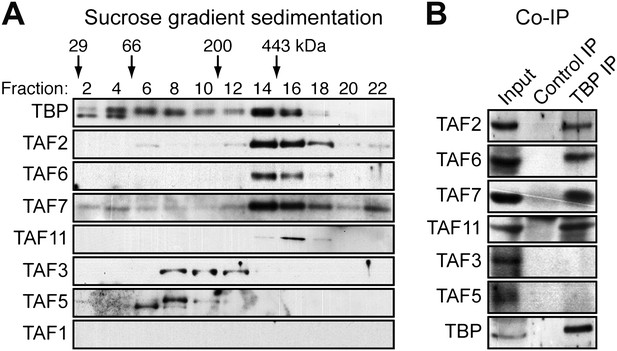
hESCs have a non-canonical TBP-containing TAF complex.
(A) Sucrose gradient sedimentation. H9 cell nuclear extract was fractionated, and individual fractions were analyzed for TAFs by immunoblotting. Arrows indicate elution peaks of protein standards. (B) Co-immunoprecipitation analysis. Nuclear extracts from H9 cells were immunoprecipitated with an anti-TBP or control (anti-RAB2A) antibody and the immunoprecipitate was analyzed for TAFs and TBP by immunoblotting.
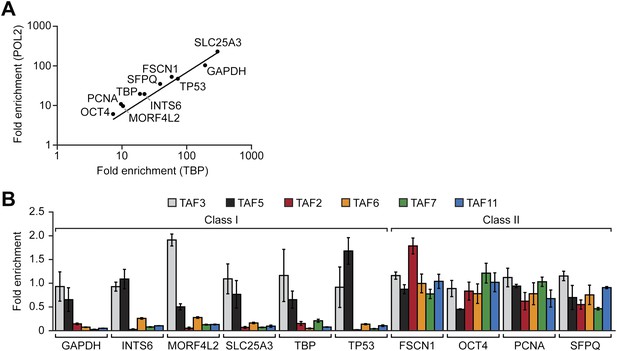
Two classes of hESC genes based on TAF promoter occupancy.
(A) Recruitment of TBP and POL2 to the promoters of 10 transcriptionally active genes (GAPDH, INTS6, MORF4L2, SLC25A3, TBP, TP53, FSCN1, OCT4, PCNA, SFPQ), each represented by a data point, were monitored by ChIP in H9 cells. For each gene, enrichment of TBP or POL2 binding to the promoter was normalized to a no antibody control and for non-specific recruitment at a control locus. (B) ChIP analysis monitoring TAF recruitment to the promoters of the 10 genes in H9 cells. TAF recruitment is specified relative to TBP recruitment (which was set to 1), after normalizing to a no antibody control and for non-specific recruitment to a control gene desert locus. Data are represented as mean ± SD.
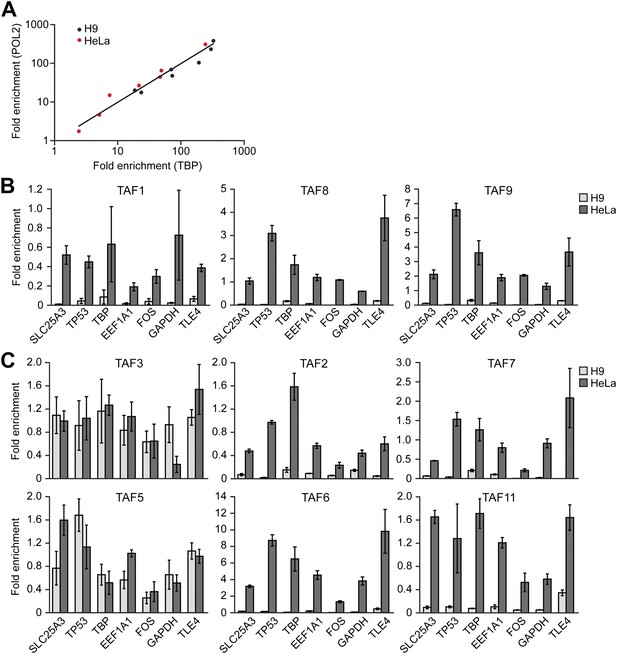
Comparison of TAF promoter occupancy on an identical set of transcriptionally active genes in HeLa and H9 cells.
(A) Recruitment of TBP and POL2 to the promoters of seven class I genes (EEF1A1, FOS, GAPDH, SLC25A3, TBP, TLE4 and TP53) were monitored by ChIP in HeLa and H9 cells. Data were normalized as described in Figure 3A. (B) ChIP analysis monitoring recruitment of TAFs 1, 8 and 9 to the promoters of the seven class I genes in HeLa and H9 cells, as described in (A). Data are represented as mean ± SD. (C) ChIP analysis monitoring recruitment of TAFs 2, 3, 5, 6, 7 and 11 to the promoters of the seven class I genes in HeLa and H9 cells, as described in (A). Data are represented as mean ± SD.
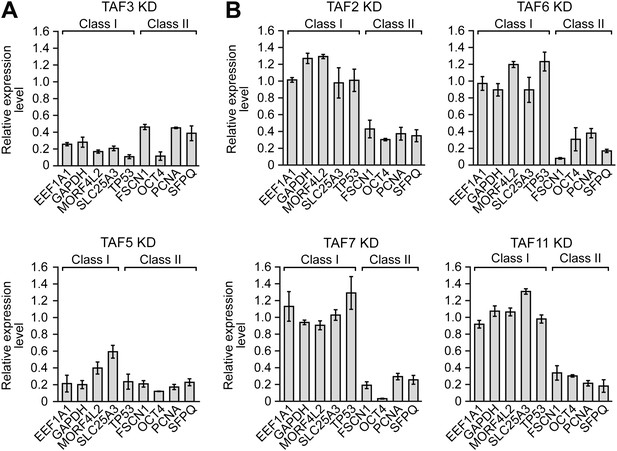
A strong relationship between TAF occupancy and transcriptional activity in hESCs.
(A) qRT-PCR analysis monitoring expression of class I and II genes in H9 TAF3 or TAF5 knockdown (KD) cells. Normalized Ct values were analyzed after subtracting the signal obtained with the control RN18S1 shRNA (see ‘Materials and methods’). Data are represented as mean ± SEM. (B) qRT-PCR analysis monitoring expression of class I and II genes in H9 TAF2, 6, 7, or 11 KD cells. Data are represented as mean ± SEM.
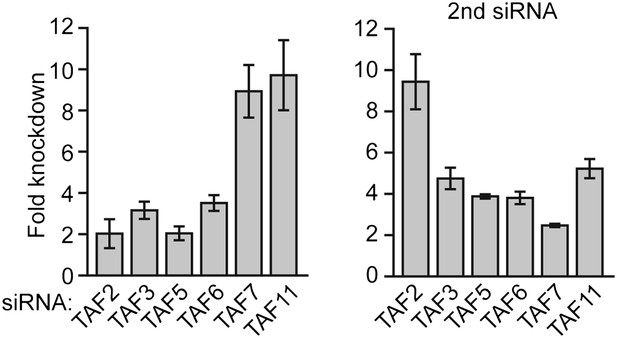
siRNA-mediated knockdown efficiency of TAFs in H9 hESCs.
qRT-PCR analysis monitoring TAF expression in H9 cells treated with two independent siRNAs (A) directed against the indicated TAF. TAF expression is specified relative to that obtained with a control luciferase siRNA, which was set to 1. Data are represented as mean ± SD.
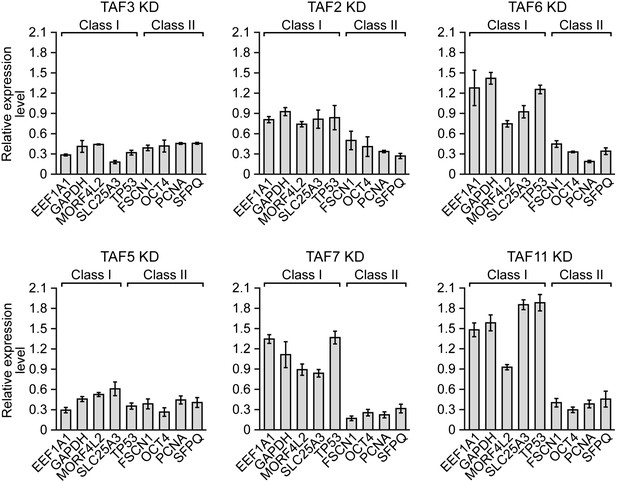
Confirmation of TAF requirement for transcriptional activity using a second, unrelated siRNA.
qRT-PCR analysis monitoring expression of class I and class II genes in H9 cells treated with a TAF3, TAF5, TAF2, TAF6, TAF7 or TAF11 siRNA. Expression of each gene is specified relative to that obtained with a control luciferase siRNA, which was set to 1. Data are represented as mean ± SEM.
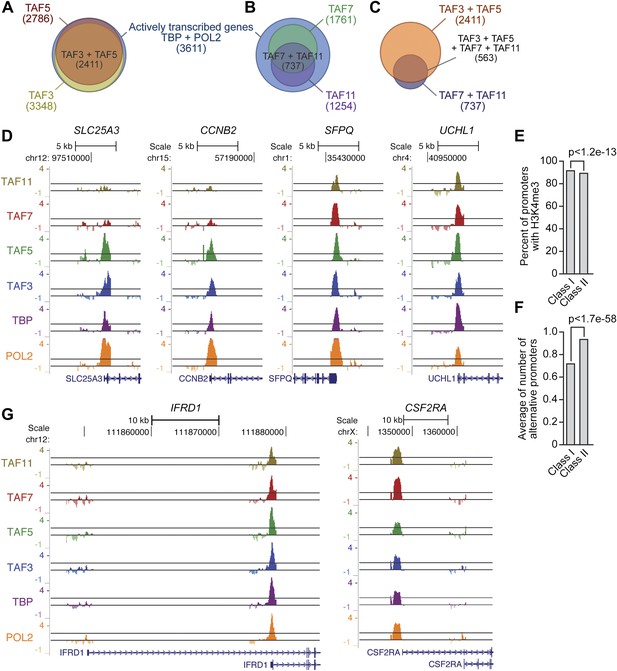
Global ChIP-chip analysis of TAF occupancy.
(A) Venn diagram showing the overlap between TBP-, POL2-, TAF3- and TAF5-bound genes. (B) Venn diagram showing the overlap between TBP-, POL2-, TAF7- and TAF11-bound genes. (C) Venn diagram showing the overlap between TAF3- and TAF5-bound genes and TAF7- and TAF11-bound genes. (D) Representative maps showing TAF3, TAF5, TAF7, TAF11, TBP and POL2 occupancy at the promoters of class I (SLC25A3 and CCNB2) and class II (SFPQ and UCHL1) genes. (E) Differences between class I and II genes with respect to promoter H3K4me3. (F) Differences between class I and II genes with respect to average number of alternative promoters per ChIP-enriched site. (G) Representative promoter occupancy maps for two class II genes with alternative promoters, IFRD1 and CSF2RA.
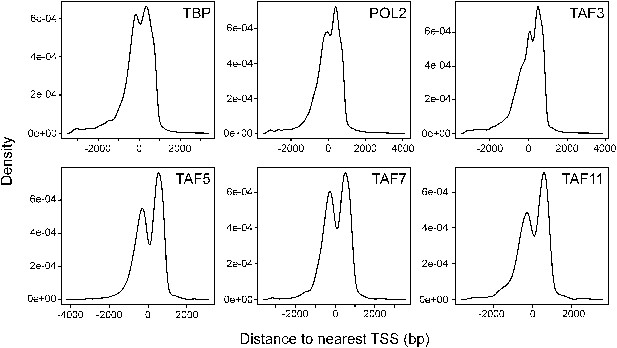
Location of TBP, POL2 and TAF occupancy relative to the transcription start site.
Histograms showing binding of TBP, POL2 and TAFs as a function of distance to the nearest transcription start site (TSS). The results show that binding of TBP, POL2 or a TAF occurred predominantly near the TSS.
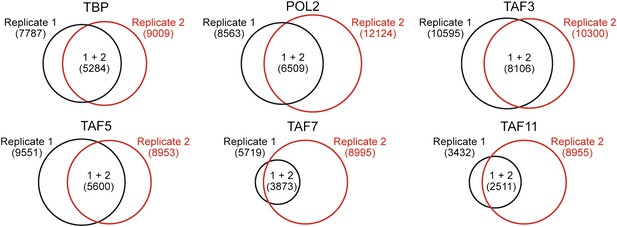
ChIP-chip peak overlap in independent replicates.
Venn diagrams showing the degree of overlap between two independent replicates of the ChIP experiment. The number of peaks in each group are indicated by brackets.
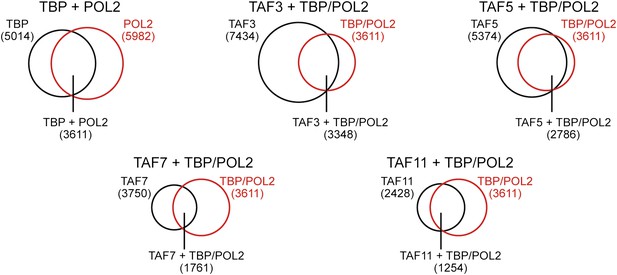
Co-occupancy of TBP and POL2 with TAFs.
Venn diagrams showing the degree of overlap between the number of genes whose promoters are bound by TBP and POL2, and the number bound by a given TAF.
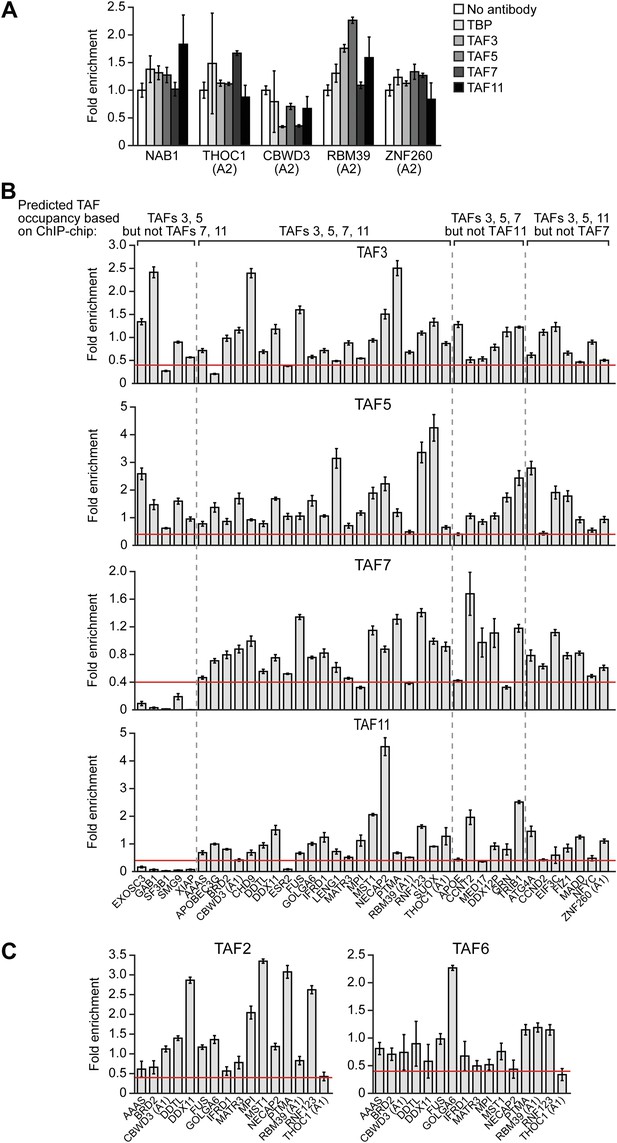
Validation of ChIP-chip results by directed ChIP experiments using promoter-specific primer pairs.
(A) ChIP analysis monitoring binding of TBP and TAFs 3, 5, 7 and 11 to a representative set of promoters that, based on ChIP-chip analyses, were predicted not to be bound by these factors. The results were normalized to a no antibody control (which was set to 1). Data are represented as mean ± SEM. For four genes (THOC1, CBWD3, RBM39 and ZNF260), the inactive promoter (A2) was analyzed in (A), and the active promoter (A1) was analyzed in (B) and (C). (B) ChIP analysis monitoring binding of TAFs 3, 5, 7 and 11 to the promoters of a representative set of 47 genes predicted by the ChIP-chip analyses to be bound by some or all of the factors. Data are normalized to TBP. Cutoff for a ‘positive’ is >0.4-fold enrichment vs TBP (red line). Data are represented as mean ± SEM. (C) ChIP analysis monitoring binding of TAF2 and TAF6 to 19 promoters that are bound by TAF7 and TAF11. Data are represented as mean ± SEM.
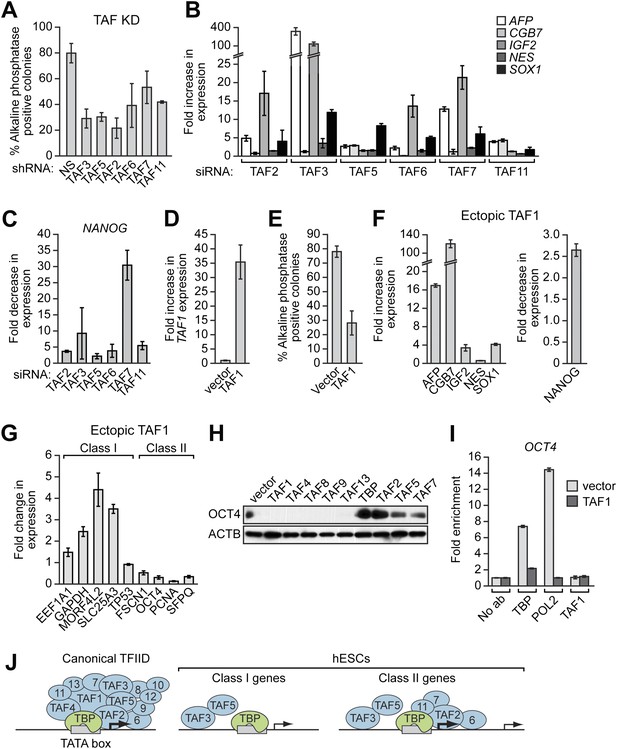
The composition of hESC TAFs is required for maintenance of the undifferentiated state.
(A) Percent of H9 TAF knockdown (KD) colonies staining with alkaline phosphatase. Data are represented as mean ± SD. (B) qRT-PCR analysis monitoring expression of differentiation markers (AFP, CGB7, IGF2, NES and SOX1) in H9 cells treated with a TAF siRNA. Values are relative to those obtained with a control luciferase siRNA, which was set to 1. Data are represented as mean ± SEM. (C) qRT-PCR analysis monitoring expression of NANOG in H9 cells treated with a TAF siRNA. Values are relative to those obtained with a control luciferase siRNA, which was set to 1. Data are represented as mean ± SEM. (D) qRT-PCR analysis monitoring TAF1 expression in H9 cells transfected with a plasmid expressing TAF1 or, as a control, empty vector. Expression of TAF1 was monitored 48 hr following transfection. TAF1 expression is specified relative to that obtained with the empty vector, which was set to 1. Data are represented as mean ± SEM. (E) Alkaline phosphatase staining of H9 colonies ectopically expressing TAF1 or, as a control, vector. Data are represented as mean ± SD. (F) qRT-PCR monitoring expression of differentiation markers (AFP, CGB7, IGF2, NES and SOX1) in H9 cells ectopically expressing TAF1. Values are relative to those obtained in H9 cells expressing vector, which was set to 1. Data are represented as mean ± SEM. (G) qRT-PCR monitoring expression of class I and II genes in H9 cells ectopically expressing TAF1. Data are represented as mean ± SEM. (H) Immunoblot analysis showing OCT4 levels in H9 cells over-expressing TAFs, TBP or vector. (I) ChIP analysis monitoring recruitment of TBP, POL2 and TAF1 to the OCT4 promoter in H9 cells ectopically expressing TAF1 or vector. Data are represented as mean ± SD. (J) Schematic model. Some of the protein interactions shown are arbitrary.
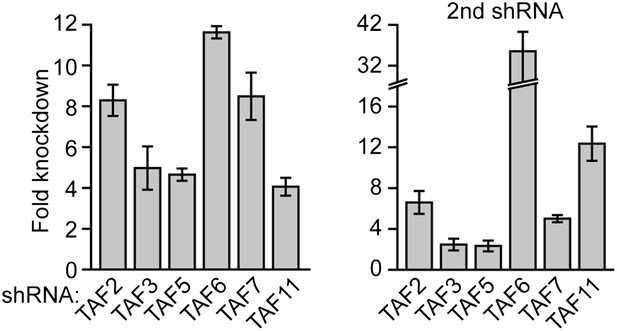
shRNA-mediated knockdown efficiency of TAFs in H9 hESCs.
qRT-PCR analysis monitoring TAF expression in H9 cells treated with two independent shRNAs directed against the indicated TAF. TAF expression is specified relative to that obtained with a control non-silencing shRNA, which was set to 1. Data are represented as mean ± SD.
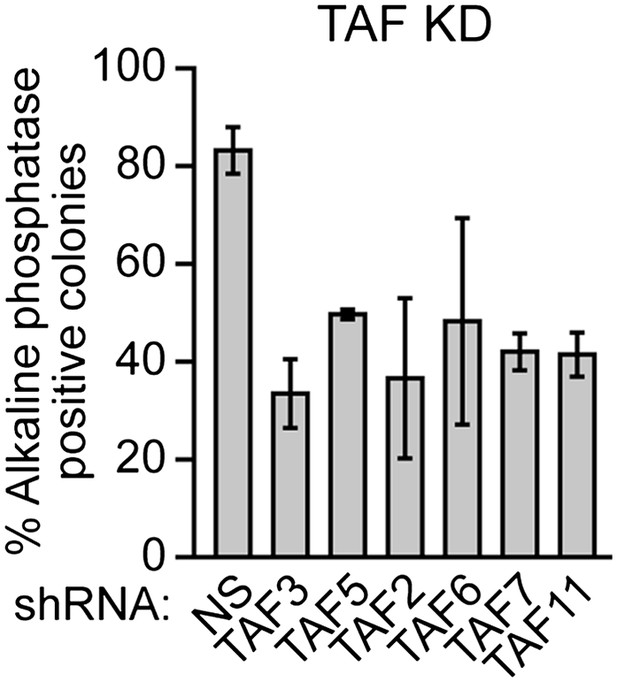
Validation of results presented in Figure 8A using a second, unrelated shRNA.
Percent of H9 TAF knockdown (KD) colonies staining with alkaline phosphatase. Data are represented as mean ± SD.
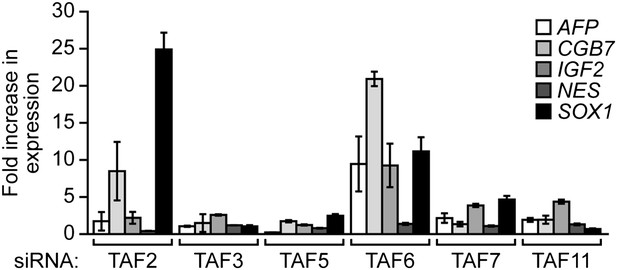
Validation of results presented in Figure 8B using a second, unrelated siRNA.
qRT-PCR analysis monitoring expression of differentiation markers in H9 cells treated with a TAF siRNA. Values are given relative to that obtained with a control luciferase siRNA, which was set to 1. Data are represented as mean ± SEM.
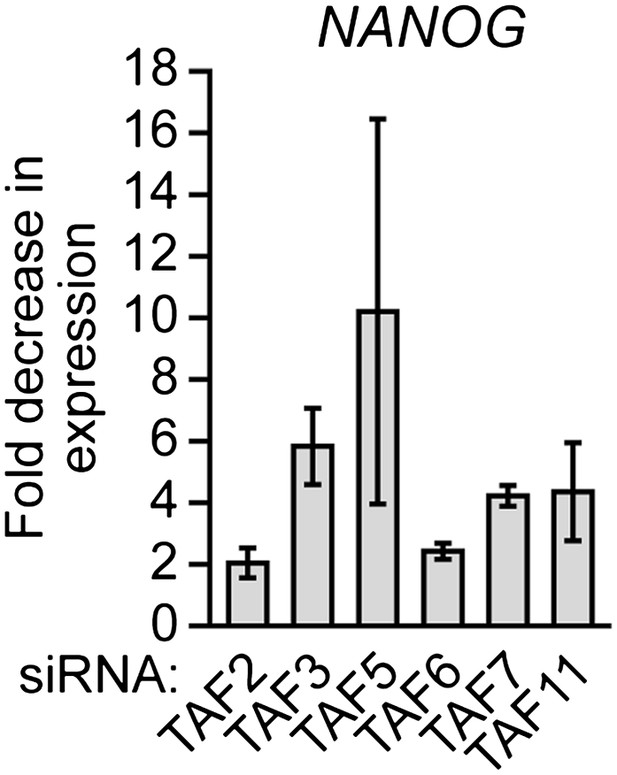
Validation of results presented in Figure 8C using a second, unrelated siRNA.
qRT-PCR analysis monitoring expression of NANOG in H9 cells treated with a TAF siRNA. Values are given relative to that obtained with a control luciferase siRNA, which was set to 1. Data are represented as mean ± SEM.
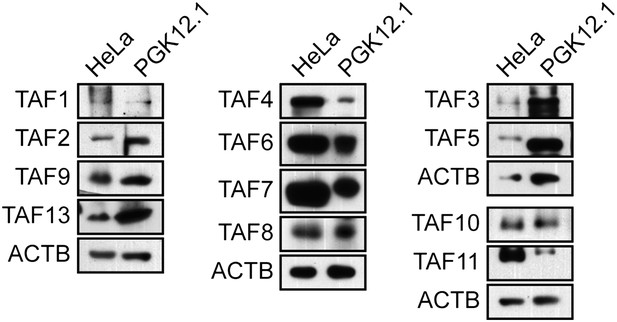
TAF expression in mouse ESCs.
Immunoblot analysis monitoring TAF levels in PGK12.1 mouse ESCs and, as a control, HeLa cells. ACTB was monitored as a loading control. The results show that TAFs 1, 4, 8, 9, 10 and 13, which are not expressed in hESCs, are expressed in mouse ESCs.
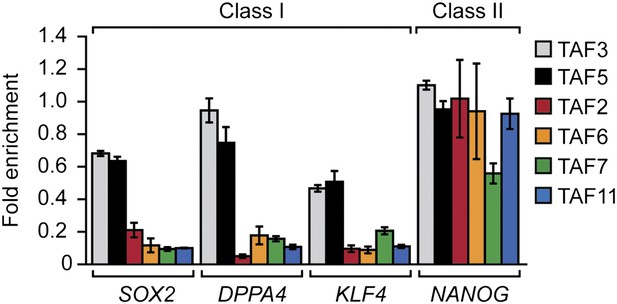
Classification of additional pluripotency genes as either class I or class II.
ChIP analysis monitoring TAF recruitment to the promoters of four pluripotency genes in H9 cells. TAF recruitment is specified relative to TBP recruitment (which was set to 1), after normalizing to a no antibody control and for non-specific recruitment to a control gene desert locus. Data are represented as mean ± SEM.
Tables
Statistical analysis of ChIP-chip data
| Feature | All TBP + POL2 sites | Class I genes (p-value vs TBP + POL2) | Class II genes (p-value vs TBP + POL2) | p-value (Class I vs Class II) |
| Alternative promoters (average number) | 0.717 | 0.686 (0.008) | 0.934 (2.64 × 10−67) | 1.73 × 10−58 |
| Bidirectional promoters (percent) | 23.961 | 25.315 (0.005) | 28.186 (0.007) | 0.110 |
| Promoter occupancy (percent) | ||||
| H3K4me3 | 91.482 | 89.214 (8.72 × 10−05) | 82.721 (4.24 × 10−20) | 1.16 × 10−13 |
| OCT4 | 3.684 | 3.919 (0.728) | 3.064 (1.000) | 0.293 |
| NANOG | 12.438 | 13.448 (0.006) | 13.725 (0.698) | 0.861 |
| SOX2 | 10.243 | 11.471 (9.64 × 10−05) | 11.642 (0.474) | 0.901 |
| Promoter elements (percent) | ||||
| TATA box | 4.834 | 6.365 (5.15 × 10−15) | 7.353 (0.012) | 0.335 |
| BRE | 88.999 | 92.413 (6.27 × 10−24) | 92.892 (0.001) | 0.650 |
| Initiator | 99.608 | 99.928 (0.000) | 100.000 (1.000) | 0.402 |
| MTE | 0.679 | 0.611 (1.000) | 0.735 (1.000) | 0.626 |
| DCE | 99.843 | 99.964 (0.422) | 99.877 (1.000) | 0.402 |
Additional files
-
Supplementary file 1
List of antibodies, primer sequences and siRNA/shRNAs used in this study. (A) List of antibodies used in this study. (B) Primers sequences for qRT-PCR, ChIP and ChIP-chip validation experiments. (C) Sequences of synthesized siRNAs, and clone ID numbers for shRNAs obtained from Open Biosystems (the luciferase siRNA sequence was previously reported (Elbashir et al., 2001)).
- https://doi.org/10.7554/eLife.00068.026





