Elba, a novel developmentally regulated chromatin boundary factor is a hetero-tripartite DNA binding complex
Figures
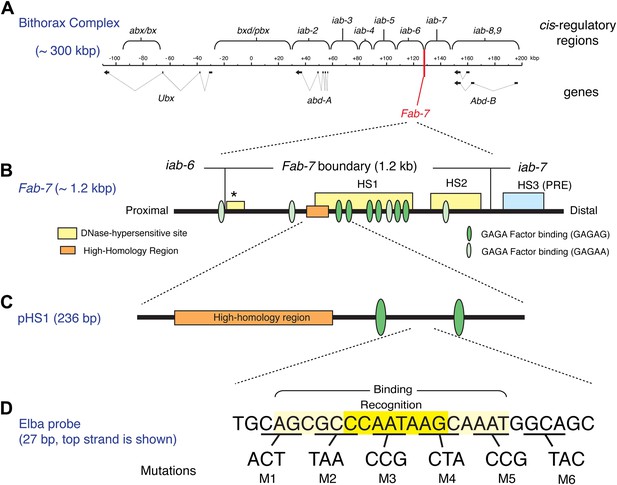
The Bithorax Complex, the Fab-7 boundary and the Elba recognition element.
(A) Drosophila Bithorax complex (BX-C). BX-C spans ∼300 kb and includes three Hox-family genes Ultrabithorax, abdominal-A and Abdominal-B. Parasegment specific expression of these three homeotic genes is generated by a series of functionally autonomous cis-regulatory domains: abx/bx, bxd/pbx and iab-2-iab8, 9. Functionally autonomy depends upon boundary elements that lie between each cis-regulatory domain (Maeda and Karch, 2010). One of these boundary elements is Fab-7, which is located in between the iab-6 and iab-7 cis-regulatory domains. Both in the context of BX-C and in transgene assays, the Fab-7 boundary can block the action of enhancers/silencers at all stages of development, apparently irrespective of tissue or cell type (Galloni et al., 1993; Hagstrom et al., 1996; Mihaly et al., 1997; Schweinsberg et al., 2004). (B) The Fab-7 boundary spans a sequence of 1.2 kb and consists of two prominent and one minor (*) chromatin specific nuclease hypersensitive regions (shown as yellow boxes). There is a third prominent nuclease hypersensitive region (blue) just distal to the boundary, which corresponds to a Polycomb Response Element (PRE) for the iab-7 cis-regulatory domain (Maeda and Karch, 2010). The orange box is a ∼100 base pair (bp) high-homology region which is conserved among Drosophila species (>90%) (Aoki et al., 2008). The ovals are binding sites for Trithorax-like (GAGA factor). (C) pHS1 is a 236-bp fragment from the proximal side of HS1 which has enhancer-blocking activity only in early embryos (Schweinsberg and Schedl, 2004). pHS1 includes the high-homology region and two GAGA-binding sites. These two GAGA sites are important for the early boundary activity of Fab-7, while GAGA sites elsewhere in Fab-7 are needed later in development (Schweinsberg et al., 2004). In addition to the GAGA sites, the enhancer-blocking activity of pHS1 in early embryos also depends upon an 8-bp sequence, CCAATAAG, called Elba (Early boundary activity). Mutations in this sequence compromise the blocking activity of a 4×pHS1 multimer, while multimerization of a 27-bp oligo spanning the Elba sequence (8×Elba) [see (D)] is sufficient to confer early blocking activity. The Elba sequence is recognized by the stage-specific Elba DNA-binding factor. Elba factor binding is detected in 0–6 hr nuclear extracts, but it is absent in 6–12 hr (and 6–18 hr) nuclear extracts (Aoki et al., 2008). (D) Sequence of the 27-bp oligo used as the Elba probe in the EMSA experiments shown in Figures 3A, 4, 5B, and 6. The Elba factor in 0–6 hr nuclear extracts recognizes the 8-bp Elba sequence (shaded by yellow) and requires an additional 5 bp both upstream and downstream for full binding activity (shaded by light yellow). The bases underlined were altered as indicated in the mutant oligos, M1–M6. These mutant oligos were used as cold competitors in Figures 4C and 6C as indicated. For the DNA affinity beads, a 27-bp oligo containing the mutation M3 was used as the mutant Elba sequence.
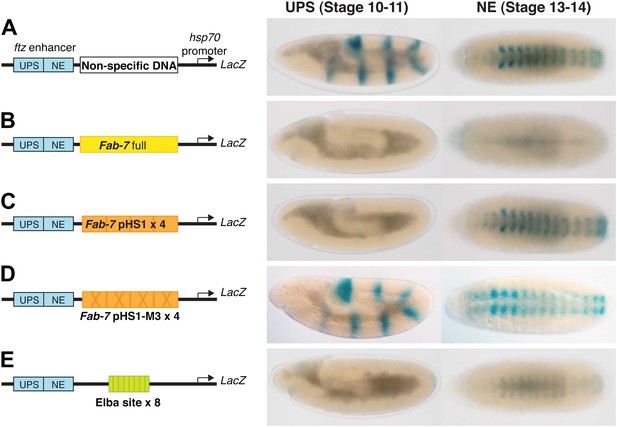
Enhancer blocking activity of Fab-7, pHS1×4 and the Elba×8 multimer.
In this enhancer blocking assay, putative boundary elements are placed in between two fushi tarazu (ftz) enhancers (UPS and NE) and an hsp70:LacZ reporter. The UPS enhancer is active in early embryos and drives LacZ expression a seven stripe pair-rule pattern. The NE enhancer is active later in development and drives LacZ expression in the central nervous system. (A) When a ‘non-specific’ DNA sequence is placed between the ftz enhancers and the hsp70:LacZ reporter, no blocking is observed. In early embryos (stage 10–11 in this assay), the ftz UPS enhancers drive LacZ expression in seven stripes. In older embryos (stage 13–14) the NE enhancer drives LacZ expression in the CNS. (B) When Fab-7 is place between the ftz enhancers and the reporter, it blocks both enhancers from activating the hsp70 promoter and there is little if any LacZ expression in either early or late embryos. (C) When pHS1×4 (four copies of the 236-bp pHS1: see Figure 1) is placed between the enhancers and the reporter, it blocks the UPS enhancer from activating stripe expression. However, boundary activity is stage specific and pHS1×4 does not block the NE from activating LacZ expression in the CNS in older embryos (Aoki et al., 2008). (D) When mutations (M3 in Figure 1) that disrupt Elba binding in nuclear extracts are introduced into the Elba sequence of pHS1, the enhancer blocking activity of the mutated pHS1×4 is compromised. In this case, both UPS stripe and NE CNS LacZ expression is observed (Aoki et al., 2008). (E) When a 27-bp oligo containing the Elba binding sequence is multimerized (Elba×8) and placed in between the ftz enhancers and the reporter, it blocks UPS driven LacZ stripe expression. It also causes a reduction in the extent of NE activation. This effect is position dependent. The blocking activity seen at this stage with the 8× multimer is likely due to the presence of some residual Elba activity in older embryos as seen in EMSA experiments with 6–12 hr nuclear extracts (Aoki et al., 2008).
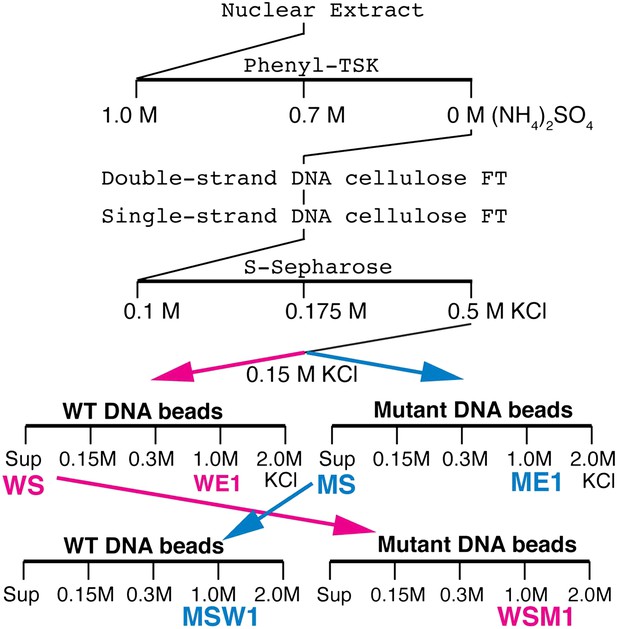
Elba purification scheme.
Details provided in ‘Materials and methods’.
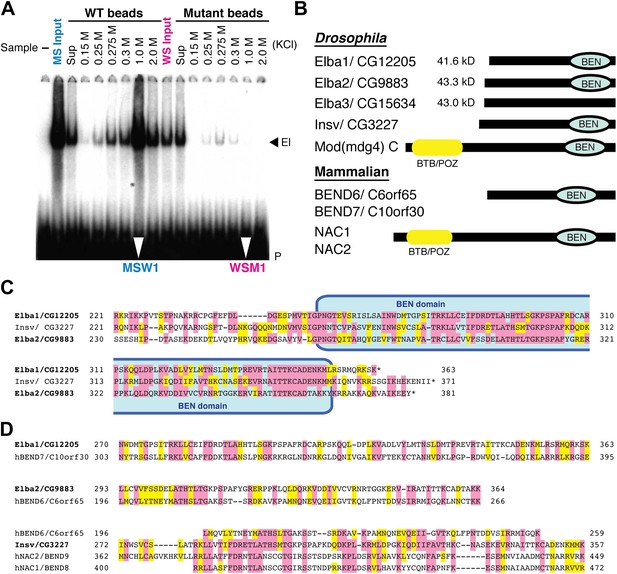
EMSA and Elba factor proteins.
(A) EMSA of fractions from the cross-affinity purification. The 32P-labeled Elba probe was incubated with fractions as indicated and subjected to 4% acrylamide-gel electrophoresis. El: Elba shift. P: probe. (B) Schematic of the Elba factors and BEN domain-containing (green) orthologs. BTB/POZ (yellow) domain is absent from Elba factors but is present in related proteins. (C) Sequence alignment of the C-terminal half of the Drosophila Elba1, Elba2 and a third Drosophila Ben protein Insv (Insensitive). The sequences of the three proteins were aligned according to the results of NCBI (National Center for Biotechnology Information) blast search (bl2seq). The amino acid residues conserved in more than two proteins are shaded with red. The residues that have similarities with each other are shaded with yellow. The predicted BEN domain region is boxed with pale blue. (D) Sequence alignment of Drosophila and mammalian (human) orthologs of BEN domain proteins. The C-terminal sequences of Elba1, Elba2 and Insv were subjected to blast search with human databases. Within the BEN domain sequences, the closest human ortholog of Elba1 is BEND7 (BEN domain-containing 7)/C10orf30 (29% identical, 46% positive), whereas for Elba2 the closest ortholog is BEND6/C6orf65 (29% identical, 53% positive). The BEN domain of Insv is most similar to three proteins: NAC2 (Nucleus accumbens-associated protein 2, NACC2)/BEND9, BEND6 and NAC1 (Nucleus accumbens-associated protein 1, NACC1)/BEND9. The amino acid residues conserved with each Drosophila protein are shaded with red. The residues that have similarities with each Drosophila protein are shaded with yellow.
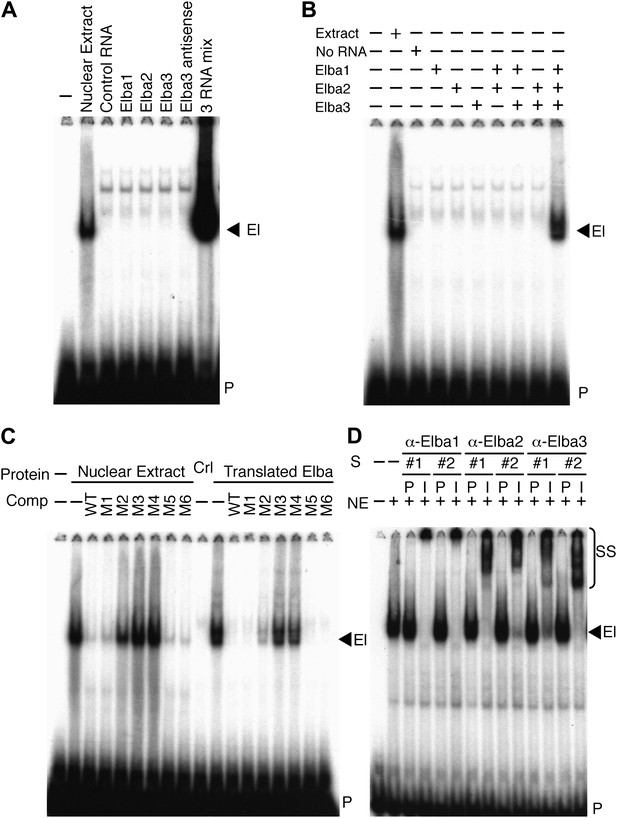
Elba is a hetero-tripartite complex.
(A) and (B) All three Elba proteins are required to reconstitute DNA binding activity. In vitro translated proteins either singly or in combination as indicated were incubated with the Elba probe. El: Elba shift. P: probe. (C) Reconstituted Elba has the same sequence specificity as nuclear Elba. Reconstituted and nuclear extract shifts with or without (minus) a 100-fold excess of competitor (Comp) as indicated above each lane. WT: wild-type probe. M1–M6: mutant probes (see Figure 1D). Ctl: no-RNA control. (D) Nuclear Elba factor has all three Elba proteins. Nuclear extracts (NE) incubated with preimmune (P) or immune (I) polyclonal rabbit serum as indicated. #1, #2: serum from different rabbits. SS: antibody supershifts. El: Elba shift. P: free probe.
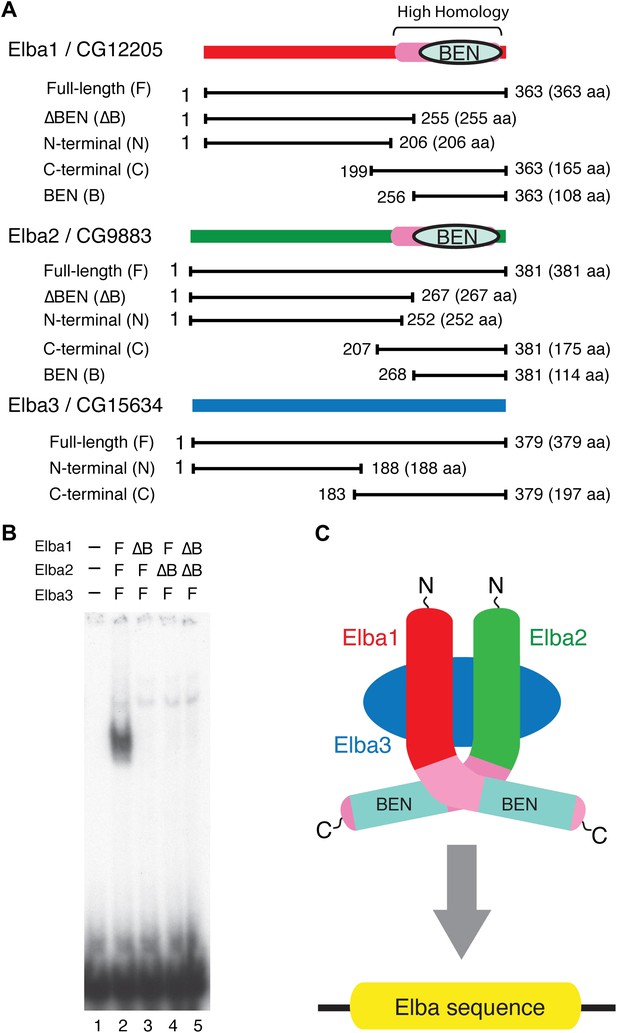
Elba protein deletions and functional organization of the Elba complex.
(A) Elba proteins and deletion mutants. The bars under each diagram indicate the sequences retained in the mutant protein. The numbers correspond to the amino acid residues at the N and C terminal ends of the protein. The letter in parentheses is used to designate proteins added to the lanes in panel B and in Figure 6. Each protein was expressed either with an N-terminal FLAG tag or with an N-terminal FLAG tag plus a GST tag. The FLAG tag was used to determine the relative amount of each protein so that the input of the translated proteins in the gel shift experiments could be adjusted. (B) BEN domain is required for the DNA-binding activity of Elba. EMSA experiments were performed as described in Figure 4 by translating the full-length (F) or ∆BEN (∆B) mutant Elba subunits in vitro and by mixing each as indicated above the lanes. (C) A schematic structure of proposed model for the Elba complex.
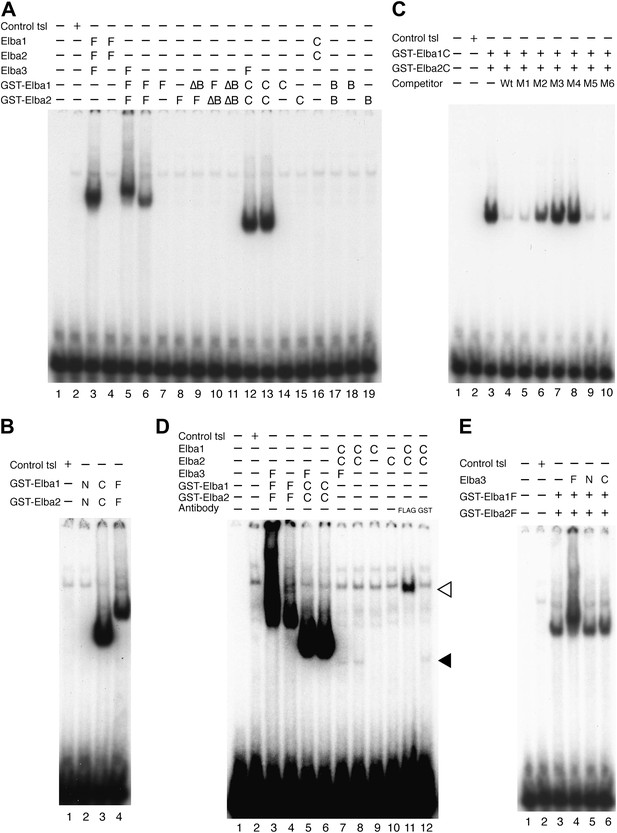
Characterization of Elba complex.
(A) DNA binding activity of different Elba protein variants. The proteins were translated in vitro from the mixed mRNAs shown above the lanes. All proteins used in this figure are FLAG tagged and approximately the same amounts of the translated proteins were added to each lane. The identities of the proteins added to each lane are indicated above the lane. For example, lane 3 has all three full length Elba proteins, while in lane 5 full length Elba3 is combined with full length Elba1 and Elba2 proteins that have an N-terminal GST tag. Lane 13 has GST fused to the C-terminal half of Elba1 and Elba2, while in lane 16 the C terminal halves of Elba1 and Elba2 lack the GST moiety. (B) GST-fused Elba1F:Elba2F (F) or Elba1C:Elba2C (C) binds to the Elba probe while Elba1N:Elba2N (N) does not. (C) Sequence specificity of the GST-Elba1C:GST-Elba2C dimer is the same as the nuclear/reconstituted hetero-tripartite complex (Figure 4). The EMSA experiment with GST-Elba1C:GST-Elba2C were performed in the absence (lanes 1–3) or presence of 100-fold excess of non-labeled competitors as indicated above the lanes. The sequences of competitor DNAs are shown in Figure 1. (D) Elba1C and Elba2C proteins lacking the N-terminal GST moiety have a weak DNA binding activity. Position of Elba1C:Elba2C shift is indicated by closed arrowhead. Position of the Elba1C:Elba2C FLAG supershift is indicated by open arrowhead. Proteins added to each lane including FLAG and GST antibodies are indicated above the lane. Note that the same shift was detected in lane 16 of (A) when the X-ray film was exposed for a longer period of time. (E) Supershifts of the GST-Elba1F:GST-Elba2F generated by the addition of Elba3. Proteins corresponding to the full length Elba3 (F), or the N-terminal (N) and C-terminal (C) halves of Elba3 were added to an EMSA reaction mix containing the GST-Elba1:GST-Elba2 dimer. Proposed structures of the native Elba complex, the artificial GST-dependent dimer by GST-Elba1:GST-Elba2 and GST-Elba1C:GST-Elba2C are shown in Figure 6—figure supplement 1–3, respectively.
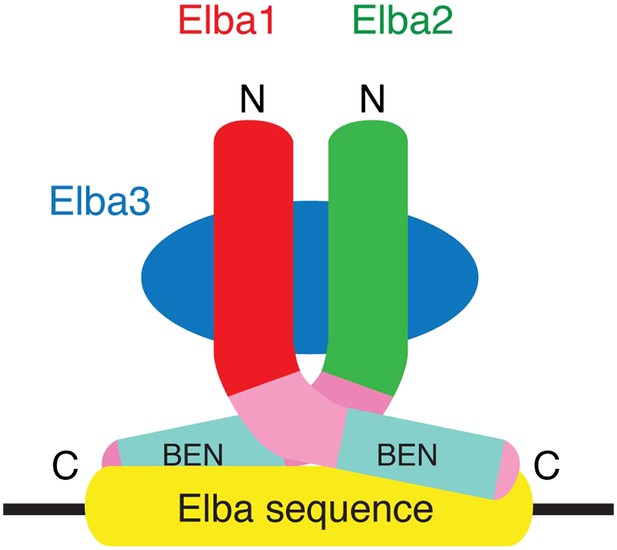
A schematic model for the tripartite Elba complex.
Elba3 links Elba1 and Elba2 through sequences in their N-terminus. This brings the C-terminal halves of Elba1 and Elba2 together to form a ‘DNA binding pocket’ which binds to the asymmetric Elba recognition sequence.
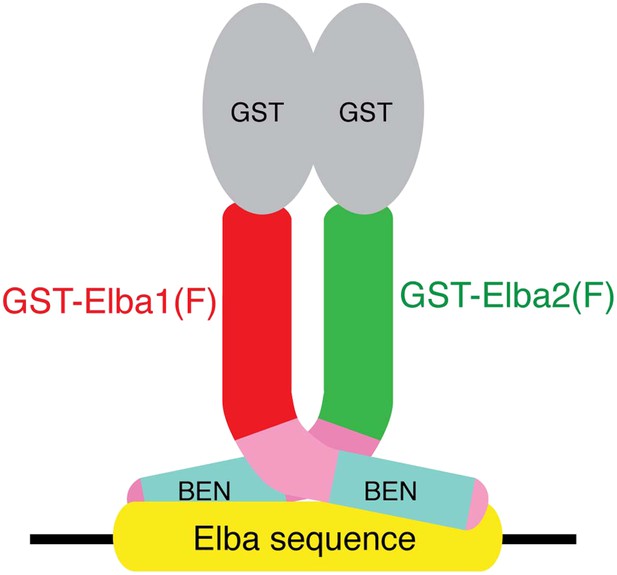
The artificial GST-Elba1:GST-Elba2 heterodimer binds to the asymmetric Elba recognition sequence.
The N-terminal GST moiety mediates dimerization of Elba1 and Elba2, mimicking the coupling of these two proteins by Elba3.
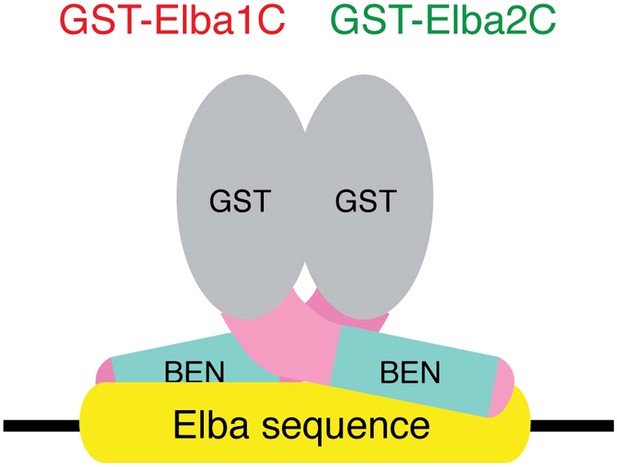
The artificial heterodimer of GST-Elba1C:GST-Elba2C binds to the asymmetric Elba recognition sequence.
The C-terminal 165–175 residues of Elba1 and Elba2 retain DNA-binding activity with the same sequence-specificity as hetero-tripartite Elba complex when they are dimerized by GST moieties.
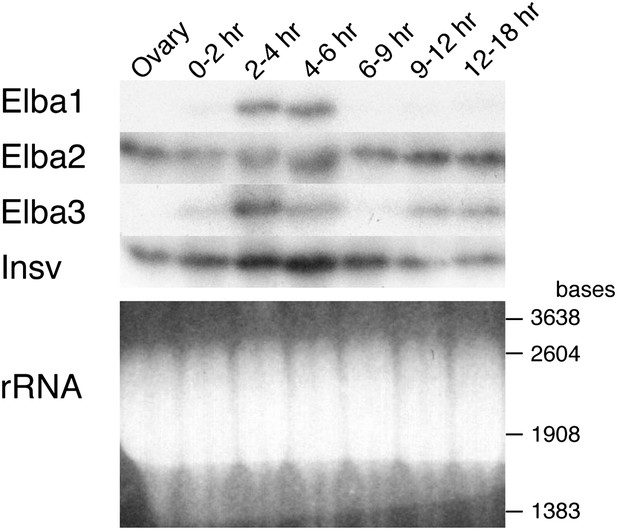
Expression of mRNAs encoding Elba factors and Insv.
Total RNA from ovaries and staged embryos as indicated were probed with cDNAs encoding each protein. Shown on the bottom is the ethidium bromide staining of a gel for the Northern blotting as a loading control. The positions of the RNA markers are indicated on the right.
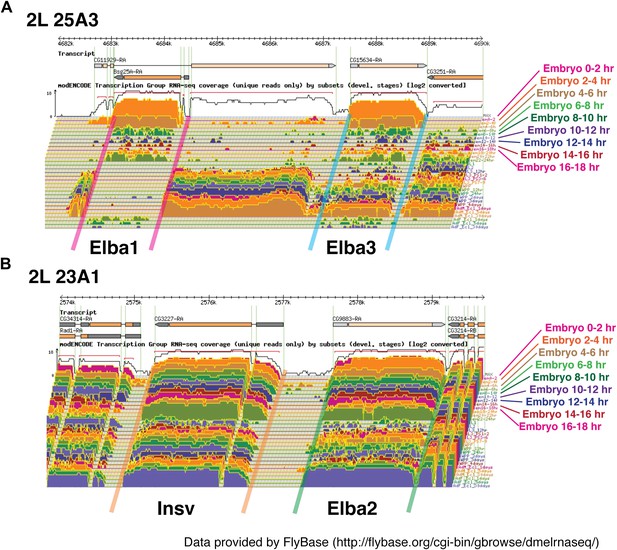
Loci of Elba proteins and the expression patterns of their mRNAs during the development.
(A) modEncode expression profiles of mRNAs encoding Elba1 and Elba3. As indicated in the diagram, elba1 (CG12205) and elba3 (CG15634) are transcribed in opposite directions and are separated from each other by sequences encoding much of the CG11929 open reading frame http://flybase.org/reports/FBgn0000227.html/, http://flybase.org/reports/FBgn0031621.html. As seen in the Northerns, there is little or no maternal deposition of elba1 or elba3 mRNAs. These two genes are not expressed until the mid-blastula transition and the highest levels of elba1 and elba3 mRNAs are found in 2–4 hr embryos. Relatively high levels of both mRNAs are found in 4–6 hr embryos and then both largely but not completely disappear during the remainder of embryogenesis. In our Northern experiments transcript levels also drop dramatically after 6 hr, but in older embryos both elba1 and elba3 are detected again. However, it is likely that the amounts of elba1 and elba3 in these later stages are overestimated in our experiments. Our gels were overloaded, and transfer of the two elba mRNAs (which migrate very near the bottom of the rRNA band) in the 2–4 hr samples does not appear to be as efficient as it is at later stages when their levels are greatly reduced. The presence of only quite low levels of elba1 and elba3 mRNAs at these later stages would be consistent with the modEncode data. (B) modEncode expression profiles of mRNAs encoding Elba2 and Insv. Like elba1 and elba3, elba2/CG9883 and insensitive/CG3227 are closely linked and transcribed in opposite orientations. As seen in our Northern experiments, the modEncode temporal expression patterns of the mRNAs encoding these two proteins are quite different from that of elba1 and elba3 http://flybase.org/reports/FBgn0031434.html/, http://flybase.org/reports/FBgn0031435.html. Both are maternally deposited and high levels are present throughout embryogenesis. Also unlike elba1 and elba3, both transcripts are expressed at later stages of development.
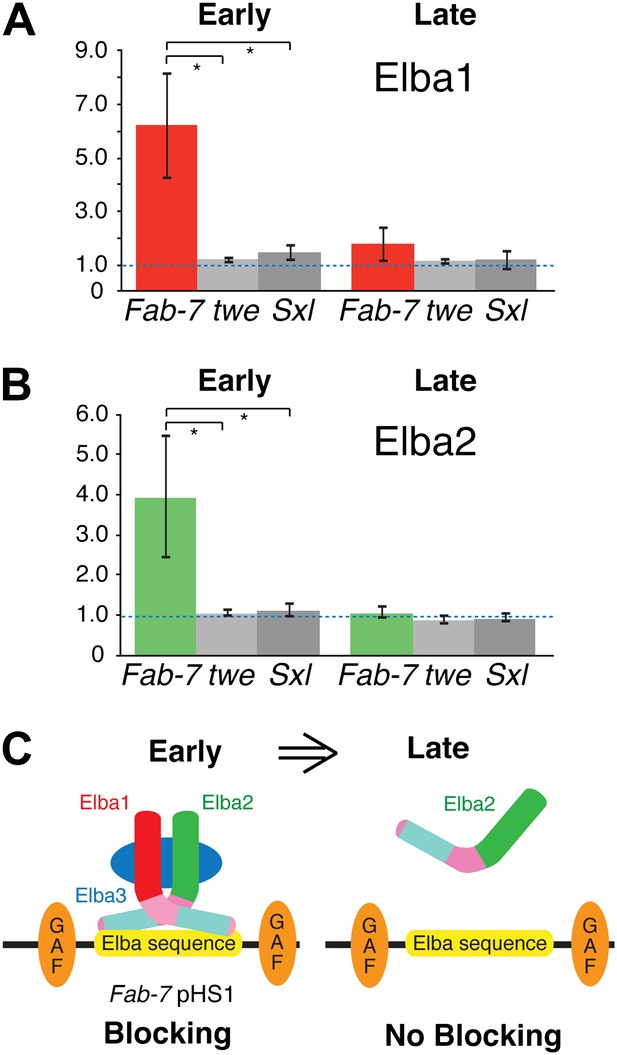
Elba proteins are bound to Fab-7 in early but not late embryos.
(A) and (B) Elba1 and 2 ChIPs. Early (2–5 hr) or late (9–12 hr) embryos were cross-linked and after processing immunoprecipitated with Elba1 or Elba2 antibodies, or pre-immune serum. Sequences from the pHS1 region of Fab-7 or control twine and Sex-lethal sequences were detected by qPCR. The y-axis shows the average immune/preimmune ratio. *p<0.05. (C) Model showing binding and enhancer blocking by the Elba complex in early but not late embryos. GAF: GAGA factor and its binding sites.
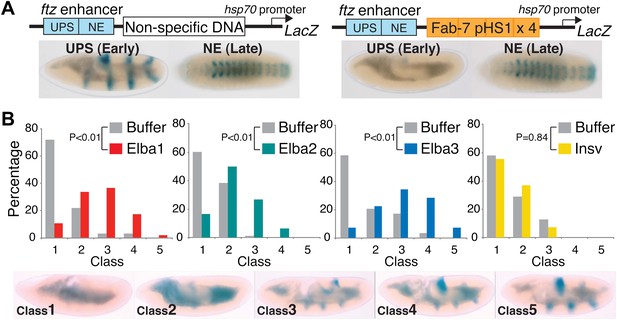
Hetero-tripartite Elba complex mediates early boundary activity.
(A) Top: fushi tarazu (ftz) enhancers drives stripe expression (UPS) in stage 10–11 embryos and central nervous system expression (NE) in stage 13–14 embryos. Bottom: Four copies of Fab-7 pHS1 (which contains the Elba sequence) blocks the UPS, but not the NE enhancer. elba1-3 and insv dsRNAs or buffer alone were injected into embryos transgenic for the ftz-4×pHS1-LacZ enhancer blocking reporter and blocking activity examined. In each experiment ∼100 embryos were photographed and categorized into Class 1–5 as indicated. Graphs show the percentage of embryos in each class for RNAi and the buffer control in that experiment. Three independent injection experiments were done for each protein. All gave similar results and only one is shown here. p values from the t-tests are shown in the graphs.
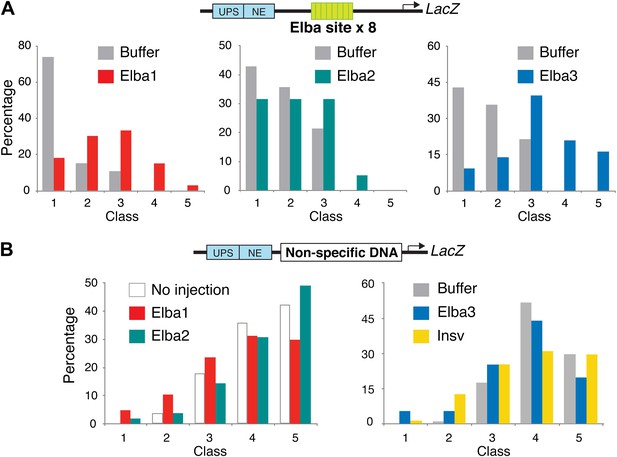
Double-stranded RNA injection into embryos transgenic for the Elba×8 multimer reporter and a reporter containing non-specific DNA in the blocking position.
(A) Embryos transgenic for the ftz-Elba×8 (multimer)-Hsp70:LacZ reporter (see Figure 1—figure supplement 1) were injected with dsRNAs specific for each of the elba genes. The injected embryos were allowed to develop until stage 10–11 and then stained for LacZ expression. In each experiment, buffer-injected control embryos were prepared and processed in parallel with the dsRNA injected embryos. Embryos were photographed and then categorized into Class 1–5 according to the intensity of ftz UPS-enhancer stripes. (Figure 9 shows representative examples of each class.) The graphs show the percentage of embryos (y-axis) that fell into each class (x-axis) in the elba dsRNAi and the corresponding control buffer injection. As was observed in the pHS1×4 injection experiments, the blocking activity of the Elba×8 multimer is compromised by injection of dsRNA for each of elba gene. The loss of blocking activity can be seen by comparing the class distribution for the dsRNA injection with the class distribution for the buffer control. (Note that the elba2 and elba3 dsRNA injections shown here were part of the same injection experiment and have the same buffer injected control.) The dsRNA injections for the two mid-blastula transition genes elba1 and elba3 have a greater effect on the blocking activity of the Elba×8 multimer than the dsRNA injections for elba2. This was observed in other Elba×8 multimer injection experiments and was also seen for the pHS1×4 reporter (see Figure 9). The smaller reduction in blocking activity observed with elba2 dsRNA injections is most likely due to the presence of substantial amounts of maternal elba2 mRNA. (B) elba and insv dsRNA injections do not affect LacZ expression from a control reporter which has a non-specific DNA sequence in the blocking position. The expressions of the UPS LacZ stripes in dsRNA and in the buffer injected or no injection controls give a similar class distribution.
Tables
List of proteins unique to the 1.0 M KCl fraction from wild type DNA affinity beads (MSW1) in the third, cross-affinity, Elba purification
| Hit protein | Sequence count | Spectrum count | Sequence coverage | Mol. wt. | |
| 1 | CG12205 (Bsg25A)-PA/gi|7295685* | 23 | 372 | 56.5%/49%* | 41,583/47,368* |
| 2 | CG15634-PA | 16 | 150 | 32.20% | 43,008 |
| 3 | CG9883-PA | 17 | 63 | 26.80% | 43,310 |
| 4 | CG12052 (lola) 17 subtypes | 3 | 21 | 10.8–5.2% | 49,320–98,162 |
| 5 | CG12052 (lola)-PY | 3 | 21 | 8.70% | 62,746 |
| 6 | CG12052 (lola), unknown subtype | 3 | 21 | 5.10% | 107,303 |
| 7 | CG12052 (lola)-PD, -PE | 3 | 21 | 6.70% | 79,439 |
| 8 | CG14339-PA | 2 | 17 | 2.00% | 117,908 |
| 9 | CG2368(psq)-PD, -PE, -PF, PG, -PH/-PB, -PC | 6 | 15 | 12.7%/7.7% | 70,298/114,984 |
| 10 | CG1249 (snRNP2)-PA | 3 | 13 | 31.90% | 13,504 |
| 11 | CG6944 (Lam)-PA, -PB, -PC | 8 | 12 | 19.10% | 71,249 |
| 12 | CG16973 (msn)-PE | 4 | 11 | 4.50% | 115,386 |
| 13 | CG16973 (msn)-PC, -PD/-PB /-PA | 4 | 11 | 4.3%/3.9%/3.1% | 120,610/130,341/162,378 |
| 14 | CG11700 (CR11700)-PA | 2 | 11 | 10.60% | 34,335 |
| 15 | Reverse_CG31284 | 2 | 11 | 3.10% | 110,298 |
| 16 | CG3561 (KH1)-PA | 8 | 10 | 15.30% | 59,597 |
| 17 | CG10067 (Act57B)-PA/CG7478 (Act79B)-PA/CG18290 (Act87E)-PA, -PB | 5 | 10 | 14.40% | 41,835/41,787/41,802 |
| 18 | CG5178 (Act88F)-PA | 5 | 10 | 14.40% | 41,700 |
| 19 | CG1759 (cad)-PA, -PB | 4 | 9 | 4.40% | 51,306 |
| 20 | CG12154 (oc)-PA | 3 | 9 | 5.20% | 69,666 |
| 21 | CG6801 (l(3)j2D3)-PA | 8 | 8 | 19.20% | 44,830 |
| 22 | CG18124 (mTTF)-PA | 7 | 8 | 16.30% | 48,281 |
| 23 | CG7583 (CtBP)-PD, -PB, -PC, -PA | 6 | 8 | 20.50% | 42,252 |
| 24 | CG13634-PA | 2 | 8 | 4.30% | 52,594 |
| 25 | CG3143 (foxo)-PC, -PB | 5 | 7 | 9.60% | 67,413 |
-
*
An alternative transcript of CG12205 (gi:7295685) that encoded a slightly larger protein was listed in the Genbank database at the time of this experiment. However, that sequence was subsequently removed from the CG12205 sequence list.
-
In the third, cross-affinity purification, there were 176 proteins in the MSW1 fraction (see Figure 2) which were not present in the WSM1 fraction. These 176 proteins were sorted according to the descending order of ‘Spectrum Count’ numbers. The 25 proteins with the highest ‘Spectrum Count’ are shown here. The top three proteins have a high spectrum count, and also have a high sequence count.
Additional files
-
Supplementary file 1
The list of oligonucleotides used in this study. See also ‘Materials and methods’ for the use of each DNA.
- https://doi.org/10.7554/eLife.00171.019





