Rapid localized spread and immunologic containment define Herpes simplex virus-2 reactivation in the human genital tract
Figures
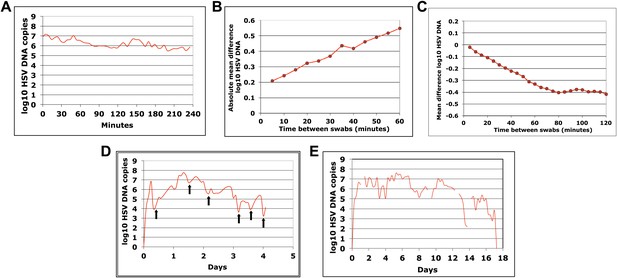
HSV-2 levels in the genital tract are stable over minutes, expand and decay markedly over hours, and fluctuate rapidly and unpredictably over days.
(A) Shedding quantity in a participant, who performed genital swabs every 5 min over 4 hr during a lesion, reveals low swab-to-swab variation in viral quantity. Using data from panel (A and B), absolute mean difference (R2 = 0.99), and (C) mean difference (R2 = 0.87), in HSV DNA copies between swabs, are a function of time between swabs. (D) Shedding quantity in a participant, who performed 10 genital swabs per day during a lesion over 4 days (swabs every 2–4 hr), shows a characteristic saw-tooth pattern; arrows denote rapid viral re-expansion; the participant had a negative swab performed before episode onset. (E) Shedding quantity in a participant, who performed four genital swabs per day over 17 days demonstrates that rapid and frequent viral re-expansion allows for shedding prolongation; four missing data points are left blank.
-
Figure 1—source data 1
Source data for Figure 1, Figure 1—figure supplement 1, Figure 1—figure supplement 2 and Figure 1—figure supplement 3.
- https://doi.org/10.7554/eLife.00288.008
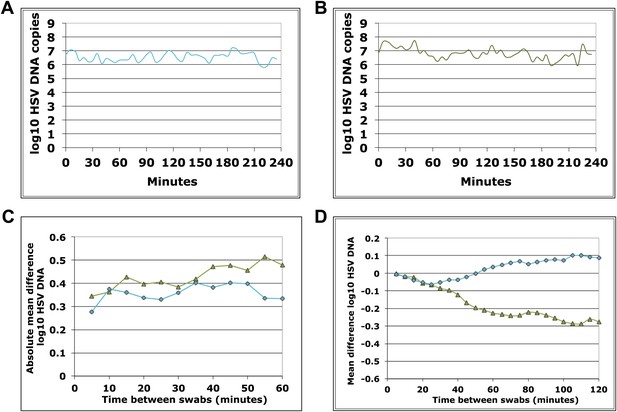
Dynamics of HSV-2 shedding over 5-min time intervals.
(A) and (B) Shedding quantity in two participants, who performed genital swabs every 5 min over 4 hr during a lesion, reveals low swab-to-swab variation in viral quantity. (C) Using data from panels (A and B), absolute mean difference in HSV DNA copies between swabs, correlated moderately with time between swabs in one participant (green line R2 = 0.60) due to steady viral decay, but was more stable as a function of time in the other participant due to peaking overall viral load (blue line, R2 = 0.16). (D) Using data from panels (A and B), mean difference in HSV DNA copies between swabs, was a function of time between swabs in the participants (green line R2 = 0.89, and blue line R2 = 0.85) presumably because viral load was generally decaying during most of the 4-hr window (green) or gradually expanding during most of the 4-hr window (blue).
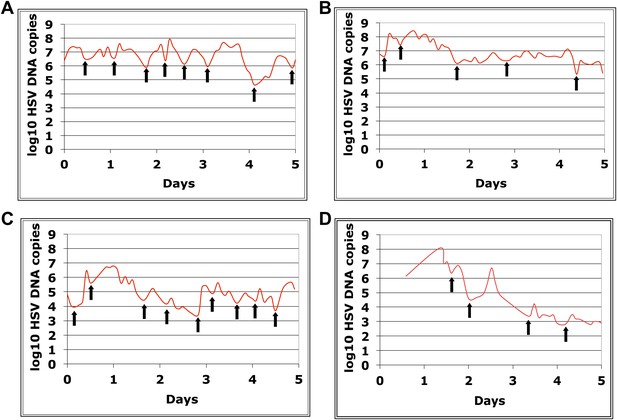
Dynamics of HSV-2 shedding with every 2-4 hr sampling.
Shedding quantity in four participants, who performed 10 genital swabs per day over 4–5 days during a lesion reveal a characteristic saw-tooth pattern; arrows denote re-expansion. Participants had swabbing initiated upon visualization of lesions. The participant in panel d initiated swabbing ∼16 hr after lesion detection.
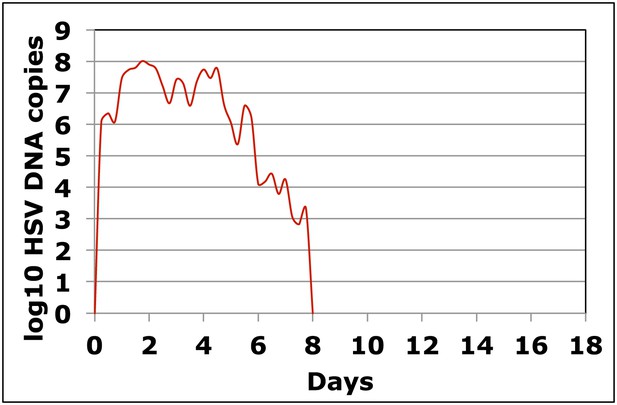
Dynamics of HSV-2 shedding with every 6-hr sampling over 8 days.
Shedding quantity in episodes detected in a participant who performed four genital swabs per day over 60 days demonstrates that viral re-expansion allows for shedding prolongation.
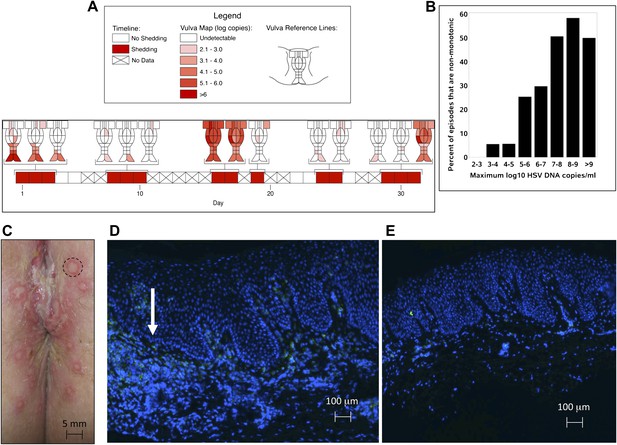
HSV-2 replicates and is contained in widely dispersed microenvironments across the genital tract.
(A) HSV shedding quantity in a participant, who underwent daily swabs in 23 regions across the genital tract for 30 days; days without sampling are marked with an X; stars denote days with a lesion; virus is widely dispersed and several prolonged episodes with heterogeneous viral loads across the genital tract are noted. (B) Increasing probability of episode re-expansion (nonmonotonic episodes) as a function of peak episode copy number among 1020 episodes from 531 study subjects; individual peaks during episodes may represent virus from a single ulcer that can seed other regions. (C) A genital lesion consists of numerous round ulcers (black dotted circle) clustered in space; contemporaneous presence of multiple ulcers may indicate concurrent viral expansion in decay in multiple regions. (D) and (E) Immunofluorescent staining of biopsies performed (D) at the edge, and (E) 1 cm away from an ulcer 3 days post-healing; CD8+ T cells (green) at the dermal–epidermal junction (arrow) are highly localized to ulcer edge (287/mm2) and are fourfold less dense 1 cm away (72/mm2).
-
Figure 2—source data 1
Source data for Figure 2 and Figure 2—figure supplement 1.
- https://doi.org/10.7554/eLife.00288.014
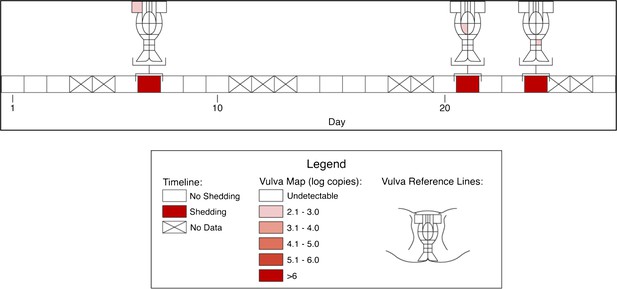
Spatial features of HSV genital tract shedding.
HSV shedding quantity in a study participant, who underwent daily swabs in 23 regions across the genital tract for 30 days; days without sampling are marked with an X. The participant had three brief localized episodes with low viral copy number in three separate localized regions.
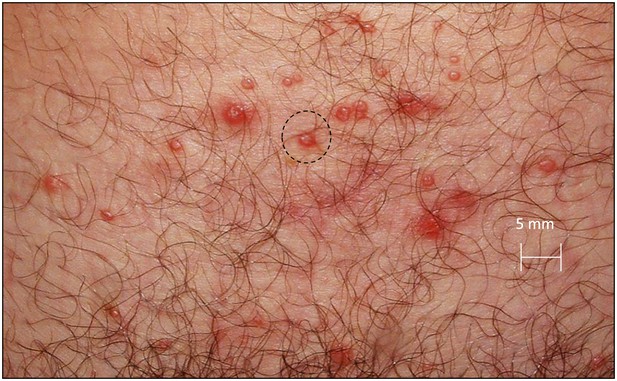
Spatial features of HSV-2 lesions.
A genital lesion consists of numerous round ulcers or vesicles (black dotted circle), clustered in space.
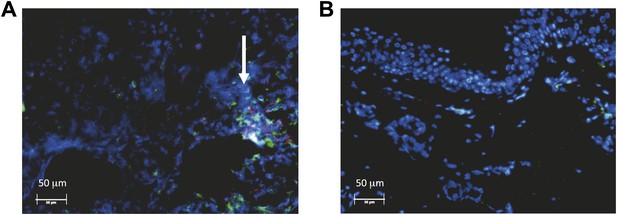
Spatial features of CD8+ T-cell response in genital skin.
Immunofluorescent staining of a biopsy performed (A) at the edge, and (B) 1 cm away from an ulcer 3 days post-healing. CD8+ T-cells (red) and CD4+ T-cells (green) at the dermal epidermal junction (arrow) are highly localized to ulcer edge (132/mm2 and 447/mm2, respectively), and are less dense 1 cm away (91/mm2 and 132/mm2, respectively).
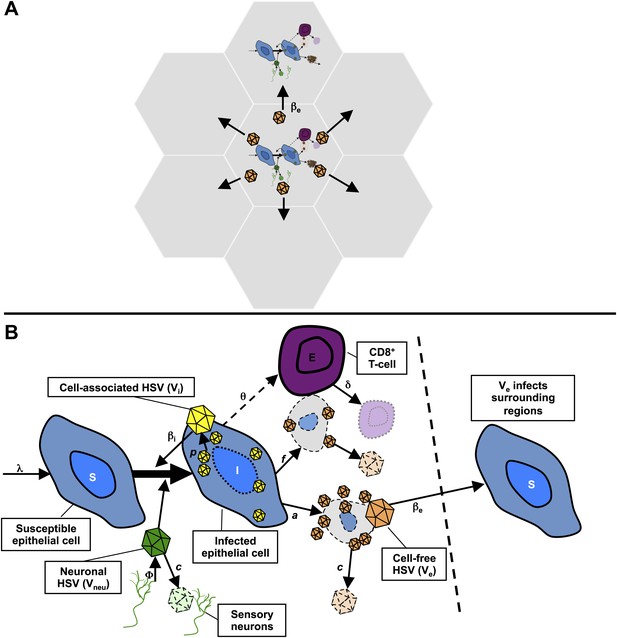
Mathematical model.
(A) Microregions are linked virally because cell-free HSV-2 can seed surrounding regions, and immunologically based on overlapping CD8+ T-cell densities between regions (not shown). (B) Schematic for HSV-2 infection within a single genital tract microenvironment. Equations capture seeding of epithelial cells by neuronal HSV-2, replication of HSV-2 within epithelial cells, viral spread to other epithelial cells, cytolytic CD8+ T-cell response to infected cells, transition of cell-associated HSV-2 to cell-free HSV-2 following lysis of infected cells, and elimination of free virus and infected cells.
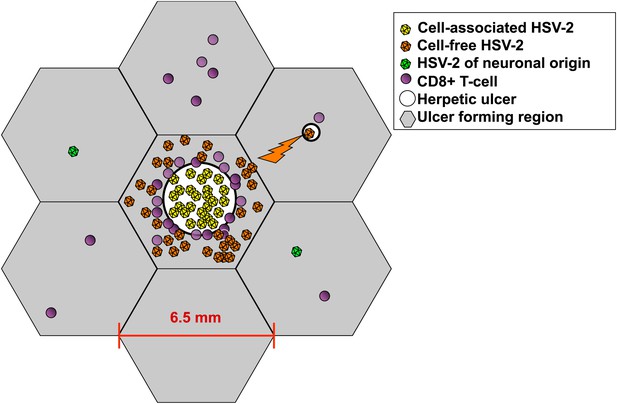
Spatial mathematical model.
Viruses produced from neurons (green), cell-associated viruses from epidermal cells (yellow), and cell-free viruses (orange) that form after rupture of epidermal cells, are distinguished in the model. Neuron-derived viruses are released throughout the genital tract and are responsible for ulcer initiation within specific regions (grey hexagons). Cell-associated HSV particles contribute to ulcer expansion (white circle) within a region. Cell-free particles initiate secondary ulcers in adjacent regions (upper right) leading to concurrent ulcers where HSV production occurs. Cytolytic CD8+ T-cell (purple circles) response is localized within each region. Regions have a maximum diameter of 6.5 mm. However, distance between regions is considered in terms of immunologic co-dependence rather than a physical distance. Seven of 300 total model regions are illustrated.
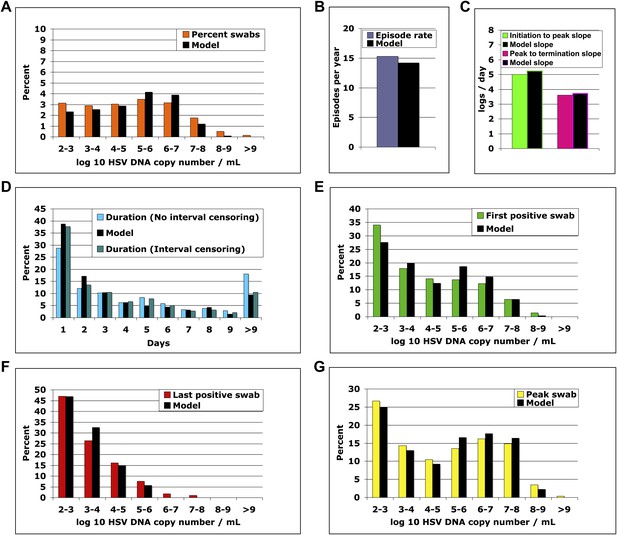
The spatial model reproduces all shedding episode characteristics.
Colored bars represent results from (A) 14,685 genital swabs and (B–G) 1020 shedding episodes from 531 study participants. The model simulation, represented with black bars in each panel, continued until 1020 episodes were generated; model sampling occurred every 24 hr as in the clinical protocol. Model output reproduced (A) quantitative shedding frequency as well as (B) rate, (C) median initiation to peak and peak to termination slopes, (D) Duration, (E) first HSV DNA copy number, (F) last HSV DNA copy number, and (G) peak HSV DNA copy number of episodes.
-
Figure 4—source data 1
Source data for Figure 4.
- https://doi.org/10.7554/eLife.00288.022
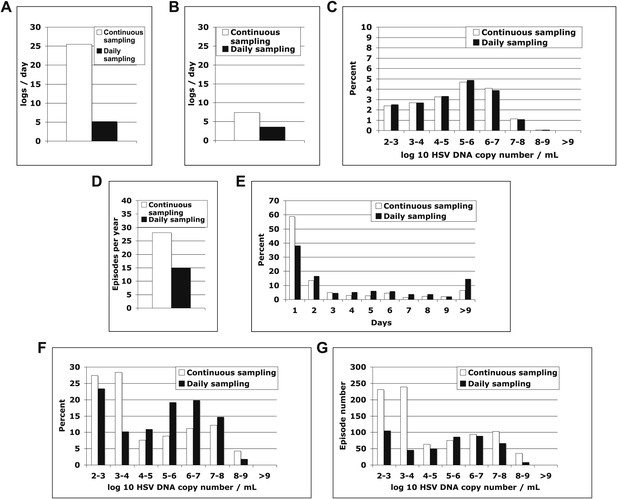
Continuous sampling of spatial model output reveals more accurate measures of episode characteristics.
We subjected a 30-year simulation to daily and continuous sampling. (A) Median initiation to peak slope, and (B) peak to termination slopes increased substantially with continual sampling. (C) Shedding frequency was similar regardless of sampling frequency. (D) Continuous sampling detected 842 episodes (28.1/year) vs 450 episodes (15.0/year) with daily sampling. The 392 additional episodes were all less than a day in duration and mostly <104 peak HSV DNA copies per milliliter, skewing the distributions of (E) episode duration and (F) peak HSV DNA copy number. (G) Total number of episodes at low and high-peak copy numbers increased with continual sampling.
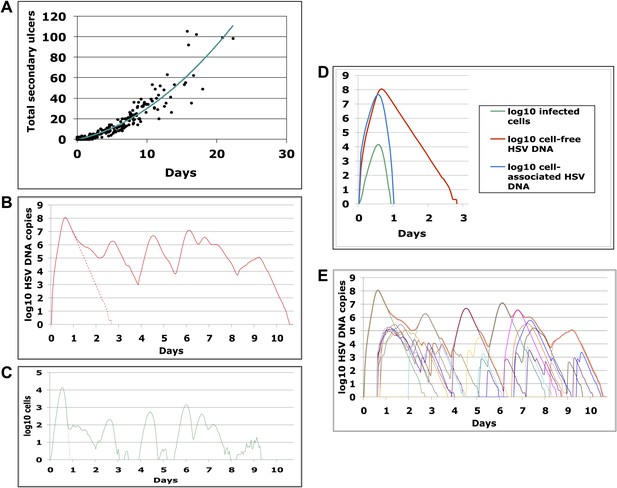
Containment of infected cells within a single ulcer is extremely rapid, although secondary ulcers explain prolonged episodes.
(A) Episode duration was a function of the number of ulcers before episode termination during 500 simulated episodes. (B)–(E) A 10-day simulated episode consisting of 24 ulcers: (B) Total cell-free virus (red) over time reflects the saw-tooth pattern of prolonged episodes; virus produced from the initial ulcer (red dotted line) was eliminated within 3 days. (C) Infected cells were eliminated from the initial viral ulcer (green dotted line) within 1 day and there were four periods during the episode when no infected cells were present. (D) Cell-free virus (red), cell-associated virus (blue), and infected cells (green) were eliminated from the primary ulcer with different kinetics; infected cells peaked at 13 hr and were extinguished in <24 hr (E) Secondary ulcers prolonged episodes; each thin line represents HSV-2 production from a specific region.
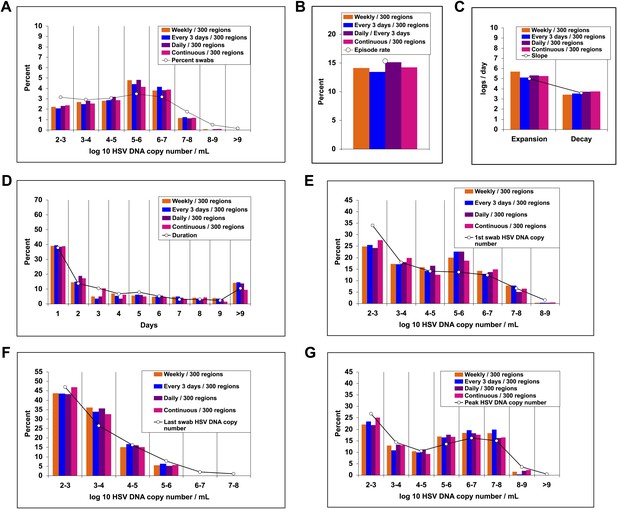
Random spatial dispersion of viral particles from neurons reproduced the full diversity of episode characteristics if particles were released continuously, daily, or weekly from neurons.
White circles represent results from (A) 14,685 genital swabs and (B–G) 1020 shedding episodes from 531 study participants (Figure 4). The model simulations represented with colored bars in each panel continued until 1020 episodes were generated. Sampling occurred every 24 hr as in the clinical protocols. Model output with release of virus randomly throughout the 300 regions on a continuous (pink), daily (purple), every 3 days (blue), and weekly (orange) basis at an average rate of 82 HSV DNA particles per day reproduced (A) quantitative shedding frequency and episode, (B) rate, (C) median initiation to peak slope and peak to termination slopes, (D) Duration, (E) first HSV DNA copy number, (F) last HSV DNA copy number, and (G) peak HSV DNA copy number.
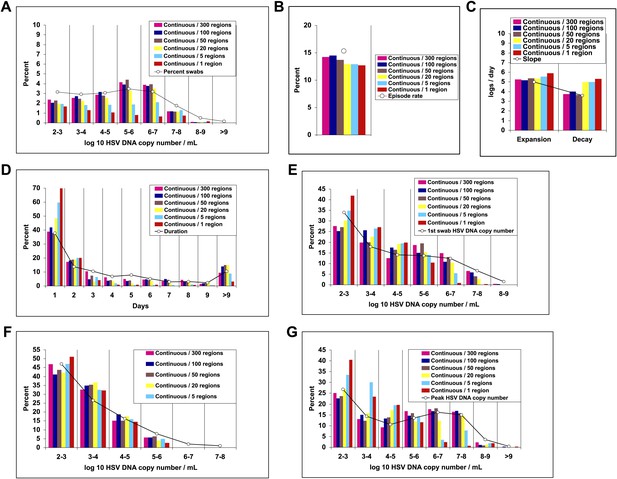
Random spatial dispersion of viral particles from neurons reproduced the full diversity of episode characteristics during simulations in which particles were released into only a minority of the 300 modeled regions.
White circles represent results from (A) 14,685 genital swabs and (B–G) 1020 shedding episodes from 531 study participants (Figure 4). The model simulations represented with colored bars in each panel continued until 1020 episodes were generated. Sampling occurred every 24 hr as in the clinical protocols. Model output with continuous release (82 HSV DNA particles per day) of virus randomly to 300 (pink), 100 (blue), and 50 (brown) regions reproduced (A) quantitative shedding frequency and episode, (B) rate, (C) median initiation to peak slope and peak to termination slopes, (D) Duration, (E) first HSV DNA copy number, (F) last HSV DNA copy number, and (G) peak HSV DNA copy number. Model output with continuous release (82 HSV DNA particles per day) of virus randomly to 20 (yellow), 5 (light blue), and 1 (red) region underestimated (A) quantitative shedding frequency and episode, (B) rate, (D) Duration, (E) first HSV DNA copy number, and (G) peak HSV DNA copy number.
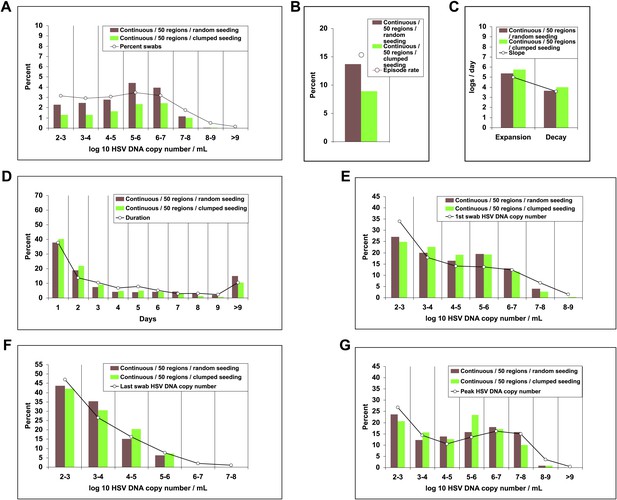
Dispersion of viral particles from neurons reproduced the full diversity of episode characteristics during simulations in which particles were released into a minority of modeled regions, provided that dispersion was random rather than clustered within a single geographic region.
White circles represent results from (A) 14,685 genital swabs and (B–G) 1020 shedding episodes from 531 study participants (Figure 4). The model simulations represented with colored bars in each panel continued until 1020 episodes were generated. Sampling occurred every 24 hr as in the clinical protocols. Model output with continuous release (82 HSV DNA particles per day) of virus to 50 randomly dispersed regions (brown) reproduced (A) quantitative shedding frequency and episode, (B) rate, (C) median initiation to peak slope and peak to termination slopes, (D) Duration, (E) first HSV DNA copy number, (F) last HSV DNA copy number, and (G) peak HSV DNA copy number. Model output with continuous release (82 HSV DNA particles per day) of virus to 50 clustered regions (green) underestimated (A) shedding frequency due only to (B) too low of an episode rate with daily sampling.
Videos
Spatiotemporal demonstration of a 10-day episode.
The left panel represents total cell-free HSV DNA copies per milliliter present over time. The right panel represents spatial spread of virus during the episode, each hexagon contains one region of shedding and virus spreads to contiguous regions. Amount of virus within a single region is displayed according to a heat map adjacent to the spatial map.
Spatiotemporal demonstration of a 14-day episode according to viral production within each single region.
The left panel represents cell-free HSV DNA measured over time with each region's production demonstrated with a different color. The right panel represents spatial spread of virus during the episode; colors in the right panel correspond to those in the left panel. Amount of virus within a region is displayed according to a heat map adjacent to the spatial map.
Spatiotemporal demonstration of infected cell and viral spread during a 6-day episode.
The upper left panel represents spatial spread of cell-free virus during the episode; the upper right panel represents spatial spread of cell-associated virus during the episode; the bottom left panel represents spatial spread of infected cells during the episode; the bottom right panel represents ulcer formation during the episode, ulcers turn from black to red when diameter exceeds 1 mm; quantities are displayed according to a heat map adjacent to each spatial map. There is more rapid decay of infected cells and cell-associated particles than of cell-free particles within each region. Visible ulcers persist after viral production terminates within a region.
Spatiotemporal demonstration of 365 days of simulated shedding.
The left panel represents total cell-free HSV DNA copies per milliliter present over time. The right panel represents spatial spread of virus during the episode; each hexagon contains one region of shedding and virus spreads to contiguous regions. Amount of virus within a single region is displayed according to a heat map adjacent to the spatial map. The simulation is notable for episodes of variable duration and peak HSV DNA copy number. Prolonged episodes at days 8, 38, 136, 158, and 288, display several re-expansion phases.
Spatiotemporal demonstration of 365 days of simulated shedding with noninfectious cell-free particles.
The left panel represents total cell-free HSV DNA copies per milliliter present over time. The right panel represents spatial spread of virus during the episode; each hexagon contains one region of shedding. Amount of virus within a single region is displayed according to a heat map adjacent to the spatial map. The simulation is notable for lack of prolonged episodes and lack of episode re-expansion.
Spatiotemporal demonstration of immune response to a pair of 2-day episodes.
The upper left panel represents total cell-free HSV DNA copies per milliliter present over time. The upper right panel represents spatial spread of cell-free virus during the episode. The lower left panel represents CD8+ T-cell density within each region. The lower right panel indicates reproductive number within each region. Quantities are displayed according to a heat map adjacent to each spatial map. Areas with high CD8+ T-cell levels and low reproductive numbers do not support high-level viral production. The simulated episodes are short because virus does not spread from the initial plaque to adjacent regions with high CD8+ T-cell density and reproductive numbers less than one.
Spatiotemporal demonstration of immune response to a 7-day episode.
The upper left panel represents total cell-free HSV DNA copies per milliliter present over time. The upper right panel represents spatial spread of cell-free virus during the episode. The lower left panel represents CD8+ T-cell density within each region. The lower right panel indicates reproductive number within each region. Quantities are displayed according to a heat map adjacent to each spatial map. The simulated episode is medium length because virus spreads from the initial region to adjacent regions with low CD8+ T-cell density and reproductive numbers less than one, but is ultimately contained when it reaches an anatomic edge region outside of the genital tract, and regions of high CD8+ T-cell density and low reproductive number within the genital tract.
Spatiotemporal demonstration of immune response to a 14-day episode.
The upper left panel represents total cell-free HSV DNA copies per milliliter present over time. The upper right panel represents spatial spread of cell-free virus during the episode. The lower left panel represents CD8+ T-cell density within each region. The lower right panel indicates reproductive number within each region. Quantities are displayed according to a heat map adjacent to each spatial map. The simulated episode is prolonged because virus spreads from the initial region to adjacent regions with low CD8+ T-cell density and reproductive numbers less than one, and is not contained until many regions of the genital tract are infected.
Spatiotemporal demonstration of immune response over 20 years.
The left panel represents CD8+ T-cell density within each region. The right panel indicates reproductive number within each region. Quantities are displayed according to a heat map adjacent to each spatial map. CD8+ T-cells expand rapidly at sites of episodes and then decay slowly over time, correlating with decreases and increases in reproductive number respectively. CD8+ T-cell and reproductive number spatial patterns cycle intermittently between a patchwork of heterogeneous density, broad low density, broad high density, and stark division between regions of high and low density. However, CD8+ T-cell density is virtually always high in at least some regions of the genital tract.
Tables
Five cohorts of HSV-2 genital tract shedding
| Cohort | Subjects | Total swabs | Swabbing frequency | Total episodes | Swabbing duration | Anatomic swabbing region | Purpose |
| A | 3 | 96 | Every 5 min | 3 | 4 hr when lesion present | Total genital tract | Swab-to-swab sampling/assay variability |
| B | 5 | 200 | 10 times/day (every 2 hr during the days and 4 hr overnight) | 5 | 4–5 days when lesion present | Total genital tract | Episode expansion, clearance and re-expansion kinetics |
| C | 25 | 4706 | 4 times/day | 109 | 30–60 days without or with a lesion | Total genital tract | Accurate estimates for expansion/decay slopes for clinical and subclinical episodes |
| D | 2 | 216 | Daily | 4 | 30 days without or with a lesion | 23 separate regions | Spatial dispersion of HSV |
| E | 531 | 14,685 | Daily | 1020 | >30 days with or without a lesion | Total genital tract | Model fit |
Parameter ranges that result in accurate reproduction of model outcomes
| Parameter | Units | Symbol | Best fit value | Good fit | Average fit | ||
| Lower limit | Upper limit | Lower limit | Upper limit | ||||
| Cell-associated HSV infectivity | DNA copy days/cell (viruses needed per day to infect one adjacent cell) | βi | 5.4e−8 (111) | 4.86e−8 (123) | 7.83e−8 (76) | 3.78e−8 (158) | 1.32e−7 (45) |
| Cell-free HSV infectivity | DNA copy days/cell (viruses needed per day to initiate one new ulcer) | βe | 2.65e−11 (2.26e5) | 1.73e−11 (3.46e5) | 2.78e−11 (2.15e5) | 3.98e−12 (1.50e6) | 5.04e−11 (1.19e5) |
| Epidermal HSV replication rate | HSV DNA copies per cell per day | p | 1.03e5 | 7.21e4 | 1.7e5 | 5.15e4 | 1.96e5 |
| Neuronal release rate | HSV DNA copies per day per genital tract | ϕ | 82 | 45 | 90 | 41 | 123 |
| Free-viral decay rate | Per day (half-life, hours) | c | 8.8 (1.9) | 7.0 (2.4) | 9.7 (1.7) | 6.2 (2.7) | 12.3 (1.4) |
| Maximal CD8+ T-cell expansion rate | Per day | θ | 2.84 | 1.85 | 3.27 | 1.85 | 5.25 |
| CD8+ T-cell decay rate | Per day (half-life, days) | δ | 1.47e−3 (471) | 1.12e−3 (619) | 1.69e−3 (409) | 6.64e−4 (1040) | 2.21e−3 (314) |
| CD8+ T-cell local recognition | Infected cells at which θ is half maximal | r | 42 | 30 | 47 | 4 | 74 |
| CD8+ regional codependence | 0 = no codependence, 1 =full codependence | ρ | 0.69 | 0.59 | 0.86 | 0.38 | 0.86 |
| Viral production lag | Days | ε | 0.96 | 0.53 | 1.1 | 0.34 | 1.1 |
Predictive model parameters for key model outcomes
| Single episode features | Long-term shedding features | |||
| Peak viral load | Duration | Shedding rate | Episode rate | |
| CD8+ T-cell density at reactivation site | −0.56 | −0.47 | NA | NA |
| Cell-associated HSV infectivity | — | 0.12 | — | 0.13 |
| Cell-free HSV infectivity | — | — | — | — |
| Epidermal cell replicate rate | 0.13 | 0.14 | −0.25 | −0.31 |
| Neuronal release rate | — | — | 0.43 | 0.55 |
| Free-viral decay rate | — | — | −0.2 | — |
| Maximal CD8+ T-cell expansion rate | −0.09 | — | 0.37 | 0.51 |
| CD8+ T-cell decay rate | — | 0.09 | — | −0.16 |
| CD8+ T-cell local recognition | — | — | — | — |
| CD8+ regional co-dependence | — | — | 0.32 | 0.34 |
| Viral production lag | — | — | 0.24 | 0.23 |
-
Partial correlation coefficients are listed only for parameters that are found to improve predictive effect on outcomes using Akaike information criteria models. Episode features are from 500 single episode simulations. Long-term shedding outcomes were measured over 10-years during 500 simulations.
Spatial model simulations that varied only according to duration of sampling (30 days, 60 days, 365 days, and 10 years)
| Simulation duration | Percent of time with HSV DNA > 150 copies per mL | Percent of time with lesions* present | Episodes per year | Lesions per year | |
| 30 day | Mean | 13.7 | 7.6 | 11 | 3.2 |
| Median | 3.4 | 0 | 12 | 0 | |
| Range | 0–82.8 | 0–58.8 | 0–36.5 | 0–24.3 | |
| 60 day | Mean | 19.0 | 9.8 | 13.4 | 4.4 |
| Median | 18.6 | 3.4 | 12 | 6 | |
| Range | 0–54.2 | 0–46.9 | 0–42.6 | 0–18 | |
| 365 day | Mean | 19.6 | 10.1 | 14.3 | 4.6 |
| Median | 19.9 | 9.9 | 14 | 4 | |
| Range | 7.1–36.8 | 2.3–22 | 8–24 | 1–9 | |
| 10 year | Mean | 17.5 | 9.1 | 14.5 | 4.3 |
| Median | 17.6 | 9.0 | 14.5 | 4.2 | |
| Range | 14.8–19.8 | 1.9–12.8 | 11.5–17.1 | 1.4–6 |
-
*
Lesions were defined as > 1 mm diameter ulcers.
-
Sixty simulations were performed at each of the sampling durations. Within shorter sampling duration simulations, lesion, and shedding frequency varied significantly, while ranges narrowed with prolonged sampling.
Mathematical models of HSV-2 pathogenesis
| Model | Equations (additions to previous model are denoted in bold) | Variables (model fitting variable) | New features |
| 1 (Schiffer et al., 2009; Schiffer et al., 2010) | S, I, E, Vi Vneu, | — | |
| 2 | S, I, E, Vi, Vneu, (Ve) | Cell-free and cell-associated particles | |
| 3 | S, I, E, Vneu, Vitot, (Vetot) | Concurrent plaques from cell-free particles | |
| 4* | S, I, E, Vneu, Vitot, (Vetot) | Spatial model |
-
*Model 4 has a parameter of regional CD8+ co-dependence (ρ) within each plaque-forming region. At episode onset within a region, the CD8+ density is adjusted to infer the spatial co-dependence of CD8 density from surrounding regions: Ei (time + 0.001) = (Ei × (1 − ρ)) + (Eavg × ρ) where Eavg is average of E from the 6 surrounding regions.
-
Models are described in the ‘Methods’. Units and values of each parameter in the optimized model are listed in Table 2. Variables include: S (susceptible skin cells), I (infected skin cells), E (CD8+ T cells), Vi (cell-associated HSV DNA particles), and Ve (cell-free HSV DNA particles).
Model fit to Cohort E
| Summary measure | Summed criteria scores | Best fitting score | AIC | |||||||
| Shedding frequency | Episode duration | First positive swab | Last positive swab | Peak positive swab | Median expansion | Median decay | Episode frequency | |||
| Model 1 | 2.13 | 5.59 | 0.05 | 0.08 | 0.24 | 0.07 | 0.43 | 0.01 | 8.61 | −50 |
| Model 2 | 1.26 | 8.33 | 0.38 | 0.34 | 0.22 | 0.04 | 0.09 | 0.03 | 10.69 | −41 |
| Model 3 | 0.25 | 3.75 | 0.31 | 0.22 | 0.61 | 0.05 | 0.50 | 0.22 | 5.92 | −62 |
| Model 4 solved for 10 parameters | 0.25 | 0.13 | 0.29 | 0.15 | 0.09 | 0.01 | 0.01 | 0.03 | 0.96 | −139 |
| Model 4 solved for 5 parameters | 0.39 | 0.32 | 0.44 | 0.21 | 0.09 | 0.04 | 0 | 0.18 | 1.67 | −125 |
-
Summed criteria scores measure the degree of fit for each model according the eight individual shedding episode features using a weighted sum of squares. Model 4 is the spatial model. Models 1–3 are described in the ‘Methods’. Best fitting score is a sum of all summed criteria scores for a particular model with lower scores indicating better fit. AIC: Akaike information criteria with lower scores indicating better fit.





