A role for PVRL4-driven cell–cell interactions in tumorigenesis
Figures
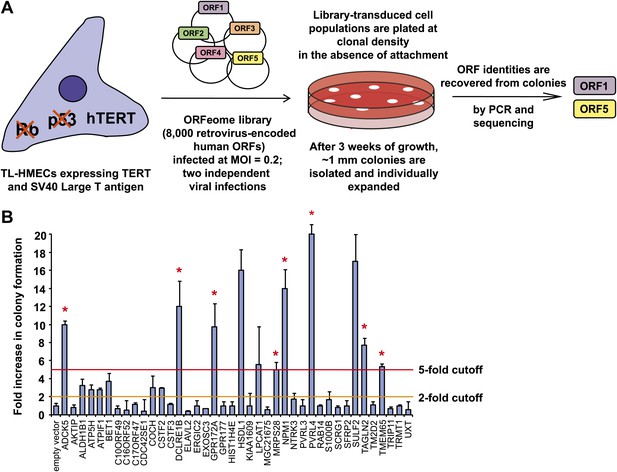
A genetic screen for drivers of anchorage-independent growth in human mammary epithelial cells.
(A) A schematic of the screen. TL-HMECs were transduced with the ORFeome library (8000 CMV promoter-driven human open reading frames [ORFs] in a retroviral vector) at a multiplicity of infection (MOI) of 0.2 and plated into semi-solid medium. Macroscopic colonies were isolated, individually expanded, and the identities of ORF inserts were determined by sequencing. (B) ORFs recovered from two independent screen replicates were individually transduced into TL-HMECs and plated into semi-solid medium. Colonies were counted and colony numbers were normalized to an empty vector-transduced sample. Asterisks denote strongly validated ORFs that localize to focal amplification peaks in at least one tumor subtype. Assays were performed in triplicate (error bars ± SD).
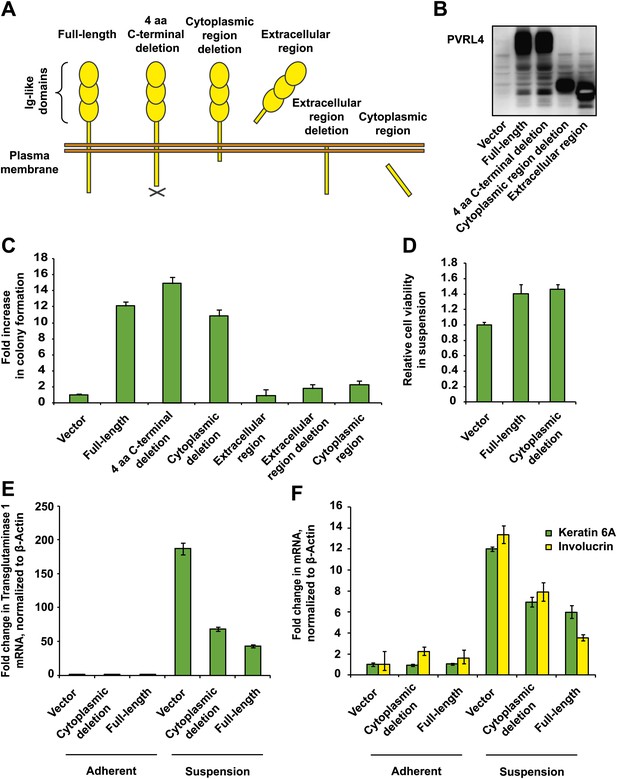
PVRL4-induced anchorage-independent colony formation is carried out through its extracellular region.
(A) and (B) A series of PVRL4 deletion constructs were designed and their expression confirmed by Western blot. (C) PVRL4 mutants from (A) were tested for their ability to induce anchorage-independent colony formation in triplicate (error bars ± SD). (D) Cells with full-length PVRL4 or the cytoplasmic region deletion mutant were assayed for viability under conditions of anchorage deprivation by measuring total ATP content in cells cultured on ultra-low attachment plates for 72 hr. Values were normalized to an empty vector-transduced sample. Assays were performed in triplicate (error bars ± SD). (E) and (F) TL-HMECs expressing empty vector, full-length PVRL4 or cytoplasmic region deletion mutant containing cells were cultured on tissue culture-treated (adherent) or ultra-low attachment (suspension) dishes for 72 hr. RNA was isolated and mRNA levels for TGM1 (E) and KRT6A and IVL (F) were measured by RT-qPCR. Transcript levels were normalized to β-actin. qPCR was performed in quadruplicate (error bars ± SD).
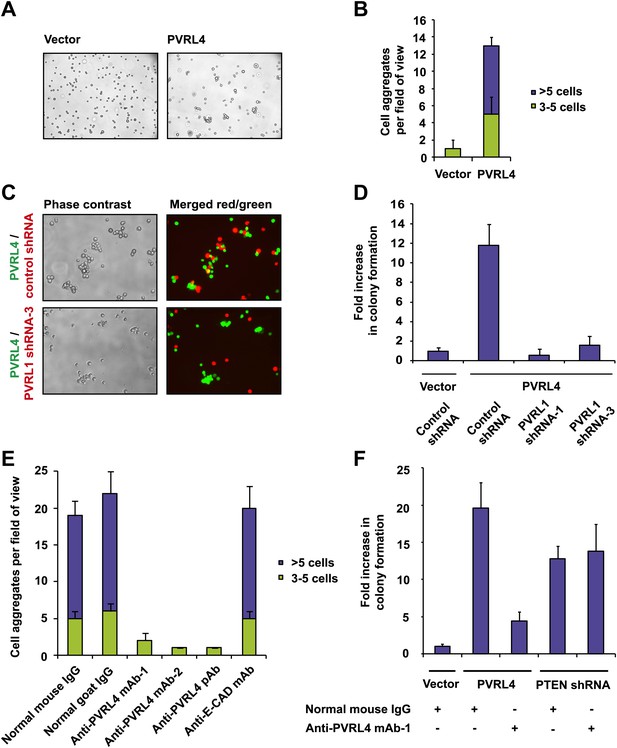
PVRL4 facilitates cell-to-cell attachment, inhibition of which suppresses anchorage-independence.
(A) and (B) PVRL4 promotes cell clustering of TL-HMECs. Cells were dissociated off the tissue culture surface with trypsin-free cell dissociation buffer and kept in suspension for 1 hr. Small (3–5 cells) and large (>5 cells) cell clusters per field of view were counted, n = 3 (error bars ± SD). (C) GFP-labeled PVRL4-expressing TL-HMECs were allowed to aggregate with dsRed-labeled cells expressing either a PVRL1-targeting shRNA or a control shRNA. Representative phase-contrast and fluorescent images (red and green channels superimposed) are shown. (D) PVRL4 was co-expressed with the indicated shRNAs and anchorage-independent colony formation in TL-HMECs was assayed. Values were normalized to an empty vector-transduced sample. Assays were performed in triplicate (error bars ± SD). (E) PVRL4-expressing TL-HMECs were assayed for clustering in the presence of the indicated antibodies or isotype controls. Cell clusters were quantified as before. (F) Anchorage-independent growth induced by PVRL4 or an shRNA against PTEN was assayed in the presence of PVRL4-targeting antibody or control IgG. Colony numbers were normalized to the control sample. Anchorage-independent colony formation assays were performed in triplicate (error bars ± SD).
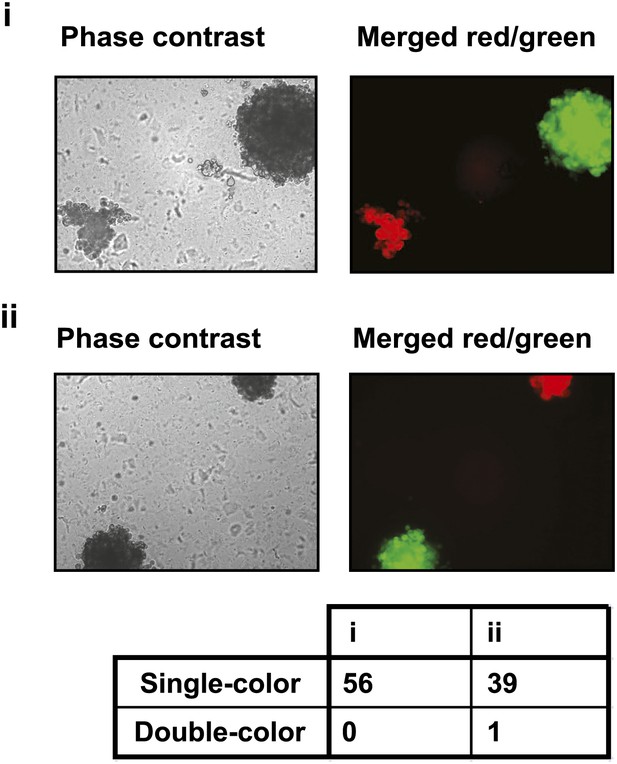
PVRL4-driven anchorage-independent colonies originate from single cells.
PVRL4 expressing TL-HMECs were stably transduced with dsRed or GFP and mixed in equal proportions, followed by (i) immediate plating into semi-solid medium, or (ii) co-culturing on an adherent surface for 2 d, followed by plating into semi-solid medium. Resulting colonies were visualized under a fluorescent microscope and each colony was assessed for the presence of red and green fluorescence. Representative phase-contrast and fluorescent images (red and green channels) are shown.
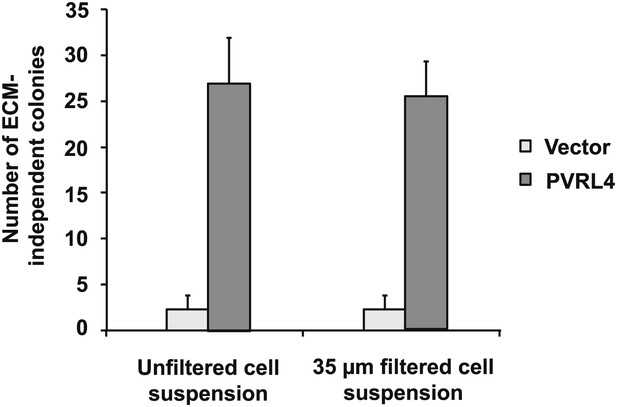
Potential preformed clusters of TL-HMECs do not contribute to anchorage-independent colony numbers.
The colony formation efficiency of TL-HMECs transduced with empty vector or vector expressing PVRL4 was compared between unfiltered cell suspensions and cell suspensions that were filtered through a 35 μm nylon mesh strainer prior to plating into methylcellulose. Anchorage-independent colony formation assays were performed in triplicate (error bars ± SD).
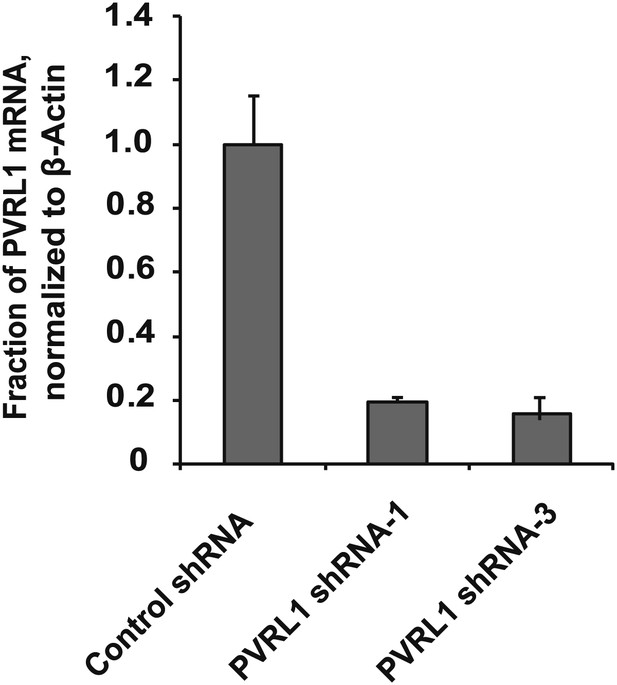
Depletion efficiency of individual anti-PVRL1 shRNAs.
efficiency of shRNA-mediated PVRL1 mRNA depletion was measured by RT-qPCR. PVRL1 transcript abundance was normalized to β-actin. qPCR was performed in quadruplicate (error bars ± SD).
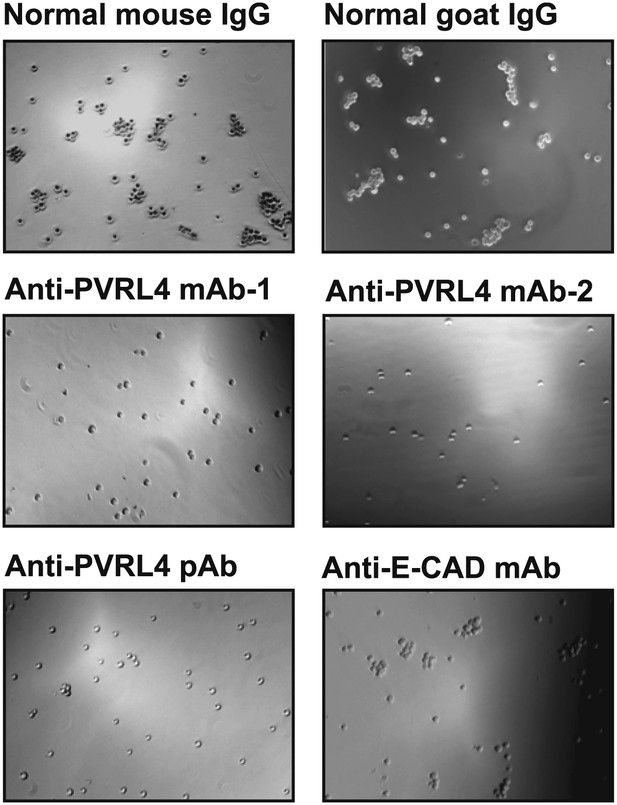
Anti-PVRL4 antibodies block PVRL4-driven cell–cell clustering.
PVRL4-expressing TL-HMECs were allowed to aggregate in the presence of the indicated antibodies or isotype controls. Representative images are shown.
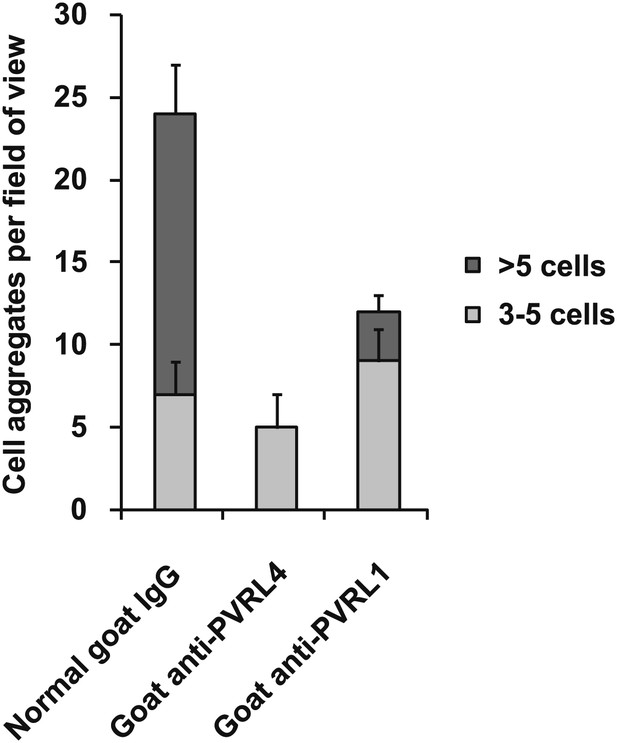
PVRL4-driven cell–cell clustering is inhibited by antibodies against PVRL1.
PVRL4-expressing TL-HMECs were assayed for clustering in the presence of the indicated antibodies or isotype controls. Cell clusters were quantified as before.
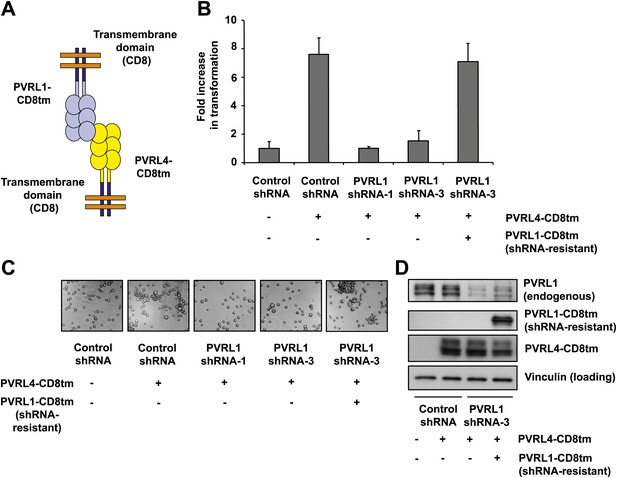
Expression of extracellular regions of PVRL4 and PVRL1 on the cell surface is sufficient for anchorage-independence.
(A) Schematics of chimeric constructs containing extracellular domains of PVRL4 or an shRNA-resistant version of PVRL1 fused to the transmembrane domain of CD8 (blue). (B) and (C) TL-HMECs were stably transduced with the indicated combinations of expression constructs and assayed for anchorage-independent growth (B) and clustering (C). Colony numbers were normalized to the control sample. Anchorage-independent colony formation assays were performed in triplicate (error bars ± SD). (D) Expression levels of endogenous and chimeric proteins were verified by Western blot.
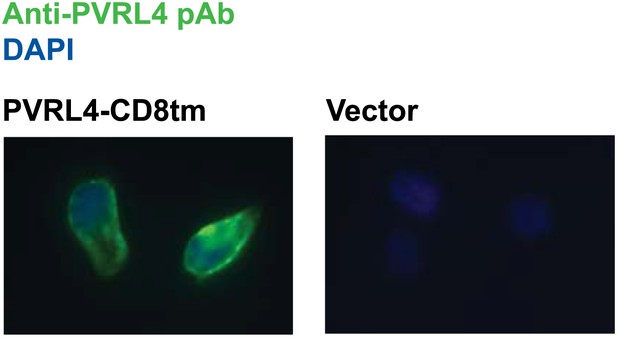
Plasma membrane localization of PVRL4-CD8tm construct in TL-HMECs.
TL-HMEC cells infected with PVRL4-CD8tm or empty vector were fixed with methanol and stained with goat polyclonal anti-PVRL4 antibody followed by anti-goat Alexa Fluor 488 secondary antibody.
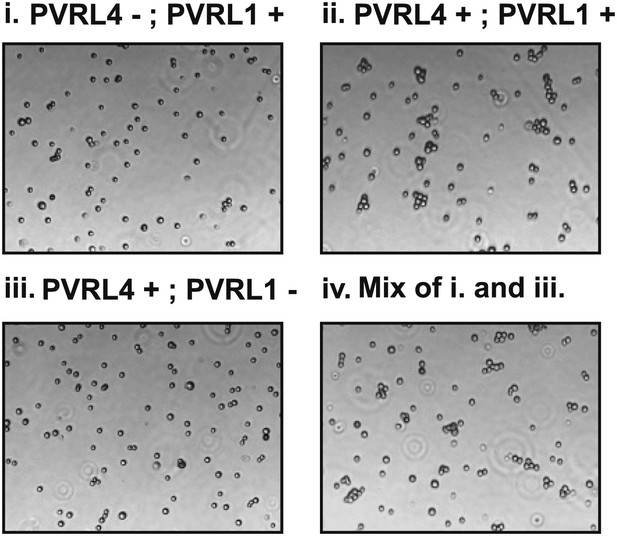
Cell–cell clustering is driven by PVRL4-PVRL1 trans-interactions between individual cells.
Clustering assays were performed with TL-HMECs expressing the following transgenes: (i) empty vector/control shRNA; (ii) PVRL4-CD8tm/control shRNA; (iii) PVRL4-CD8tm/anti-PVRL1 shRNA; and (iv) a 1:1 mixture of (iii) and (i).
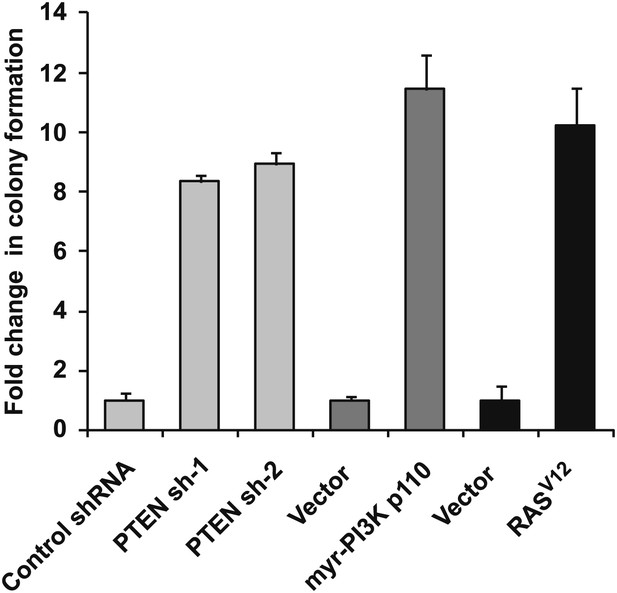
Expression of shRNA constructs against PTEN or constitutively active mutants of RAS and PI3K induces anchorage-independent growth.
TL-HMECs transduced with the indicated constructs were assayed for ability to induce anchorage-independent colony formation in triplicate (error bars ± SD).
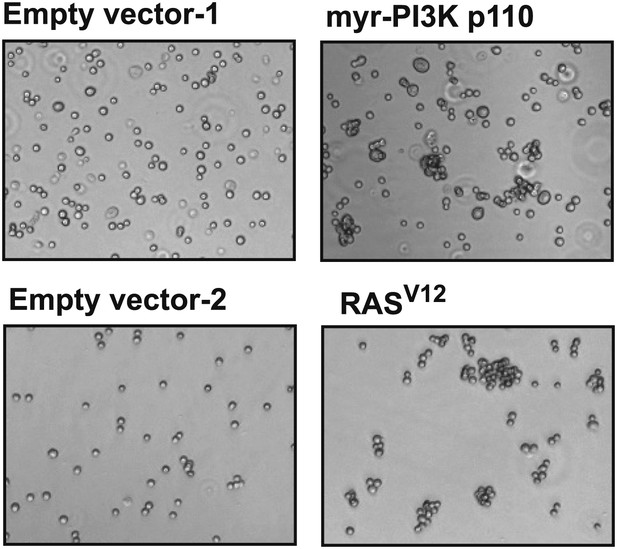
Constitutively active mutants of RAS and PI3K induce cell–cell clustering.
TL-HMECs were stably transduced with the indicated constructs and assayed for cell–cell clustering and colony formation in the absence of substratum anchorage. Representative clustering assay images are shown.
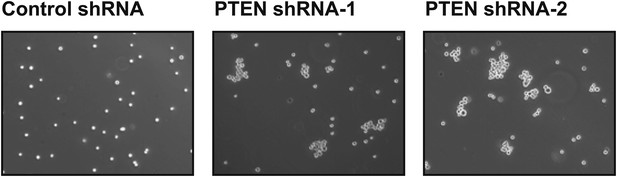
PTEN depletion induces cell–cell clustering.
TL-HMECs expressing two independent PTEN shRNAs or control shRNA were assayed for cell–cell clustering. Representative images are shown.
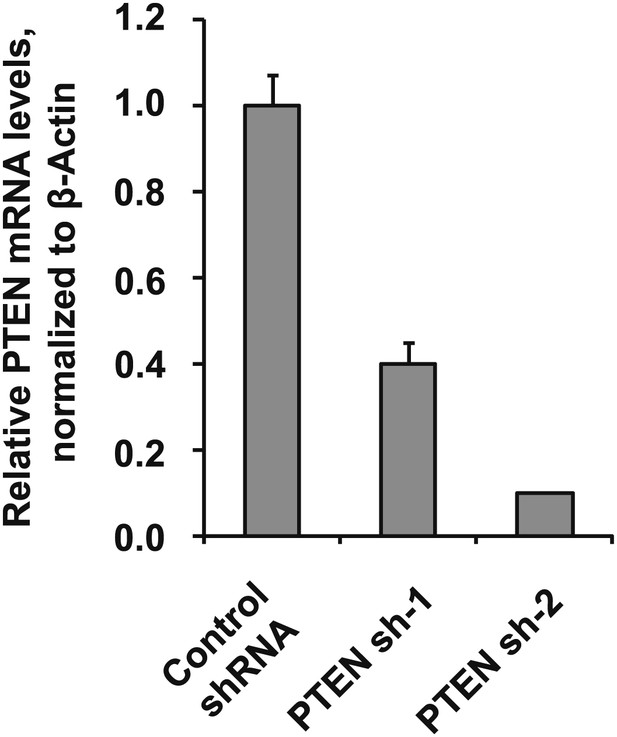
Depletion efficiency of individual anti-PTEN shRNAs.
The efficiency of PTEN mRNA depletion was measured by RT-qPCR. PTEN transcript abundance was normalized to β-actin. qPCR was performed in quadruplicate (error bars ± SD).
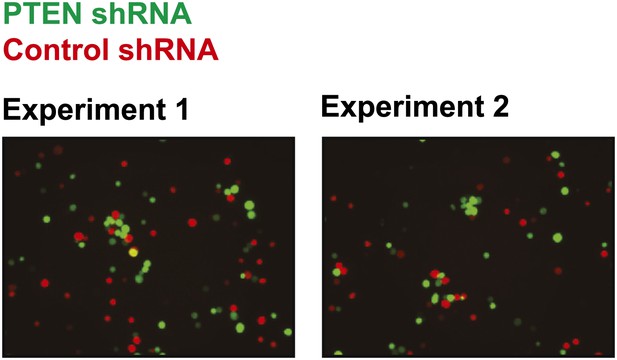
Cell–cell clustering induced by depletion of PTEN is heterotypic.
GFP-labeled PTEN-depleted TL-HMECs were allowed to aggregate with dsRed-labeled control shRNA-expressing cells. Images were taken and processed as before. Representative images are shown.
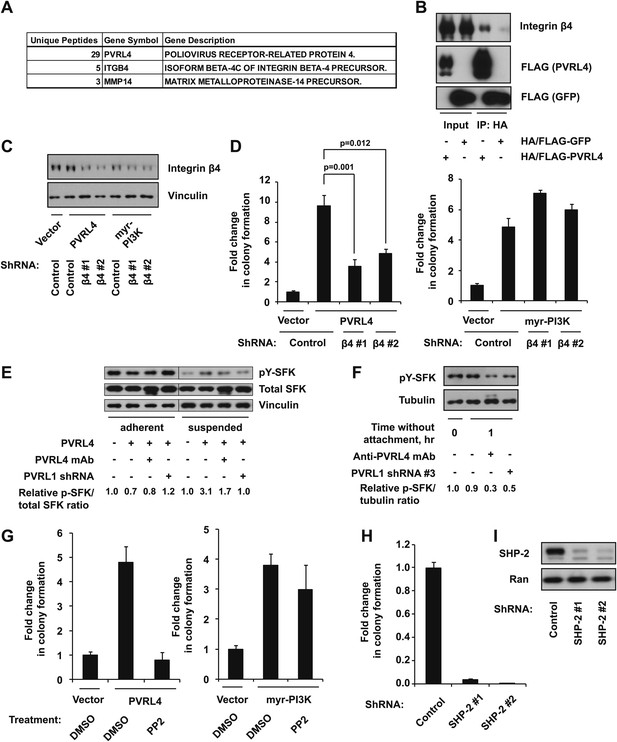
PVRL4-driven cell-to-cell attachment promotes anchorage-independence via integrin β4-associated signaling.
(A) Cell surface-localized proteins interacting with HA/FLAG-tagged PVRL4, but not with HA/FLAG-tagged GFP, as determined by mass spectrometry. (B) TL-HMECs expressing HA/FLAG tagged PVRL4 or HA/FLAG-tagged GFP were detached from the adherent surface with the enzyme-free cell dissociation buffer and incubated in suspension for 1 hr. Immunoprecipitations were performed with HA beads, followed by Western blot with FLAG and integrin β4 antibody. (C) TL-HMECs expressing vector control, PVRL4, or myr-PI3K were stably transduced with the indicated shRNA constructs and integrin β4 levels were assayed by Western blot. (D) TL-HMECs from (C) were assayed for anchorage-independent colony formation. Colony numbers were normalized to the vector control sample. Assays were performed in triplicate (error bars ± SD). (E) TL-HMECs stably transduced with the indicated constructs were detached from the adherent surface with enzyme-free cell dissociation buffer and incubated in 0.5% methylcellulose in suspension for 6 hr or cultured on an adherent surface for 48 hr. Levels of pY416-SFK (Src family kinases), total SFK, and vinculin loading control were measured by Western blot. Band intensity was measured with ImageJ software. (F) PVRL4-expressing TL-HMEC cells transduced with control or anti-PVRL1 shRNA were incubated in suspension in the conditions indicated. Levels of pY416-SFK (Src family kinases) and tubulin loading control were measured by Western blot. Band intensity was measured with ImageJ software. (G) TL-HMECs stably expressing PVRL4 or control vector were assayed for anchorage-independent colony formation in the presence of PP2 or vehicle control. Colony numbers were normalized to the vector sample. Assays were performed in triplicate (error bars ± SD). (H) TL-HMECs expressing PVRL4 were stably transduced with the indicated shRNA constructs and assayed for anchorage-independent colony formation. Colony numbers were normalized to the vector control sample. Assays were performed in triplicate (error bars ± SD). (I) SHP-2 levels were assayed by Western blot in TL-HMEC lysates from (H).
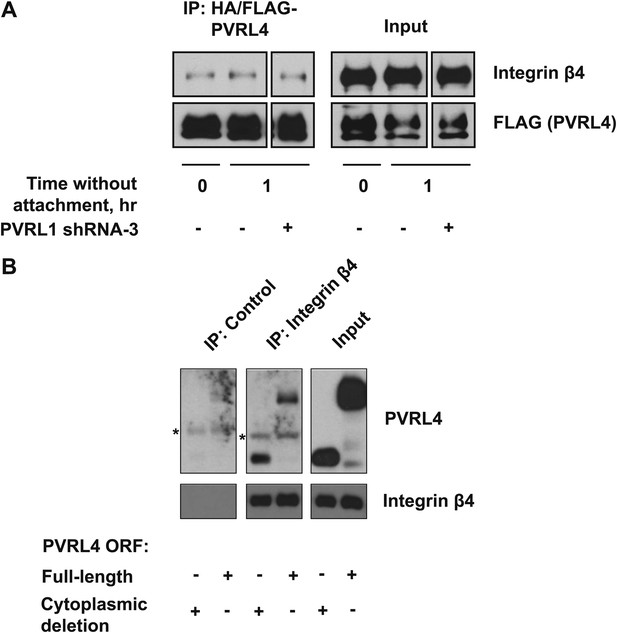
Interaction of PVRL4 with integrin β4.
(A) Anti-HA beads were used to immunoprecipitate HA/FLAG-PVRL4 from the indicated lysates. Immunoprecipitates were blotted with anti-integrin β4 and anti-FLAG antibodies. (B) Immunoprecipitations with anti-integrin β4 antibodies or with a control IgG were performed from TL-HMEC lysates expressing either full-length PVRL4 or its cytoplasmic deletion mutant. Immunoprecipitates and input lysates were blotted with anti-integrin β4 and anti-PVRL4 antibodies. Asterisks denote heavy chains of control and anti-integrin β4 antibodies, which cross-react with an anti-goat secondary antibody.
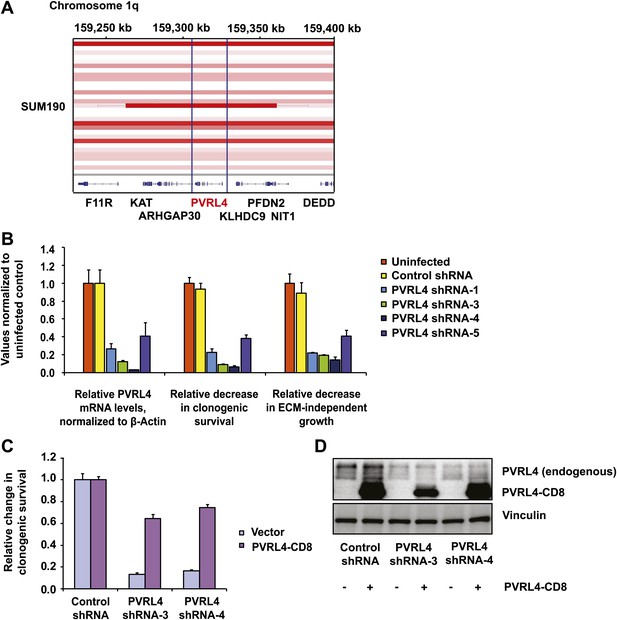
PVRL4 is amplified in breast cancer and is essential for the transformed phenotype of cancer cells.
(A) A view from the integrated Genome Viewer program showing focal amplification of the PVRL4 locus in SUM190 cells. The degree of amplification is denoted by the intensity of the color. (B) PVRL4 mRNA was stably depleted from SUM190 cells by four independent shRNAs. Transcript levels were measured by RT-qPCR and normalized to β-actin. qPCR was performed in quadruplicate (error bars ± SD). PVRL4-depleted and control cells were assayed for clonogenic survival and anchorage-independent colony formation. Assays were performed in triplicate (error bars ± SD). All values were normalized to the uninfected control sample. ECM: extracellular matrix. (C) The PVRL4-CD8 chimeric construct was used to rescue the defect in clonogenic survival observed with RNAi-mediated PVRL4 depletion. Assays were performed in triplicate (error bars ± SD). Colony numbers were normalized to the control shRNA sample. (D) Expression levels of endogenous and chimeric proteins were verified by Western blot.
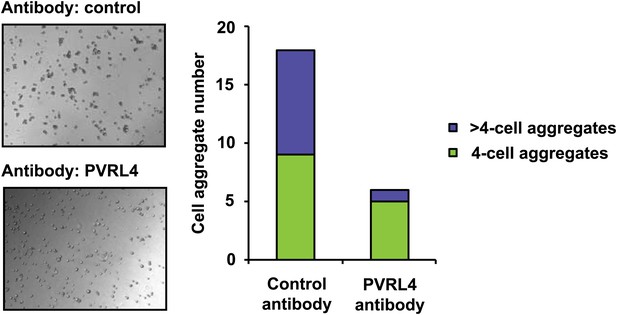
PVRL4 induces clustering of SUM190 cells which is blocked by antibodies against PVRL4.
SUM190 cells were assayed for cell–cell clustering in the presence of the indicated antibodies. Four-cell clusters and clusters with more than four cells from representative images were scored separately.
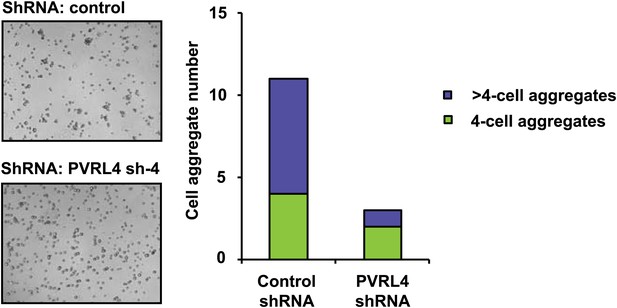
PVRL4 induces clustering of SUM190 cells which is blocked by RNAi against PVRL4.
SUM190 cells were assayed for cell–cell clustering in the presence of the indicated shRNAs. Four-cell clusters and clusters with more than four cells from representative images were scored separately.
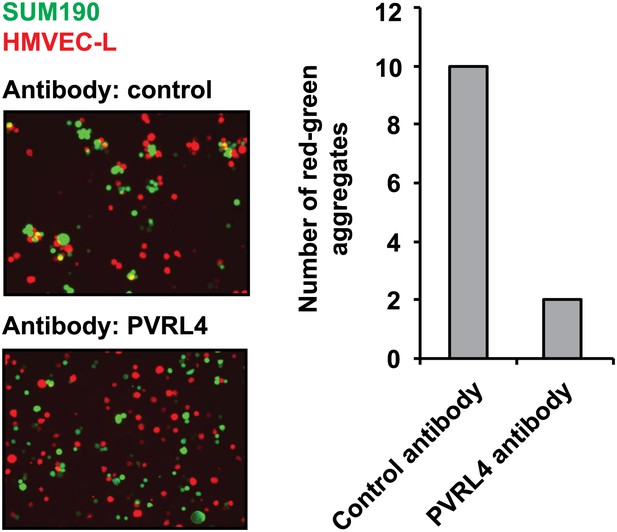
PVRL4 induces attachment of SUM190 cells to microvascular endothelial cells which is blocked by antibodies against PVRL4.
SUM190 cells (GFP) were assayed for heterotypic clustering with HMVEC-L cells (dsRed) in the presence of the indicated antibodies. Clusters with at least three cells incorporating both green and red cells from representative images were counted.
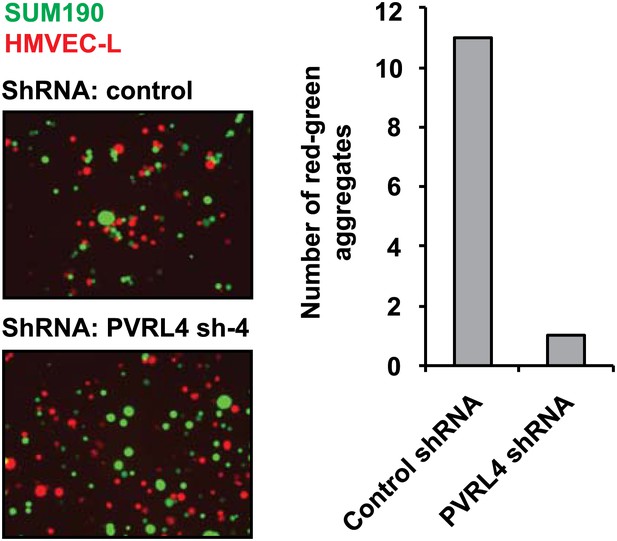
PVRL4 induces attachment of SUM190 cells to microvascular endothelial cells which is blocked by RNAi against PVRL4.
SUM190 cells (GFP) were assayed for heterotypic clustering with HMVEC-L cells (dsRed) in the presence of the indicated shRNAs. Clusters with at least three cells incorporating both green and red cells from representative images were counted.
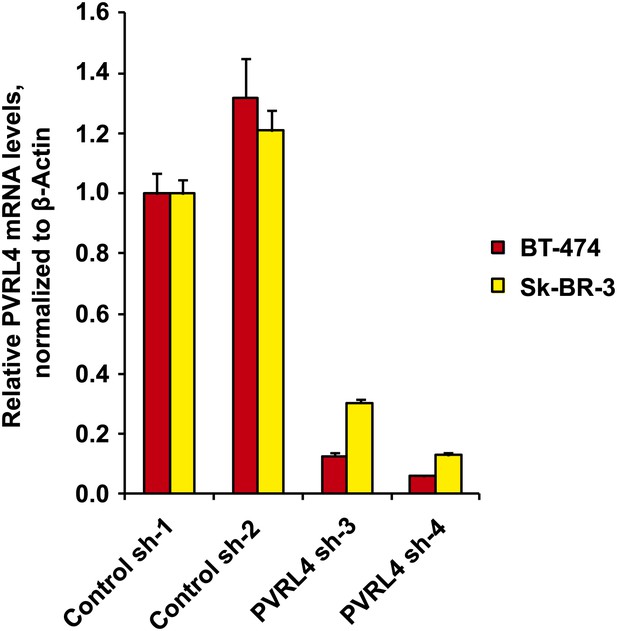
Stable depletion of PVRL4 transcript in BT-474 and Sk-BR-3 cell lines.
PVRL4 transcript levels were measured by RT-qPCR and normalized to β-actin. qPCR was performed in quadruplicate (error bars ± SD).
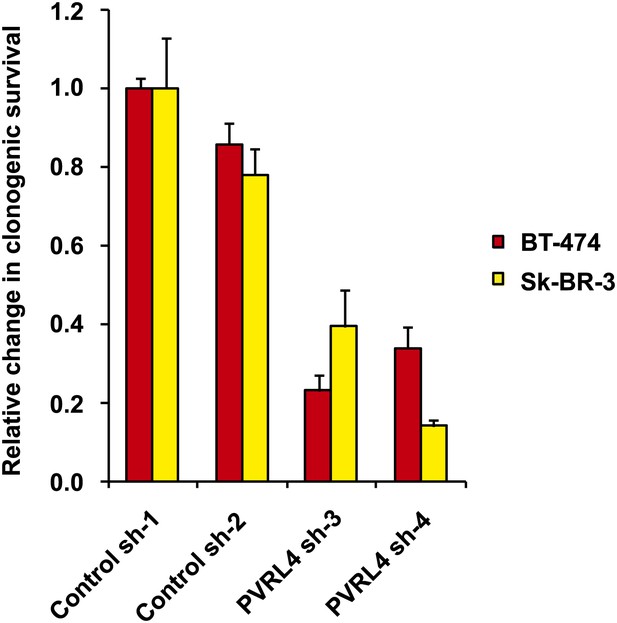
PVRL4 depletion affects clonogenic growth of BT-474 and Sk-BR-3 cell lines.
The clonogenic potential of the indicated cell lines in the presence of control or PVRL4 shRNA constructs was assessed. Assays were performed in triplicate (error bars ± SD). Colony numbers were normalized to the control shRNA sample.
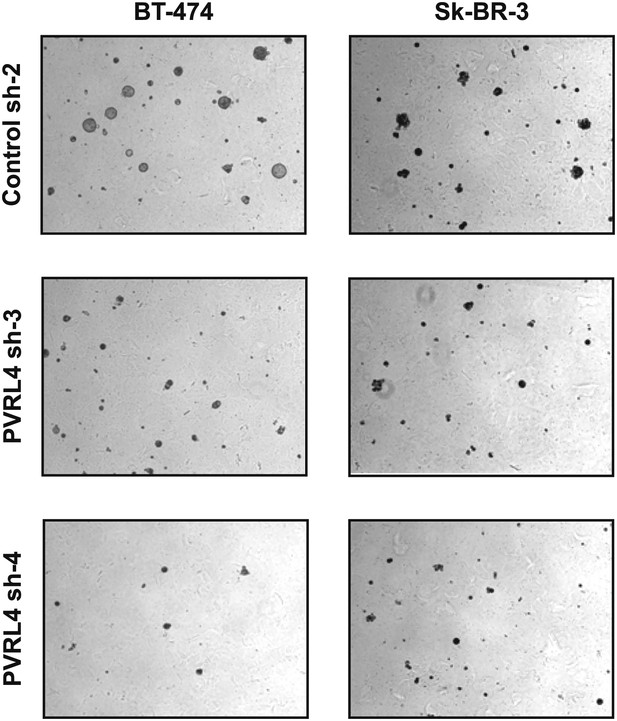
PVRL4 depletion affects anchorage-independent growth of BT-474 and Sk-BR-3 cell lines.
Cell line growth in methylcellulose-containing media on an ultra-low attachment surface was assessed in the presence of control or PVRL4 shRNA constructs. Representative images are shown.
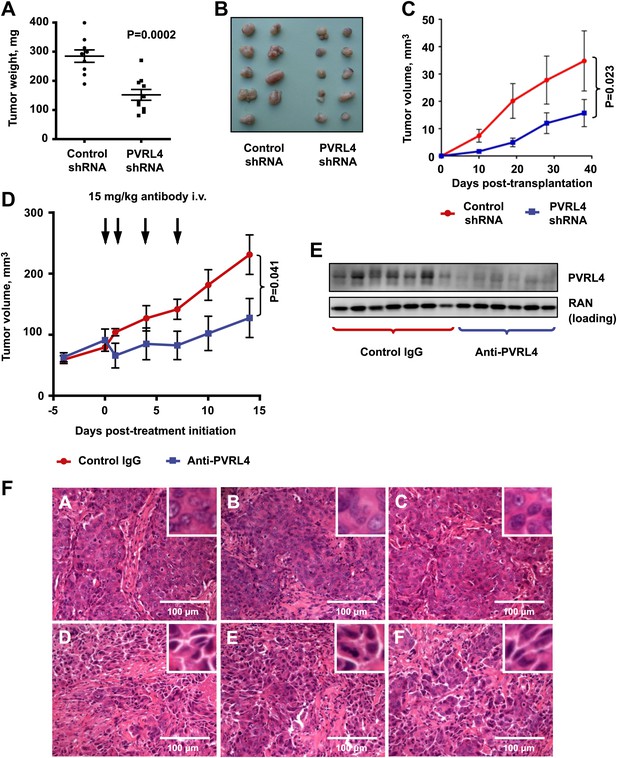
Targeting PVRL4 inhibits tumor growth.
(A) and (B) Female nude mice were injected into their mammary fat pads with SUM190 cells expressing PVRL4-targeted or control shRNA (n = 10 per group, error bars ± SEM). The resulting tumors were excised, scaled (A), and photographed (B). (C) SUM185 cells were stably transduced with PVRL4-targeted or control shRNA and injected into the mammary fat pads of female nude mice (n = 10 per group, error bars ± SEM). Tumor volume was measured with calipers at the indicated time points. (D) Female nude mice with ∼50 mm3 SUM190-eGFP xenografts were randomized into two cohorts (n = 7 per group) and injected with anti-PVRL4 monoclonal antibodies or control IgG on the indicated days. Tumor volume was measured with calipers (error bars ± SEM). (E) Levels of PVRL4 protein were measured in tumor lysates from anti-PVRL4 antibody or control-treated mice, 7 days after the last treatment. (F) Tumor sections from control IgG (A–C) or anti-PVRL4 antibody-treated (D–F) mice were stained with hematoxylin/eosin and photographed. Representative images are shown.
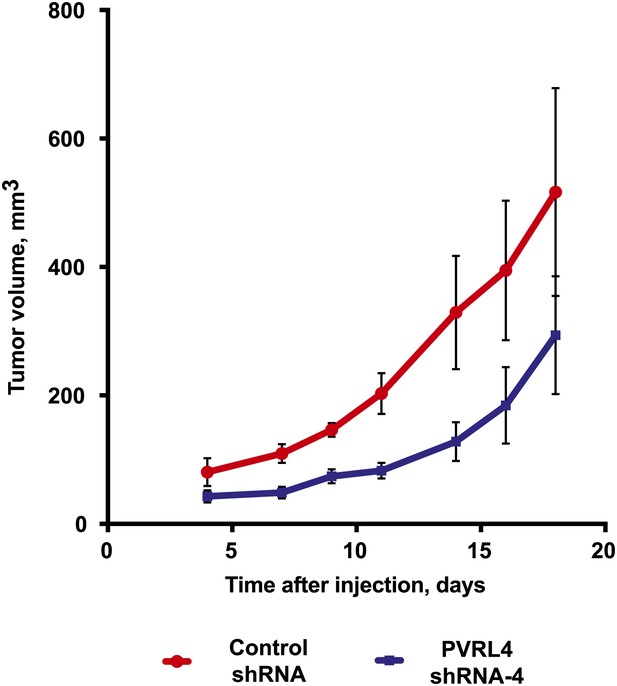
PVRL4 depletion inhibits BT-474 xenograft growth.
BT-474 cells were transduced with the indicated shRNA constructs and injected subcutaneously into female nude mice (N = 10 per group). Slow-release estrogen pellets were implanted into mice 72 hr prior to cell line injection. Tumor growth was measured at the indicated time points (error bars: SEM).
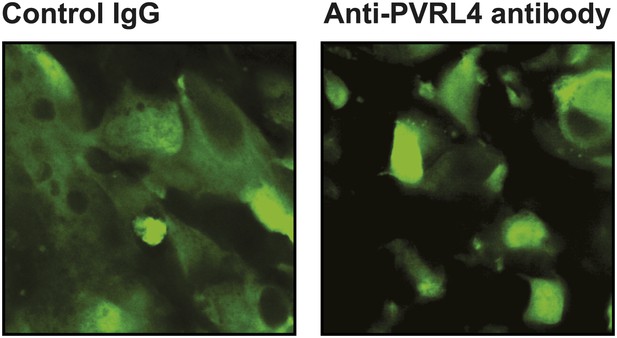
Anti-PVRL4 antibodies disrupt cell–cell contacts in xenografts in vivo.
Freshly explanted tumors from control IgG or anti-PVRL4 antibody-treated mice (N = 3 per group) were visualized using a two-photon confocal microscope. Representative images are shown.
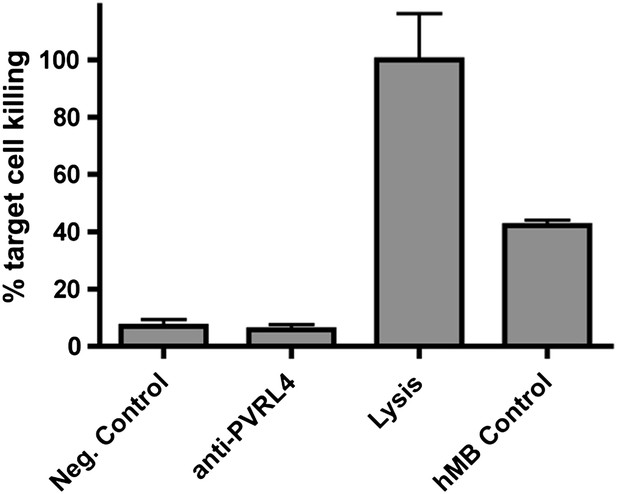
Anti-PVRL4 antibodies do not induce ADCC in vitro.
Europium-labeled SUM190 cells were incubated with fresh human NK cells in the presence of an isotype control or anti-PVRL4 antibody, and the degree of lysis was measured by the DELFIA europium assay. The maximum signal was determined by a complete lysis of labeled SUM190 cells in DELFIA lysis buffer. As a positive control, hMB humanized mouse lymphoma cells were mixed with effector cells in the presence of ADCC-competent anti-CD52 antibody. ADCC: antibody-dependent cytotoxicity.
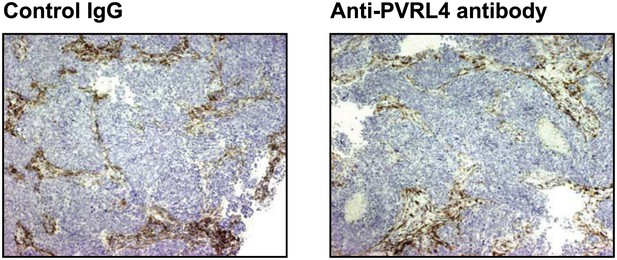
Anti-PVRL4 antibody treatment does not affect the degree of macrophage infiltration into SUM190 xenografts.
Paraffin-embedded sections of SUM190 xenografts from mice treated with either control IgG or anti-PVRL4 antibody were stained with anti-mouse F4/80 antibody. Representative images are shown.
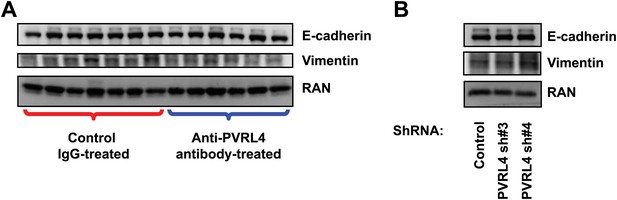
Inhibition of PVRL4 by antibodies or by RNAi does not affect expression of EMT markers.
(A) Tumor lysates from control antibody- or anti-PVRL4 antibody-treated mice were blotted for E-cadherin and vimentin. (B) PVRL4 was stably depleted by two independent shRNAs in SUM190 cells and lysates were blotted for E-cadherin and vimentin.
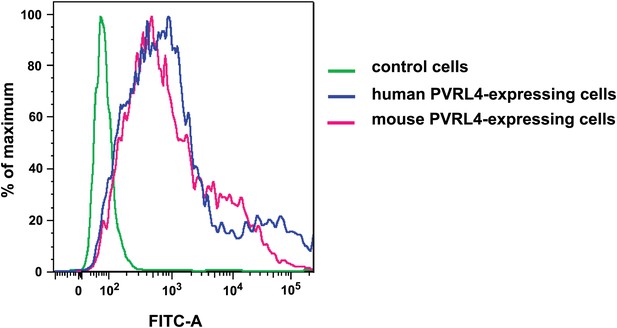
Anti-PVRL4 antibodies recognize both human and mouse epitopes.
293T cells were transfected with empty pQCXIN (green line), pQCXIN-human PVRL4 (blue line), or pCMV-SPORT6-mouse PVRL4 (magenta line). Live-cell FACS was performed with mouse anti-human PVRL4 antibody followed by anti-mouse secondary antibody conjugated to Alexa Fluor 488 fluorophore. The FITC-A fluorescent signal for three labeled cell populations is shown.
Additional files
-
Supplementary file 1
(A) List of candidate ORFs from the anchorage-independence screen. (B) List of validated ORFs which localize to statistically defined peaks of focal amplification in human tumors. (C) PVRL4 deletion series. (D) Sequences of shRNAs used in the study. (E) Sequences of qPCR primers used in the study.
- https://doi.org/10.7554/eLife.00358.036





