LIN-12/Notch signaling instructs postsynaptic muscle arm development by regulating UNC-40/DCC and MADD-2 in Caenorhabditis elegans
Figures
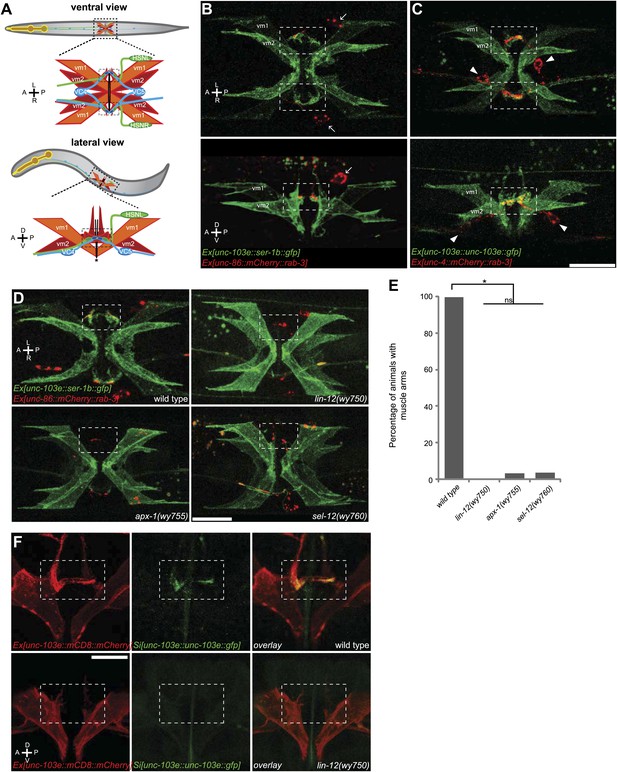
The muscle arms of type 2 vulval muscles (vm2) are missing in lin-12(wy750), apx-1(wy755) and sel-12(wy760) mutants.
(A) An illustration showing ventral (top) and lateral (bottom) views of C. elegans egg-laying circuit. vm1 (orange) and vm2 (red) are vulval muscles. HSN (green) and VC4/5 (blue) are presynaptic motoneurons. Each of the four vm2 cells extends a dendritic spine-like muscle arm laterally, on which it receives synapses with HSN and VC neurons (boxed areas). Top row of (A–C) are ventral views; bottom row of (A–C) are left lateral views. Anterior (A), posterior (P), left (L), right (R), dorsal (D), ventral (V). (B) Ventral (top) and lateral (bottom) views of vulval muscles labeled by SER-1B::GFP transgene in a young adult. HSN neurons are labeled by a mCherry::RAB-3 transgene. The boxed areas indicate synaptic regions. Arrows indicate the HSN cell bodies. Note that the laterally extended muscle arms on vm2 cells are colocalized with the HSN presynaptic specializations. (C) Ventral (top) and lateral (bottom) views of vulval muscles labeled by a UNC-103E::GFP transgene and VC4/5 neurons labeled by a mCherry::RAB-3 transgene. The boxed areas indicate synaptic regions. Arrowheads indicate cell bodies of VC4 and VC5. Note that the laterally extended muscle arms on vm2 cells are colocalized with the VC4/5 presynaptic specializations. Scale bar is 20 μm. (D) Ventral views of the vulval muscles and presynaptic regions of HSN neurons in wild-type, lin-12(wy750), apx-1(wy755) and sel-12(wy760) animals. The boxed areas indicate synaptic regions. Note the absence of vm2 muscle arms in mutants. Scale bar is 20 μm. (E) Quantification of the vm2 muscle arm defects in wild-type, lin-12(wy750), apx-1(wy755) and sel-12(wy760) animals. * p<0.0001, n.s. no significant difference, chi-squared test, n = 90–114 animals. (F) High magnification lateral views of the synaptic region in wild type and lin-12(wy750). vm2 muscle arms are visualized by a single-copy UNC-103E::GFP transgene. Vulval muscle morphology is labeled by mCD8::mCherry. The boxed areas indicate synaptic regions. Scale bar is 10 μm.
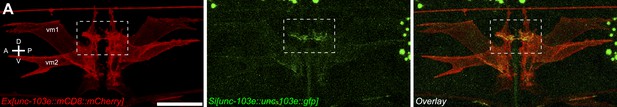
vm2 muscle arms are the postsynaptic specializations.
(A) Lateral views of the vulval muscles. The vulval muscles are labeled by mCD8::mCherry (top). vm2 muscles arms are labeled by a single-copy unc-103e::gfp transgene (middle). Note that the UNC-103E::GFP is enriched on the muscle arms. Boxed areas indicate synaptic regions. Anterior (A), posterior (P), dorsal (D), ventral (V). Scale bar is 20 μm.
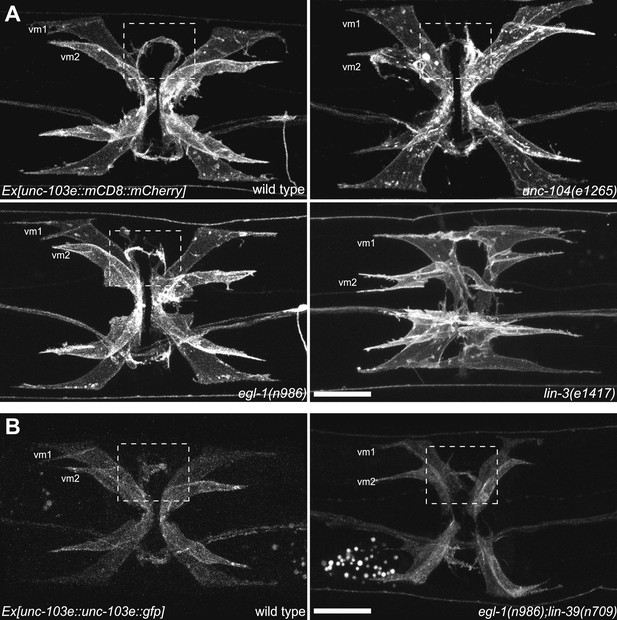
vm2 muscle arm development is a guidance event independent of presynaptic specialization.
(A) Ventral views of the vulval muscle morphology in wild-type, unc-104(e1265), egl-1(n986) and lin-3(e1417) mutant animals. Boxed areas indicate synaptic regions. Scale bar is 20 μm. (B) Ventral views of the vulval muscle morphology in wild-type (left) and egl-1(n986); lin-39(n709) (right) mutant animals. Boxed areas indicate synaptic regions. Scale bar is 20 μm.
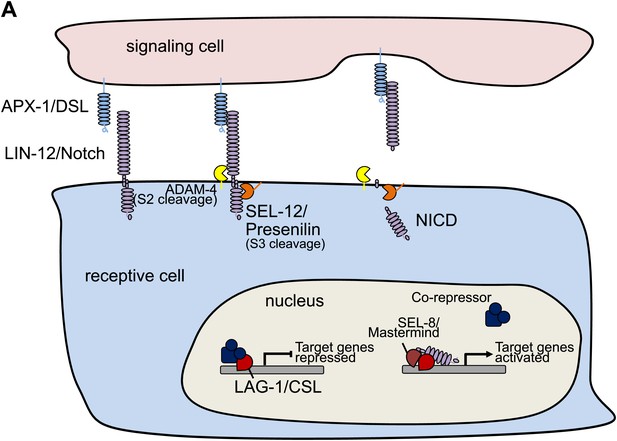
Schematic model of LIN-12/Notch signaling pathway.
(A) Schematic model showing the core LIN-12/Notch signaling pathway. The binding of the ligand APX-1/DSL to the receptor LIN-12/Notch triggers proteolytic cleavages. SEL-12/Presenilin releases the Notch intracellular domain (NICD), which enters the cell nucleus and interacts with LAG-1/CSL and other transcription factors to activate target gene transcription.
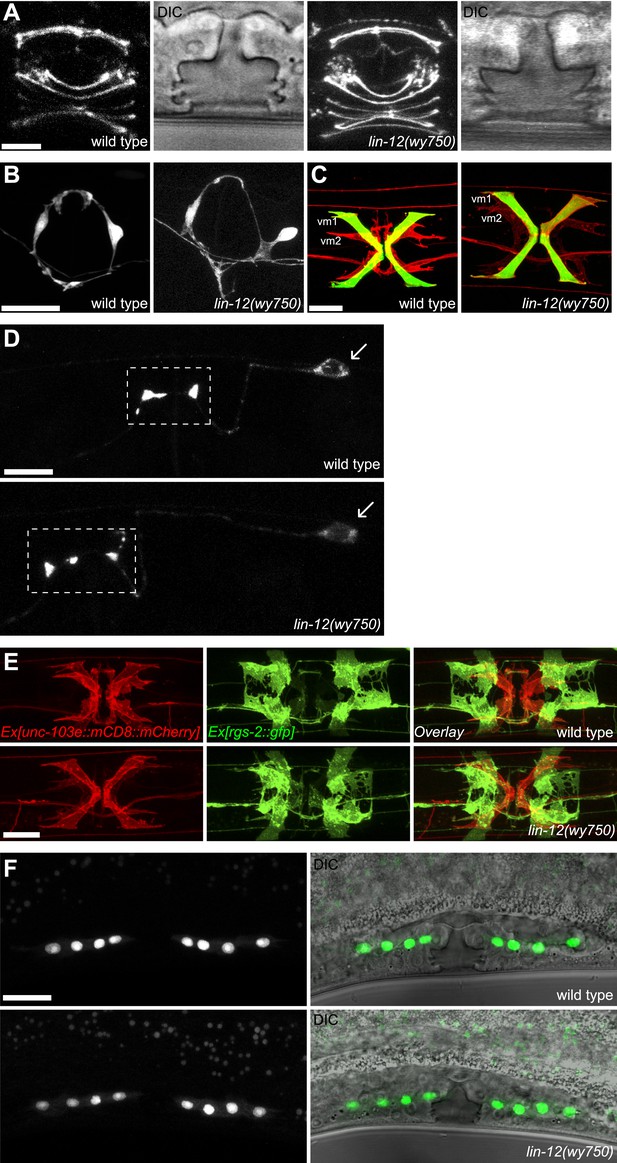
lin-12(wy750) has specific vm2 muscle arm defects.
(A) Vulval morphology in wild-type (left two panels) and lin-12(wy750) (right two panels) animals. Vulval morphogenesis is examined by visualizing the apical boundaries of vulval epithelial cells with AJM-1::GFP or by DIC microscopy. Scale bar is 5 μm. (B) VC4 and VC5 axons in wild-type (left) and lin-12(wy750) (right) animals. VC axons are labeled with CAT-1::GFP. Scale bar is 20 μm. (C) Visualization of vm1 and vm2 in wild-type (left) and lin-12(wy750) (right) animals. Vulval muscles (both vm1 and vm2) are labeled with mCD8::mCherry. egl-15::GFP labels vm1 but not vm2 in both wild type and lin-12(wy750) animals. Scale bar is 20 μm. (D) HSN presynapstic regions in wild-type (top) and lin-12(wy750) (bottom) animals. HSN presynaptic regions are labeled by SNB-1::YFP. Boxed areas indicate synaptic regions. Arrows indicate the HSN cell bodies. Scale bar is 10 μm. (E) Visualizations of undifferentiated M lineage cells in wild-type (top) and lin-12(wy750) (bottom) mutants by hlh-8::GFP in mid-L4 animals (lateral view). Note that the pattern of the M lineage cells do not change in the lin-12(wy750) mutant. Scale bar is 20 μm. (F) Ventral view of vulval muscles and uterine muscles in wild-type (up) and lin-12(wy750) (down) animals. Vulval muscles (both vm1 and vm2) are labeled with mCD8::mCherry. rgs-2::GFP labels uterine muscles in both wild type and lin-12(wy750) animals. Note that the uterine muscle differentiation and morphology do not change in lin-12(wy750) mutant. Scale bar is 10 μm.
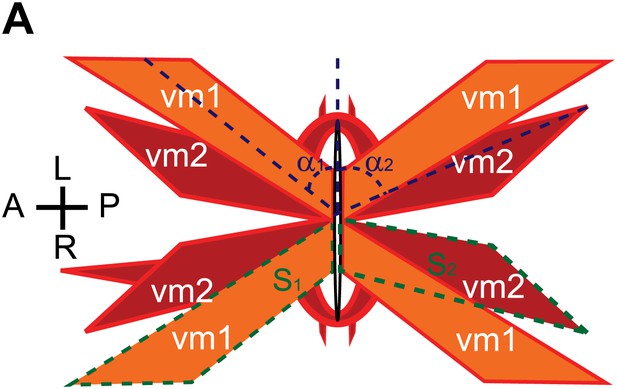
Schematic demonstrations of the quantitative measurements of vulval muscle morphology.
(A) Schematic demonstration of vulval muscles. The intersection angles between vulval opening and vulval muscles are shown with dashed blue lines. vm1 and vm2 are delineated by dashed green lines.
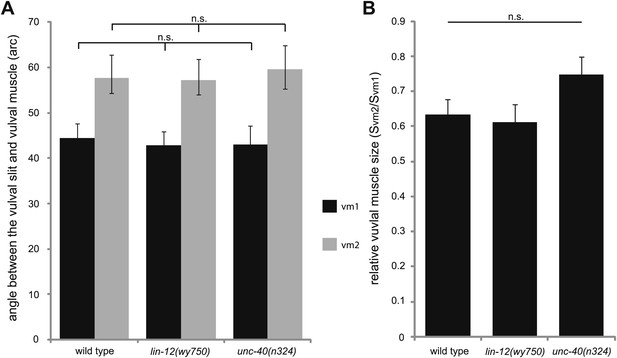
Quantifications of the vulval muscle morphology.
(A) Quantification of the intersection angles between vulval opening and vulval muscles with different genotypes indicated on the X-axis. n.s., no significant difference, t test, n = 48–80 pieces of vulval muscles. (B) Quantification of the relative size of vm2 to vm1 (Svm2/Svm1) with different genotypes indicated on the X-axis. n.s., no significant difference, t test, n = 48–80 pairs of vulval muscles.
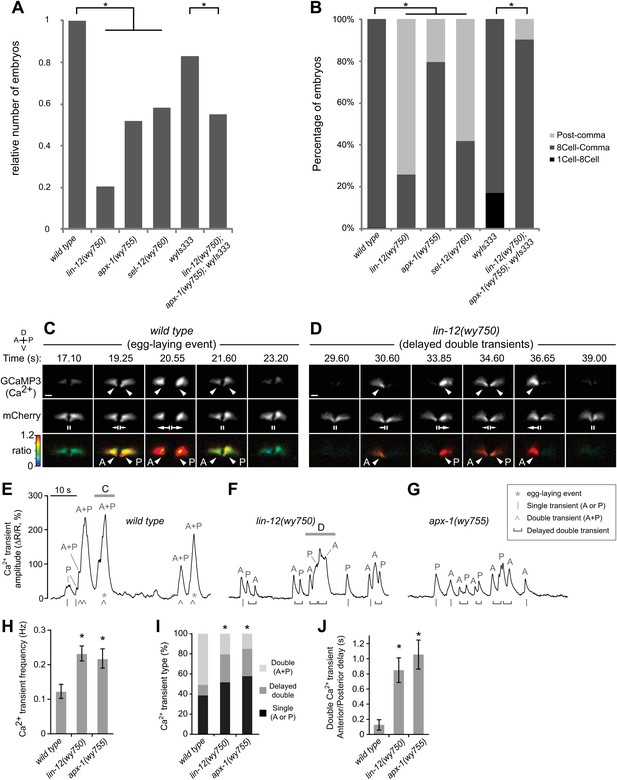
Muscle arm defective mutants have egg-laying behavioral and physiological defects.
(A) Relative numbers of embryos freshly-laid by wild-type and mutant animals. wyIs333 is the marker that double-labels the HSN presynaptic specializations and vulval muscles. *p<0.0001, Fisher's exact test, n = 62–303 embryos. (B) Percentage of different stages of embryos freshly-laid by wild-type and mutants animals. *p<0.0001, Fisher's exact test, n = 62–303 embryos. (C) and (D) Ratiometric Ca2+ imaging in the vulval muscles in behaving wild-type (C) and lin-12(wy750) (D) animals. GCaMP3 (top) and mCherry (middle) were co-expressed in the vulval muscles and the GCaMP3/mCherry fluorescence ratio (bottom) was used to record Ca2+ transients (arrowheads). Time points are shown from Video 1 (Wild type) and Video 2 (lin-12(wy750) mutant). Vertical lines (II) indicate the vulval muscles at rest, and arrows indicate vulval muscle twitches (small) and egg-laying (large) contractions. Anterior (A), posterior (P), left (L), right (R), dorsal (D), ventral (V). Scale bars are 10 μm. (E–G) traces of vulval muscle GCaMP3/mCherry ratio change (ΔR/R) from the same wild-type (E) and lin-12(wy750) mutant (F) animals shown above (horizontal bars) and in the apx-1(wy755) mutant (G). Also indicated are egg-laying events (*), single transients limited to the anterior or posterior (A or P) vulval muscles (|), double transients occurring simultaneously in both anterior and posterior (A + P) vulval muscles (^), or delayed double transient where anterior and posterior transients within a body bend are separated by a visually discernible interval (horizontal bracket). (H–J) Quantitations of vulval muscle Ca2+ signaling (6-min recording per animal, 11 or 12 animals per genotype; error bars indicate 95% confidence intervals). (H) lin-12(wy750) and apx-1(wy755) mutants have more frequent Ca2+ transients than does the wild type. *p<0.0001, one-way ANOVA, n = 215–488. (I) Fewer synchronous double (A + P) Ca2+ transients in lin-12(wy750) and apx-1(wy755) mutants. *p<0.0001, chi-squared test, n = 200–305. (J) Consecutive anterior and posterior Ca2+ transients are delayed in lin-12(wy750) and apx-1(wy755) mutants. *p<0.0001; one-way ANOVA, n = 123–190.
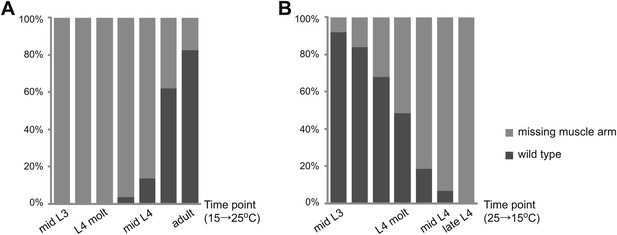
Temporal requirement of LIN-12 in vm2 muscle arm formation.
(A) and (B) Temperature up-shift (A) and down-shift (B) experiments of lin-12(n676n930). The percentages of animals with normal or defective muscle arm phenotype are indicated by dark or light gray bars, respectively. The time points when animals are shifted are shown on X-axis.
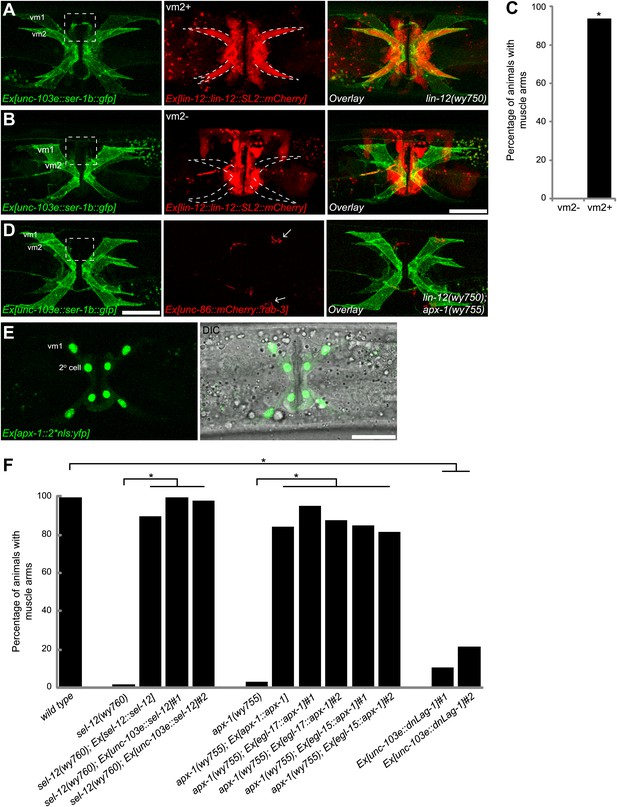
Cell-autonomous requirement of LIN-12 in vm2.
(A) and (B) Representative images of lin-12(wy750) mutant animals with non-mosaic (A) and mosaic (B) expression of lin-12::SL2::mCherry. Left column shows the vulval muscles labeled by SER-1B::GFP. Middle column shows the lin-12 expression patterns. Right column shows the overlaid images. Note that the complete lin-12 expression (vm2+) rescues muscle arm defects of lin-12(wy750) (A). Lacking of lin-12 expression in vm2 (vm2-) abolishes its capacity to rescue muscle arm defects (B). Boxed areas indicate synaptic regions. Dashed lines indicate the morphology of vm2. Scale bar is 20 μm. (C) Quantification of the vm2 muscle arm phenotypes in the mosaic transgenic animals. *p<0.0001, Fisher's exact test, n = 48–50. (D) Vulval muscle morphology in the apx-1(wy755); lin-12(w750) mutant animals. Boxed area indicates synaptic region. Arrows indicate HSN cell bodies. Scale bar is 20 μm. (E) Epifluorescence and DIC images showing the apx-1 expression pattern in young adult. The four cells in the center of the image are 2° vulval epithelial cells. The four cells at the periphery are vm1 cells. Scale bar is 20 μm. (F) Cell-autonomous requirement of apx-1, sel-12 and lag-1. Quantification of muscle arm phenotypes in animals with different genotypes indicated on the X-axis. *, P<0.0001, Fisher's exact test, n = 40–57.
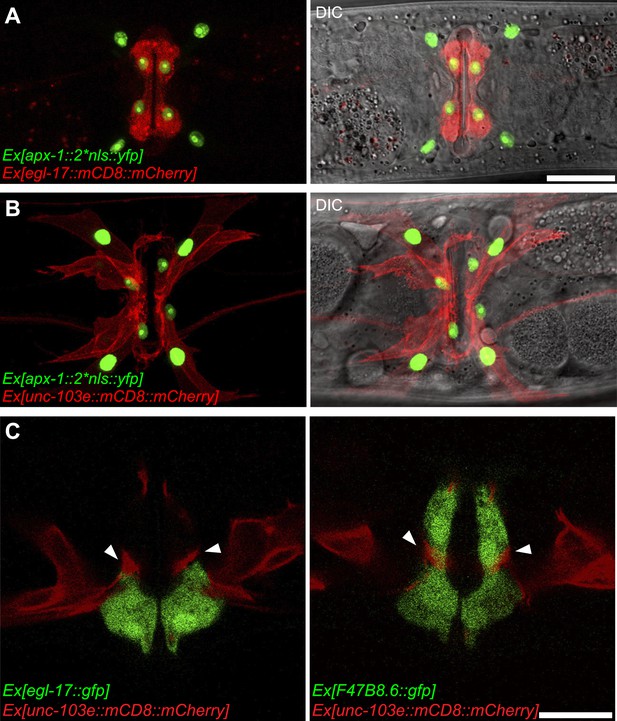
APX-1 expression pattern and the relative position of vm2 muscle arms.
(A) Overlaid epifluorescence and DIC images showing the identity of the apx-1 expressing cells. The egl-17::mCD8::mCherry transgene labels the 2° vulval epithelial cells in young adult. Scale bar is 20 μm. (B) Overlaid epifluorescence and DIC images showing the identity of the apx-1 expressing cells. The unc-103e::mCD8::mCherry transgene labels the vm1 cells in young adult . Scale bar is 20 μm. (C) Single confocal slices showing the position of muscle arms and the vulval epithelial cells. Vulval muscles are labeled by mCD8::mCherry. Secondary vulval epithelial cells are labeled by egl-17::gfp transgene (left). Both primary and secondary vulval epithelial cells are labeled by F47B8.6::gfp transgene (right). Note that the muscle arms are located between the 1o and the 2o vulval epithelial cells and directly contact these cells. Arrowheads indicate muscle arms. Scale bar is 10 μm.
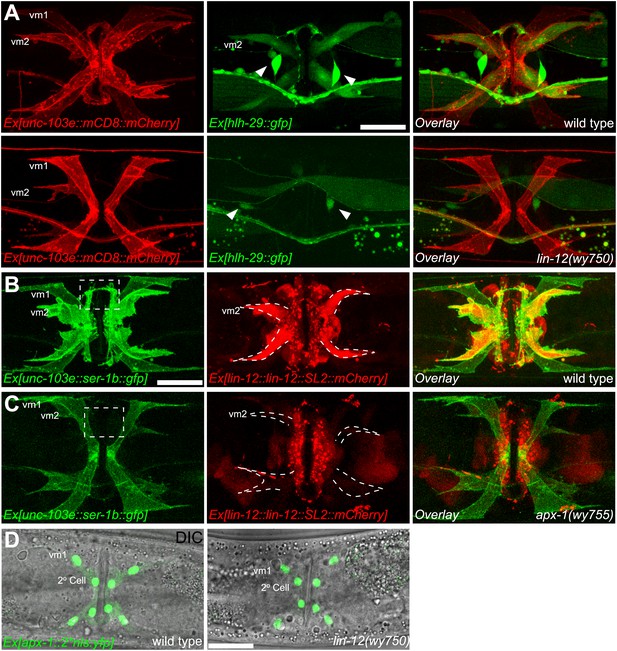
vm2 muscle arm development requires APX-1-induced LIN-12/Notch signaling activity.
(A) hlh-29::GFP expression in wild-type (top) and lin-12(wy750) (bottom) animals. Left column shows the vulval muscles labeled by mCD8::mCherry. Middle column shows the hlh-29::GFP expression patterns. Right column shows the overlaid images. Arrowheads indicate VC4/5 cell bodies. Note that hlh-29::GFP is specifically expressed in vm2 but not vm1 in the wild type, and is down-regulated in the lin-12(wy750) mutant. Scale bar is 20 μm. (B) and (C) lin-12::SL2::mCherry expression in wild-type (B) and apx-1(wy755) (C) animals. Left column shows the vulval muscles labeled by SER-1::GFP. Middle column shows the lin-12 expression patterns. Right column shows the overlaid images. Boxed areas indicate synaptic regions. Dashed lines delineate the morphology of vm2. Note that lin-12 transcription is specifically down-regulated in vm2 cells in apx-1(wy755) mutant animals. Scale bar is 20 μm. (D) apx-1 expression in wild-type (left) and lin-12(wy750) (right) animals. The four cells in the center of the image are 2o vulval epithelial cells. The four cells at the periphery are vm1 cells. Note that the apx-1 expression pattern does not change in the lin-12(wy750) mutant. Scale bar is 20 μm.
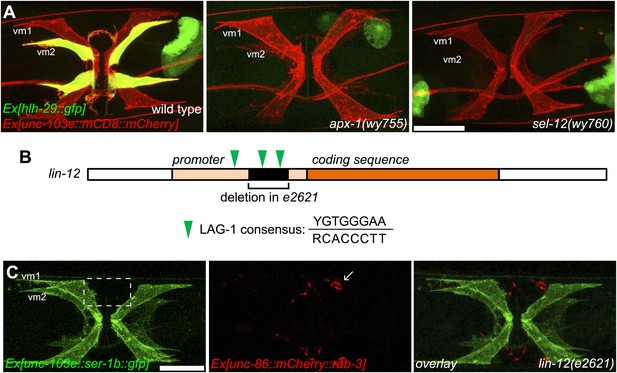
vm2 muscle arm development requires high LIN-12/Notch signaling activity.
(A) hlh-29::gfp expression in wild-type (left), apx-1(wy755) (middle) and sel-12(wy760) (right) mutant animals. Vulval muscles are labeled by mCD8::mCherry. Note that hlh-29::gfp is specifically expressed in vm2 but not vm1 in the wild type, and is down-regulated in the apx-1(wy755) and sel-12(wy760) mutants. Scale bar is 20 μm. (B) Schematic illustration of the molecular lesion (top) and muscle arm phenotype (bottom) of lin-12(e2621). lin-12 promoter and coding sequence are indicated by yellow and orange colors. Black box and bracket show the deletion in e2621. LAG-1 consensus binding sites are indicated by green arrowheads. Note that there are two LAG-1 consensus binding sites located in the deletion region. Boxed area indicates synaptic region. Arrow indicates HSN cell body. Scale bar is 20 μm.
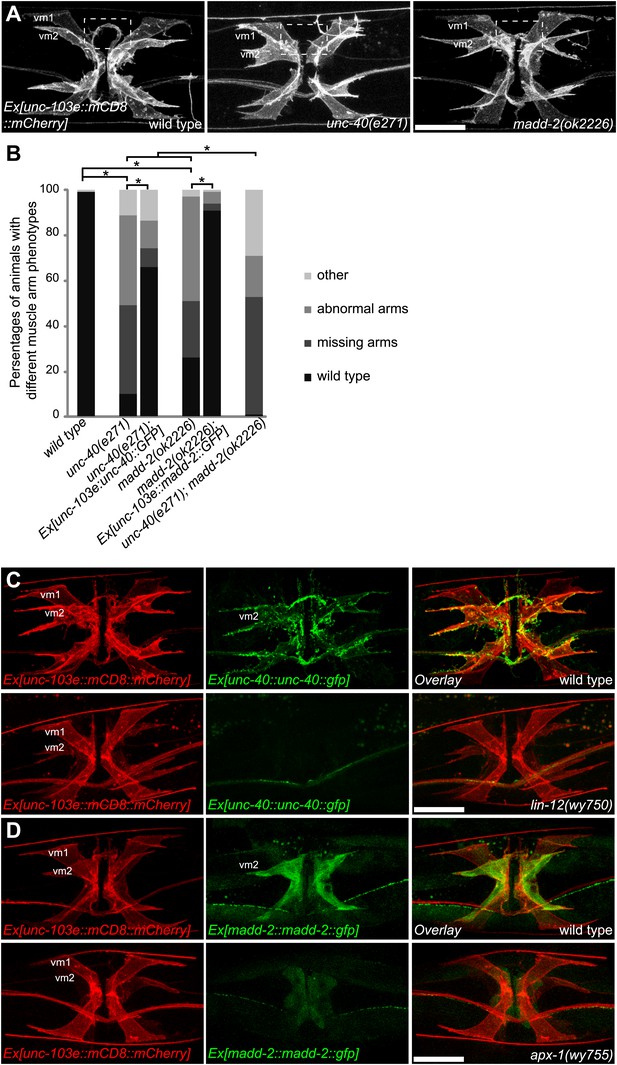
LIN-12/Notch signaling instructs vm2 muscle arm by regulating unc-40/DCC and madd-2.
(A) Representative images showing the vulval muscle morphology in wild type (left), unc-40(e271) (middle) and madd-2(ok2226) (right) animals. Boxed areas indicate synaptic regions. Note that unc-40(e271) and madd-2(ok2226) show missing (middle) or abnormal (right) muscle arm phenotypes. Scale bar is 20 μm. (B) Quantification of muscle arm phenotypes in animals with different genotypes indicated on the X-axis. ‘Wild type’ indicates the animals with normal muscle arms. ‘Missing arms’ indicates the animals with missing muscle arms. ‘Abnormal arms’ indicates the animals with flimsy muscle arms or abnormal muscle arms extending from incorrect positions. ‘Other’ indicates animals with severe vulval muscle morphology phenotype that the muscle arms could not be scored. *p<0.0001, chi-squared test, n = 73–194. (C) Double-labeling of the expression of UNC-40 and vulval muscle morphology in wild type (top) and lin-12(wy750) animals (bottom). Left column shows the vulval muscles labeled by mCD8::mCherry. Middle column shows the UNC-40::GFP expression patterns. Right column shows the overlaid images. Note that UNC-40::GFP is preferentially expressed in vm2 and enriched on the muscle arms in the wild type, and is down-regulated in the lin-12(wy750) mutant. Scale bar is 20 μm. (D) Double-labeling of the expression of MADD-2 and vulval muscle morphology in wild type (top) and apx-1(wy755) animals (bottom). Left column shows the vulval muscles labeled by mCD8::mCherry. Middle column shows the MADD-2::GFP expression patterns. Right column shows the overlaid images. Note that MADD-2::GFP is preferentially expressed in vm2 in the wild type, and is down-regulated in the apx-1 (wy755) mutant. Scale bar is 20 μm.
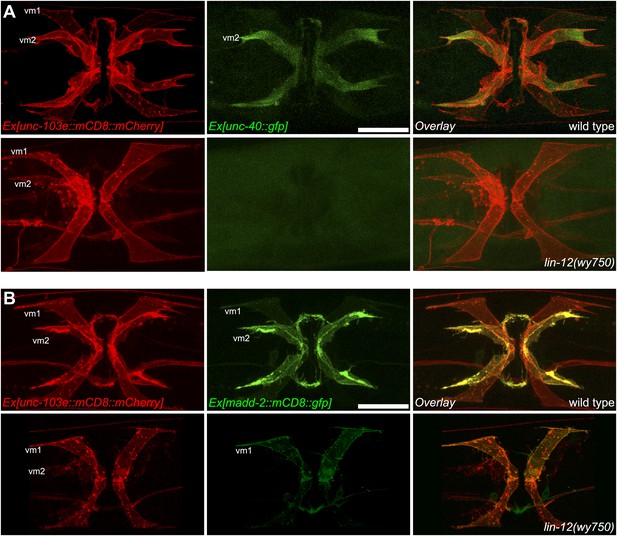
LIN-12/Notch signaling instructs vm2 muscle arm by regulating unc-40/DCC and madd-2.
(A) Ventral views of the double-labeling of the transcription of unc-40 and vulval muscle morphology in wild type (top) and lin-12(wy750) animals (bottom). Left column shows the vulval muscles labeled by mCD8::mCherry. Middle column shows the unc-40 transcription patterns. Right column shows the overlaid images. Note that unc-40 is preferentially transcribed in vm2 in wild type, and down-regulated in lin-12(wy750) mutant. Scale bar is 20 μm. (B) Ventral views of the double-labeling of the transcription of madd-2 and vulval muscle morphology in wild type (top) and lin-12(wy750) animals (bottom). Left column shows the vulval muscles labeled by mCD8::mCherry. Middle column shows the madd-2 transcription patterns. Right column shows the overlaid images. Note that madd-2 is preferentially transcribed in vm2 in wild type, and down-regulated in lin-12(wy750) mutant. Scale bar is 20 μm.
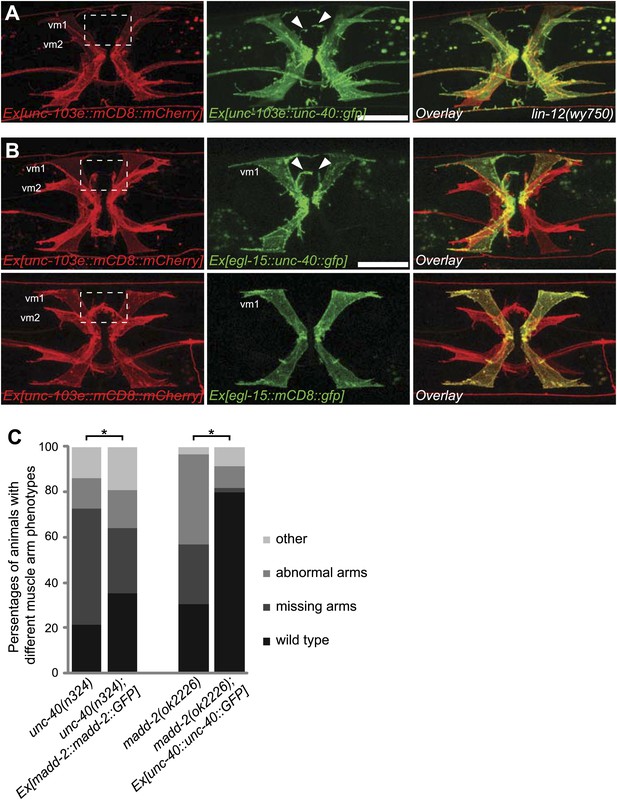
unc-40 and madd-2 function partly in parallel in regulating vm2 muscle arm development.
(A) Expression of UNC-40::GFP (middle) in vulval muscles in the lin-12(wy750) animals. Vulval muscles are labeled by mCD8::mCherry (left). Boxed area indicates synaptic region. Arrowheads indicate rescued muscle arms. Note that the forced expression of UNC-40 in vm2 from the unc-103e promoter induces the formation of muscle arms in the lin-12(wy750) mutant. Scale bar is 20 μm. (B) Ectopic expression of unc-40::gfp (top) or mCD8::gfp (bottom) transgene in vm1. Left column shows the vulval muscles labeled by mCD8::mCherry. Middle column shows forced transgene expression patterns. Right column shows the overlaid images. Boxed areas indicate synaptic regions. Arrowheads indicate ectopic muscle arm-like structures. Note that the ectopic expression of UNC-40 in vm1 induces the muscle arm-like membrane extensions. Scale bar is 20 μm. (C) Quantification of muscle arm phenotype in animals with different genotypes indicated on the X-axis. ‘Wild type’ indicates the animals with normal muscle arms. ‘Missing arms’ indicates the animals with missing muscle arms. ‘Abnormal arms’ indicates the animals with flimsy muscle arms or abnormal muscle arms extending from incorrect positions. ‘Other’ indicates animals with severe vulval muscle morphology phenotype that the muscle arms could not be scored. *p<0.0001, chi-squared test, n = 105–311.
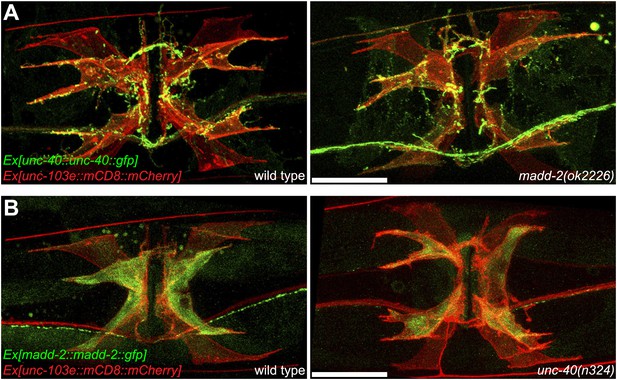
Ectopic expression of UNC-40 in vm1 induces muscle arm-like structures.
(A) UNC-40 expression in the wild-type (left) and madd-2(ok2226) (right) animals. Vulval muscles are labeled by mCD8::mCherry. Note that in both wild-type and madd-2(ok2226) mutant animals, UNC-40 are preferentially expressed in vm2 cells. Scale bar is 20 μm. (B) MADD-2 expression in the wild-type (left) and unc-40(n324) (right) animals. Vulval muscles are labeled by mCD8::mCherry. Note that in both wild-type and unc-40(n324) mutant animals, MADD-2 are preferentially expressed in vm2 cells. Scale bar is 20 μm.
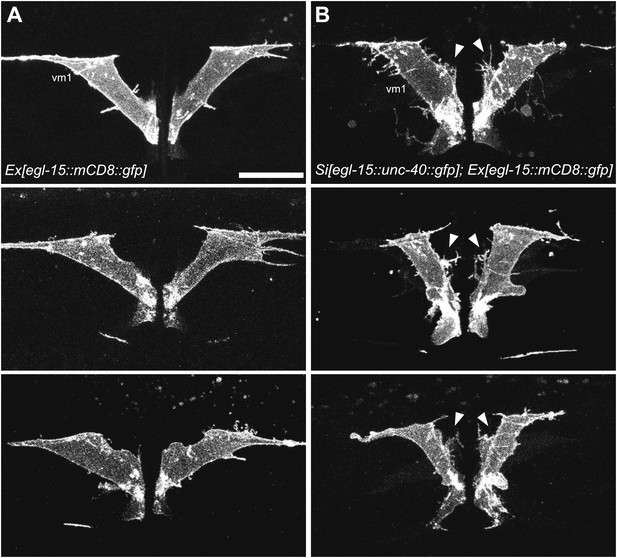
unc-40 and madd-2 are not mutually regulated.
(A) Three examples showing the vm1 morphology in animals with the expression of a single-copy transgene egl-15::unc-40::gfp. vm1 is visualized by egl-15::mCD8::gfp. Arrowheads indicate the induced muscle arm-like membrane protrusions. Scale bar is 20 μm.
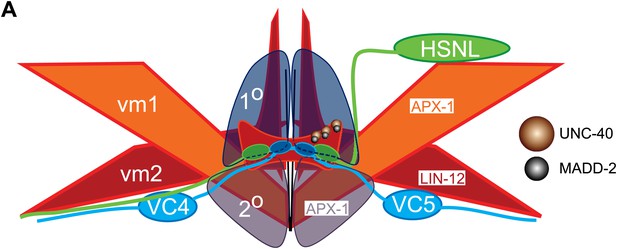
Schematic model of muscle arm development and vm2 postsynaptic specification.
(A) Schematic model showing the physical organization of the vulva organ and the mechanism of vm2 postsynaptic target selection. Note that vm2 muscle arms (red) directly contact the primary (1°, dark blue) and secondary (2°, dark gray) vulval epithelial cells. APX-1/DSL is expressed in vm1 cells (orange) and the secondary vulval epithelial cells. LIN-12/NOTCH is expressed in vm2 cells (red). UNC-40/DCC (brown circles) and MADD-2 (black circles) interact with each other and are localized on vm2 muscle arms.
Videos
Synchronous Ca2+ transients in the vulval muscles of wild-type animals drive egg-laying behavior. Intensity-modulated ratiometric imaging of wild-type C. elegans expressing the Ca2+ sensor GCaMP3 and soluble mCherry in the vulval muscles from the unc-103e promoter at 20 fps. Change in the GCaMP3 to mCherry fluorescence ratio is indicated by a rainbow scale from 0 (dark blue) to 1.2 (red). Egg-laying events are observed at 23 and 56 s. Still images from this video are shown in Figure 2C, and the traces of ΔR/R and vulval muscle area are shown in Figure 2E.
lin-12 mutants have asynchronous vulval muscle Ca2+ transients leading to uncoordinated contractions. Intensity-modulated ratiometric imaging of lin-12(wy750) mutants expressing the Ca2+ sensor GCaMP3 and soluble mCherry in the vulval muscles from the unc-103e promoter at 20 fps. Change in the GCaMP3 to mCherry fluorescence ratio is indicated by a rainbow scale from 0 (dark blue) to 1.2 (red). Still images from this video are shown in Figure 2D, and the traces of ΔR/R and vulval muscle area are shown in Figure 2F.
Asynchronous vulval muscle Ca2+ transients in lin-12 and apx-1 mutants can still lead to uncoordinated vulval opening and egg laying. Intensity-modulated ratiometric imaging of apx-1(wy755) mutants expressing the Ca2+ sensor GCaMP3 and soluble mCherry in the vulval muscles from the unc-103e promoter at 20 fps. Change in the GCaMP3 to mCherry fluorescence ratio is indicated by a rainbow scale from 0 (dark blue) to 1.2 (red). At 6 and 18 s, there was sufficient vulval opening to permit an egg-laying event.
Tables
Mutants isolated from screen for genes required for vm2 postsynaptic target selection
| Complementation group | Allele | Mutation | Homolog |
| lin-12 | wy750 | G473R(g1417a) | Notch |
| apx-1 | wy754 | G208E(g623a) | Dsl |
| wy755 | G159E(g476a) | ||
| wy766 | C217Y(g650a) | ||
| sel-12 | wy756 | W184stop(g552a) | Presenilin |
| wy760 | G373D(g1118a) |





