A novel sphingolipid-TORC1 pathway critically promotes postembryonic development in Caenorhabditis elegans
Figures
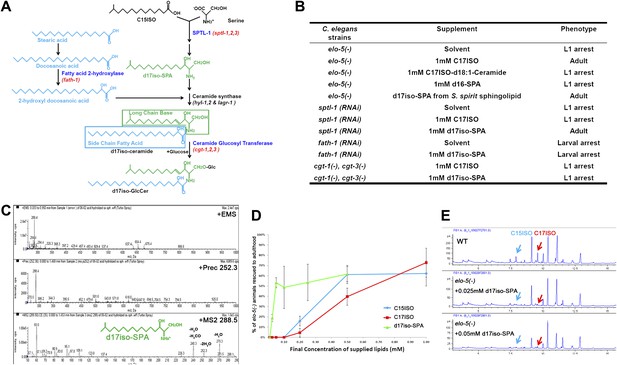
Iso-branched d17iso-sphinganine rescues elo-5(-) L1 arrest.
(A) Sphingolipid biogenesis pathway in C. elegans, including the catalytic enzymes (blue) and corresponding genes (red) used in this study. Molecular structures in green and in light blue indicate the long chain base and side chain fatty acid, respectively. (B) Summary of growth phenotype of various mutants with indicated lipid supplementations. elo-5(gk208), cgt-1(tm1027), and cgt-3(tm504) were the alleles of the (−) mutants. LCB: long chain base; SPA: sphinganine. See Supplementary file 1 for numerical data or more detailed description of the phenotypes. (C) Mass spectrometry of iso-branched d17iso-sphinganine (d17iso-SPA) isolated from the bacteria S. spiritivorum. The major peak (m/z = 288.4) corresponds to d17iso-SPA in each panel. The lower panel shows fragmentation of d17iso-SPA by MS-MS scan. The fragment peaks are labeled with the name of lost residues. (D) d17iso-SPA rescues elo-5(−) animals more efficiently than C15ISO and C17ISO. Error bar, SD. (E) Gas chromatography of methyl-esterified fatty acid extracts from rescued elo-5(−) animals fed with d17iso-SPA. Low concentration of d17iso-SPA (second and third panels) supplement did not restore C15iso or C17iso fatty acid levels.
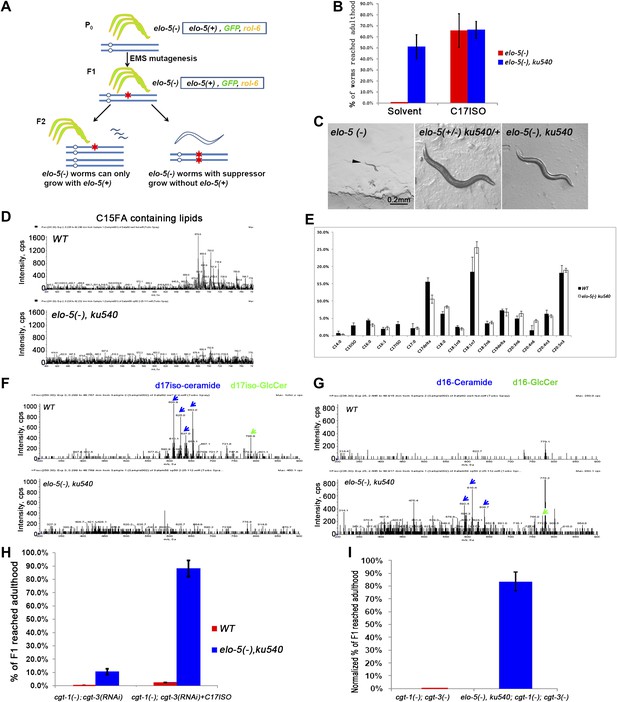
ku540 mutant suppresses L1 arrest of mmBCFA and sphingolipid biosynthetic mutants without restoring the levels of mmBCFAs.
(A) Screening strategy to isolate elo-5(−) suppressors. Green-colored C. elegans carry the extrachromosomal array containing copies of the elo-5(+), sur-5-gfp, and rol-6(dn) genes. (B) Percentages of animals reaching adulthood (bypass L1 arrest) when fed with/without C17ISO supplement. elo-5(−);ku540 mutant animals reach adulthood without C17ISO supplement. Error bar, SD. (C) Images showing that elo-5(−) animals arrested at L1 and elo-5(−);ku540 animals reached adulthood. (D) Mass spectrometry by precursor scan m/z = −241.2 showing no detectable C15FA-containing lipids in elo-5(−);ku540 animals. Numeric data for the levels of PA and PE are shown in Figure 2—figure supplement 1A. (E) Bar graph of relative FA composition by gas chromatography (GC) of methyl-esterified fatty acid extracts. These data indicate no detectable C15ISO or C17ISO was restored in elo-5(−) ku540 animals. GC graph is shown in Figure 2—figure supplement 1B. (F) and (G) Mass spectrometry of d17iso-ceramide–containing lipids by precursor scan m/z = +250.3, and d16-ceramide–containing lipids by precursor scan m/z = +236.3 (F). d17iso-ceramides and d17iso-glucosylceramides are detectable in wild-type but not elo-5(−);ku540 animals (G). In contrast, d16-ceramides and d16-glucosylceramides are present in elo-5(−);ku540, but not wild-type, animals. Numeric data for relevant peaks are shown in Figure 2—figure supplement 1C,D. (H) and (I) Percentages of animals of the indicated genotypes and treatment that reached adulthood. Error bar, SD. (H) The addition of ku540 dramatically suppressed the L1 arrest phenotype of cgt-1(−);cgt-3(RNAi) with and without the C17ISO supplement. The inclusion of elo-5(−) was due to its close linkage with ku540. When C17ISO was added to remove the negative effect of elo-5(−), about 90% of elo-5(−);ku540; cgt-1(−);cgt-3(RNAi) animals reached adulthood. C17ISO itself does not rescue the phenotype. (I). ku540 suppressed the L1 arrest phenotype the cgt-1(−);cgt-3 (−) double mutants (83.4%, n = 126). In this particular test, elo-5(−) ku540;cgt-1(−);cgt-3(−) homozygous animals were the progeny of elo-5(−/+) ku540(−/+);cgt-1(−/+);cgt-3(−) heterozygous mothers, and the presented data were obtained after normalizing against heterozygous populations (see ‘Materials and methods’). In all the other tests shown in Figure 2, elo-5(−) ku540 homozygous animals were the progeny of homozygous mothers.
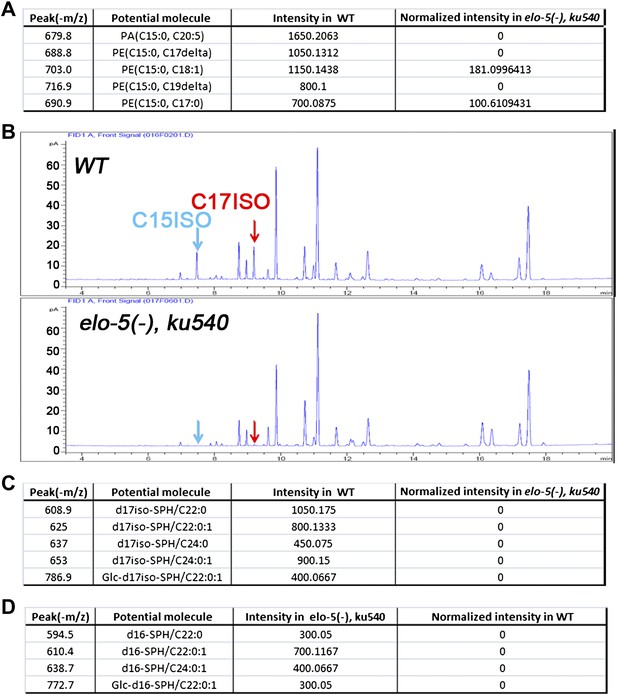
Quantification of GC and mass spectrometry analyses of elo-5(−) ku540 animals.
(A) Tables show relative intensities of the five strongest signals by mass spectrometry precursor scan (m/z = −241.2) in wild-type animals (second right column) and their normalized relative intensities in elo-5(−) ku540 animals (right column). The low intensities of these lipids in elo-5(−) ku540 animals indicate that the C15ISO mmBCFA level was not restored elo-5(−) ku540 animals. (B) Gas chromatography of methyl-esterified fatty acid extracts showing no detectable C15ISO or C17ISO in elo-5(−) ku540 animals. (C) Table shows relative intensities of normal d17iso-sphingolipid by mass spectrometry precursor scan (m/z = +250.3) in wild-type animals (second right column) and their normalized relative intensities in elo-5(−) ku540 animals (right column). The low intensities of these lipids in elo-5(−) ku540 animals indicate that the d17iso-sphingolipid level was not restored elo-5(−) ku540 animals. (D) Table shows relative intensities of abnormal d16-sphingolipid by mass spectrometry precursor scan (m/z = +238.3) in elo-5(−) ku540 animals (second right column). These sphingolipids are not present in wild-type animals (right column).
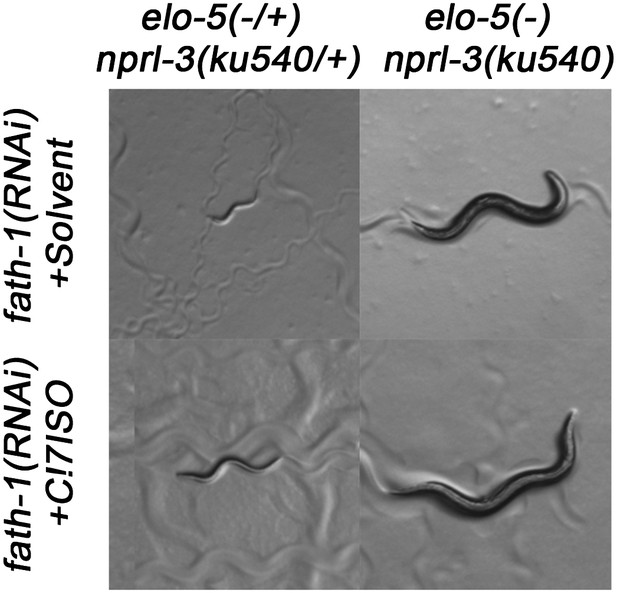
Suppression of fath-1(−) by nprl-3(ku540).
Microscopic images showing that nprl-3(ku540) suppresses the L1 arrest phenotype of fath-1 (injection RNAi) with or without C17ISO supplementation.
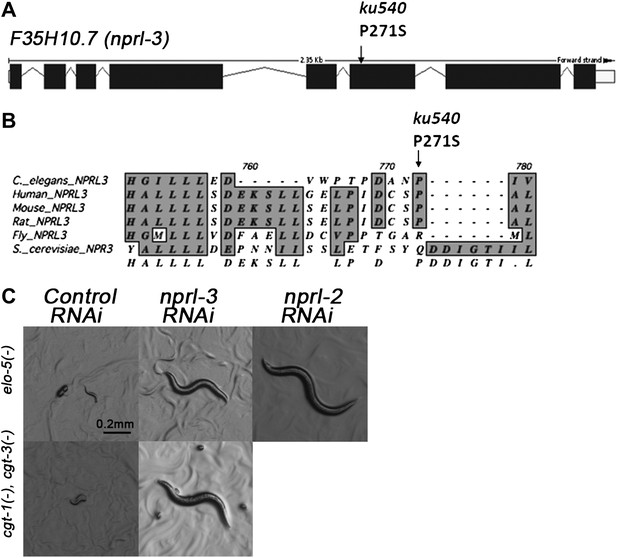
ku540 is a loss-of-function missense mutation of nprl-3.
(A) Predicted structure and position of the ku540 mutation in the nprl-3 gene (F35H10.7). (B) Abbreviated alignment of C. elegans NPRL-3 with its orthologs in other organisms. (C) C. elegans images showing nprl-3(RNAi) mimics the effect of the ku540 mutation in rescuing the L1 arrest phenotype caused by blocking mmBCFA or glucosyl-ceramide biosynthesis. nprl-3 dsRNA injection rescued 61.0% of elo-5(−) (n = 123) and 6.8% of cgt-1(−) cgt-3(−) (n = 71) animals to or beyond L3 stage. nprl-2 dsRNA injection also rescued 42.4% of elo-5(−) animals to or beyond L3 stage (n = 128). All data have been normalized to heterozygous populations (see ‘Materials and methods’).
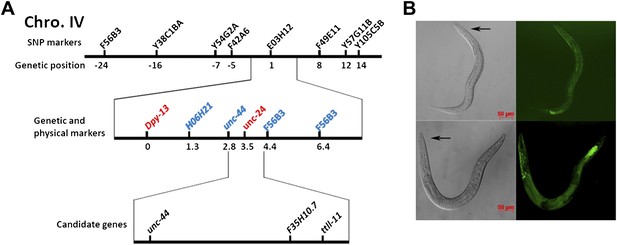
Mapping and expression of nprl-3.
(A) A simplified diagram of the mapping process. ku540 is genetically linked to the E03H12 SNP marker on chromosome IV (upper section). Further three-point mapping narrowed the ku540 locus to near unc-24, by visible markers (red) and by physical mutations (blue) found by genomic deep sequencing (middle section). The three candidate gene mutations resulting in amino acid changes are shown (lower section). (B) DIC and fluorescence images showing two representative animals carrying a nprl-3 promoter::GFP transgene. The head is indicated by a black arrow. GFP is visible in most tissues with stronger expression seen in the head and tail regions.
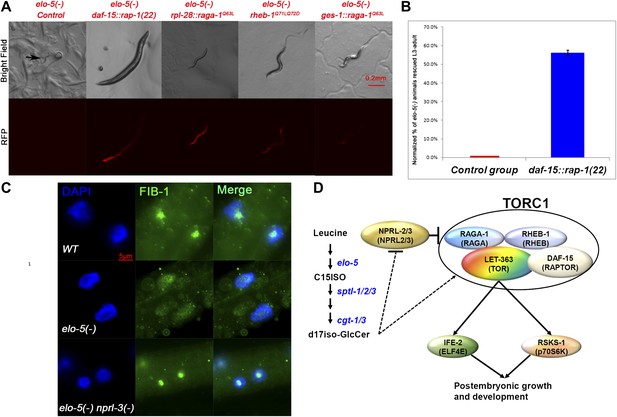
TORC1 activation is sufficient for mmBCFAs-mediated growth regulation.
(A) Representative florescent images showing that elo-5(−) animals with each of four RFP-marked transgenes, which constitutively activated TORC1, bypass L1 arrest to reach beyond L3 stage (statistical data are described in the text and Figure 4B). The rpl-28 promoter drives ubiquitous expression, whereas the ges-1 promoter drives the expression specifically in the intestine (Edgar and McGhee, 1986). Arrow in the upper left panel marks an arrested L1. (B) Percentage of homozygous elo-5(−) animals carrying the daf-15::rap-1(22) transgene reached L3-adult stages (n = 161). The data were normalized against that of elo-5(−/+) heterozygous animals. Error bar, SD. (C). Immunofluorescence images showing FIB-1 expression and localization in intestinal cells of L3 larvae. DAPI-stained nuclei are blue. Green fluorescence indicates the staining of antibody against FIB-1. FIB-1 localization in the nucleoli is largely abolished in elo-5(−) animals and restored by the nprl-3(−) mutation (the percentages of condensed nucleoli localization for the three genotypes from top to bottom are 95% [n=42], 24% [n=112] and 93% [n=68]). (D) A model for the regulation of postembryonic growth and development by mmBCFAs and GlcCer.
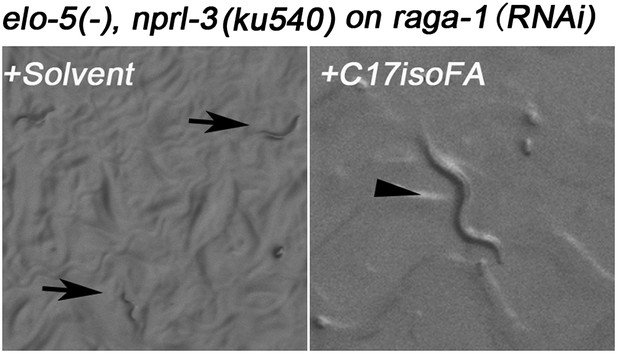
Microscopic images of C. elegans with raga-1(RNAi) treatment.
Animals without C17ISO supplement were developmentally arrested at L1 stage (arrows). Animals with C17ISO supplement reached adulthood (arrowhead), suggesting that the arrest depends on mmBCFA deficiency and is not caused by RNAi itself.
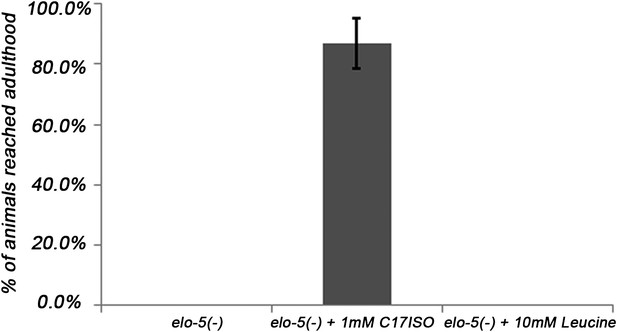
Leucine could not promote postembryonic development independent of the mmBCFA/d17isoGlcCer/TORC1 pathway.
Normalized percentages of elo-5(−) animals that reached adulthood on various supplements. While 1 mM C17ISO could suppress the L1 arrest of elo-5(−), 10 mM leucine could not. Error bar: SD.
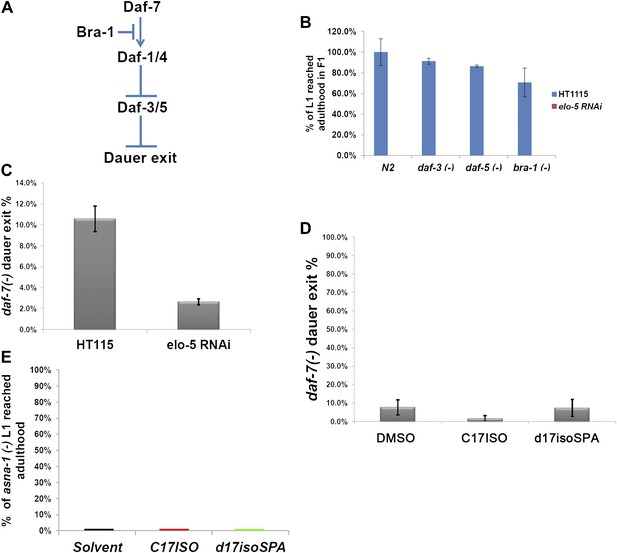
mmBCFA/GlcCer/TORC1 pathway is independent of the DAF-7/TGF-β pathway.
(A) Cartoon illustration of a simplified DAF-7/TGF-β pathway in C. elegans. Mutations in daf-3, daf-5, or bra-1 have been shown to cause constitutive activity of the TGF-β pathway and suppress dauer formation (Patterson and Padgett, 2000). (B) Percentages of animals of the indicated genotypes that reached adulthood on elo-5 (RNAi) plates. Mutation in none of these negative regulators of the daf-7 pathway permitted the elo-5(RNAi)–treated animals to bypass L1 arrest, suggesting that the TGF-β pathway does not act downstream of mmBCFAs. (C) Percentages of daf-7(−) animals that exited dauer stage to reach adulthood on indicated RNAi plates. elo-5(RNAi) enhanced the constitutive dauer formation phenotype of a daf-7(−) mutant. (D) Percentages of daf-7(−) animals that exited dauer stage to reach adulthood on various branched lipid supplements. Neither C17ISO nor d17iso-SPA could suppress the constitutive dauer formation of daf-7(−), suggesting that DAF-7/TGF-β does not act upstream of mmBCFAs or d17iso-sphingolipid. (E) Percentages of asna-1(−) animals that reached adulthood on plates with various branched-chain lipid supplements. asna-1 encodes a protein required for proper DAF-7/TGF-β function and an asna-1(−) mutation causes L1 arrest (Kao et al., 2007). Neither C17ISO nor d17iso-SPA permits asna-1(−) animals to bypass L1 arrest.
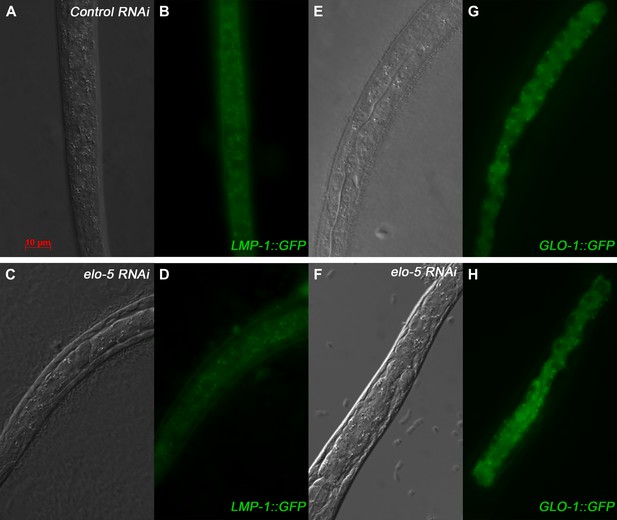
Lysosome integrity is not disrupted in mmBCFA-deficient animals.
(A)–(H) DIC and GFP images illustrating LMP-1::GFP (A–D) and GLO-1::GFP (E–H) expression patterns are similar throughout the intestine of young wild-type or elo-5(RNAi) larvae. Both LMP-1::GFP (A–D) and GLO-1::GFP are lysosomal markers in C. elegans. These data indicate that lysosomal integrity is not disrupted in mmBCFA-deficient animals.
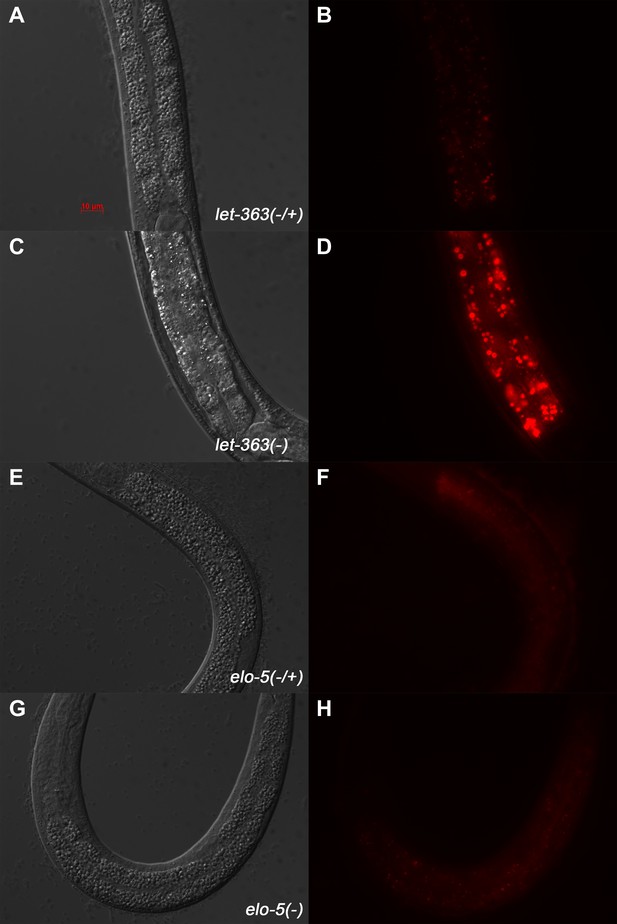
Neutral red staining of let-363 -and elo-5-deficient animals.
(A)–(H) DIC and Rhodamine channel fluorescence images of larvae stained with Neutral red. (A–D) let-363 homozygous L3 animals have increased size and intensity of Neutral red stained lysosomes (D) compared to the heterozygous control (B). (E–H) elo-5 homozygous L1 arrested animals show similar Neutral red intensity and staining pattern (H) when compared to the heterozygous control (F). These data indicate that (1) lysosome integrity is not disrupted in elo-5(−) animals, and (2) unlike let-363(−), elo-5(−) animals do not show increased Neutral red staining in the intestine.
Tables
Intact TORC1 function is necessary for mmBCFA-mediated growth regulation
| Genotype | RNAi | Dietary C17ISO | Normalized % of F1 reached adulthood | N | p |
| elo-5(−) | Vector | − | 0 | 208 | |
| elo-5(−) | Vector | + | 86.9 | 205 | 0 |
| elo-5(−) nprl-3(−) | Vector | − | 71.5 | 69 | |
| elo-5(−) nprl-3(−) | Vector | + | 111.5 | 103 | 0.24 |
| elo-5(−) nprl-3(−) | raga-1 (a) | − | 35.5 | 56 | |
| elo-5(−) nprl-3(−) | raga-1 (a) | + | 102.5 | 83 | 0.031 |
| elo-5(−) nprl-3(−) | raga-1 (b) | − | 25.5 | 214 | |
| elo-5(−) nprl-3(−) | raga-1 (b) | + | 88.0 | 279 | 0.00001 |
| elo-5(−) nprl-3(−) | rheb-1 (a) | − | 14.2 | 351 | |
| elo-5(−) nprl-3(−) | rheb-1 (a) | + | 64.6 | 526 | 0.0000001 |
| elo-5(−) nprl-3(−) | rheb-1 (b) | − | 32.5 | 201 | |
| elo-5(−) nprl-3(−) | rheb-1 (b) | + | 86.0 | 116 | 0.0037 |
| elo-5(−) nprl-3(−) | rsks-1 (a) | − | 15.0 | 68 | |
| elo-5(−) nprl-3(−) | rsks-1 (a) | + | 72.5 | 83 | 0.022 |
| elo-5(−) nprl-3(−) | rsks-1 (b) | − | 19.0 | 183 | |
| elo-5(−) nprl-3(−) | rsks-1 (b) | + | 61.0 | 180 | 0.0034 |
| elo-5(−) nprl-3(−) | ife-2 (a) | − | 10.5 | 144 | |
| elo-5(−) nprl-3(−) | ife-2 (a) | + | 71.5 | 49 | 0.003 |
| elo-5(−) nprl-3(−) | ife-2 (b) | − | 0 | 103 | |
| elo-5(−) nprl-3(−) | ife-2 (b) | + | 103.5 | 150 | 0 |
| elo-5(−) nprl-3(−) | let-363 | − | 0 | >100 | |
| elo-5(−) nprl-3(−) | let-363 | + | 0 | >100 | NA |
-
Percentages of elo-5(−);nprl-3(ku540) homozygotes with the indicated RNAi treatments that reached adulthood, where (a) and (b) indicate two different RNAi constructs targeting different parts of the same gene. The presented percentage of elo-5(−) nprl-3(ku540) animals that reached adulthood was calculated by normalizing against the percentage of elo-5(−/+) nprl-3(ku540)/+ heterozygotes (see ‘Materials and methods’ for detail). Without C17ISO supplementation, RNAi knockdown of multiple TORC1 components reverted elo-5(−);nprl-3(ku540) animals to larval arrest.
Additional files
-
Supplementary file 1
Detailed description of phenotypes listed in Figure 1B.
- https://doi.org/10.7554/eLife.00429.016





